Castanea sativa is one of the main multipurpose tree species valued for its timber and nuts. This species is susceptible to two major diseases, ink disease and chestnut blight, caused by Phytophthora spp. and Cryphonectria parasitica, respectively. The loss-of-function mutations of genes required for the onset of pathogenesis, referred to as plant susceptibility (S) genes, are one mechanism of plant resistance against pathogens. On the basis of sequence homology, functional domain identification, and phylogenetic analyses, we report for the first time on the identification of S-genes (mlo1, dmr6, dnd1, and pmr4) in the Castanea genus. The expression dynamics of S-genes were assessed in C. sativa and C. crenata plants inoculated with P. cinnamomi and C. parasitica. Our results highlighted the upregulation of pmr4 and dmr6 in response to pathogen infection. Pmr4 was strongly expressed at early infection phases of both pathogens in C. sativa, whereas in C. crenata, no significant upregulation was observed. The infection of P. cinnamomi led to a higher increase in the transcript level of dmr6 in C. sativa compared to C. crenata-infected samples. For a better understanding of plant responses, the transcript levels of defense genes gluB and chi3 were also analyzed.
- chestnut
- susceptibility genes
- Phytophthora cinnamomi
- Cryphonectria parasitica
1. Introduction
Castanea
Castanea sativa
Castanea crenata
Castanea mollissima
Castanea dentata
C. sativa
2. History
Over the last century, the number of chestnut trees decreased in growing areas in Europe due to the depopulation of mountains, climate change, and the spread of two severe diseases: ink disease and chestnut blight [2][3]. Ink disease is caused by the Oomycete
Phytophthora cinnamomi
Phytophthora cambivora
C. sativa
P. cinnamomi
Castanea
C. crenata
P. cinnamomi [6]. The disease, which affects both young and old trees, leads to subcortical necrosis of the root system and the basal part of the stem; this is followed by the appearance of wasting symptoms in the foliage until the total desiccation and death of the plant occur [7][8][9][10]. These pathogens spread mainly through the movement of soil harboring inoculum and the dissemination of asexual flagellated spores (i.e., zoospores) that can actively travel short distances or passively travel long distances in flowing water [10][11]. The use of resistant rootstocks represents one possible solution to protect against these pathogens, although, at present, only tolerant selections obtained from hybridization between
C. sativa
C. crenata
Chestnut blight stands among the most destructive fungal tree diseases ever [10][13]. The causal agent,
Cryphonectria parasitica, infects trees through dead plant tissue and wounds, including those caused by pruning, graft, and hail [13][14]. The symptoms involve bark cankers that can develop on suckers, young branches, and adult branches and trunks [15]. Chestnut blight was one of the causes of the abandonment of chestnut orchards in Europe until the end of the 1970s, when the natural spread of the hypovirulent form of the fungus favored a slow but progressive recovery of chestnut orchards and coppices. However, the fungus still represents a relevant problem in many areas of Europe. It is very harmful to young grafted trees in particular, hampering the establishment of new orchards in many areas [10][13].
C. dentata
C. parasitica
C. mollissima [17][18]. More recently, researchers discovered that the onset of the disease is associated with the release of oxalic acid by the pathogen during infection. Blight-resistant
C. dentata
Dryocosmus kuriphilus Yasumatsu) infestation, which only recently has been controlled effectively [1][21], and to the general difficulty of developing a modern chestnut industry based on quality cultivars of
C. sativa
Mildew resistance locus O
mlo1
Powdery mildew resistance 4
pmr4
Downy Mildew Resistance 6
dmr6
Defense no death
dnd1
Mlo
mlo
mlo
Hordeum vulgare
mlo
Solanum lycopersicum
Pisum sativum
Capsicum annum
Nicotiana tabacum
Triticum aestivum L.) [26][27] plants. The callose synthase encoded by
pmr4
pmr4
Oidium neolycopersici
Phytophthora infestans tolerance [28][29].
Dmr6
Phythophthora capsici
Hyaloperonospora arabidopsidis
Pseudomonas syringae
dnd1
P. infestans
mlo
C. sativa
C. sativa
C. crenata
P. cinnamomi
C. parasitica
chi3
acidic 26 kDa endochitinase
gluB
glucan endo-1,3-beta- glucosidase), were also determined. Our analysis revealed the strong activation of
pmr4
dmr6 genes
P. cinnamomi
C. parasitica
3. Development and Findings
C. sativa
is a European woody tree species commonly used across the globe in the food and timber industries. This chestnut species is susceptible to the two major pathogens,P. cinnamomi
andC. parasitica [10][35]. In contrast, the Asian chestnut species
[10,36]. In contrast, the Asian chestnut speciesC. crenata
andC. mollissima
exhibit higher tolerance toP. cinnamomi
andC. parasitica [6]. Achieving tolerance or resistance to pathogens is the major aim of rootstock breeding. Blight-resistant trees were obtained through backcross breeding of introgression genes from Asian species into American chestnut trees. [36]. However, this approach, although successful in developing blight-resistant American chestnut selections has been slowed by a lack of genetic tools. In Europe, ink disease tolerant hybrids were obtained through interspecific crosses between
[6]. Achieving tolerance or resistance to pathogens is the major aim of rootstock breeding. Blight-resistant trees were obtained through backcross breeding of introgression genes from Asian species into American chestnut trees. [37]. However, this approach, although successful in developing blight-resistant American chestnut selections has been slowed by a lack of genetic tools. In Europe, ink disease tolerant hybrids were obtained through interspecific crosses betweenC. sativa
andC. crenata, although the nut quality produced by these hybrids is below current market standards [37][38].
, although the nut quality produced by these hybrids is below current market standards [38,39]. It has long been recognized that a deep understanding of a pathogen’s biology, host–pathogen interactions, and the resistance mechanisms are fundamental to improving breeding programs. Genomic and transcriptomic analyses have provided the first genetic insights into mechanisms underlying susceptible and resistant chestnut species responses toP. cinnamomi
andC. parasitica [36][37][39][40][41]. Santos et al. [39] reported the upregulation of a set of candidate genes (e.g.,
[37,38,40,41,42]. Santos et al. [40] reported the upregulation of a set of candidate genes (e.g.,Cast_Gnk2-like
andCalcium-dependent protein kinase
) afterP. cinnamomi
infection, which may trigger HR-like cell death inC. crenata cells. A significant number of genes involved in the defense against chestnut blight were identified [36].
cells. A significant number of genes involved in the defense against chestnut blight were identified [37].Traditionally, the introduction of resistance gene analogues into plants was the most promising approach to facilitate the acquisition of resistance. However, it did not prove to be durable enough because the widespread use of R genes caused the selection of pathogens capable of overcoming them [24]. Susceptibility (S) genes can be interesting candidates to be used in target breeding programs [22][23][24]. Furthermore, on the basis of previous studies, it was highlighted that the disabling of susceptibility genes may facilitate durable resistance since the pathogen needs to gain a new function to replace the lost host factor it was exploiting [42].
Traditionally, the introduction of resistance gene analogues into plants was the most promising approach to facilitate the acquisition of resistance. However, it did not prove to be durable enough because the widespread use of R genes caused the selection of pathogens capable of overcoming them [24]. Susceptibility (S) genes can be interesting candidates to be used in target breeding programs [22,23,24]. Furthermore, on the basis of previous studies, it was highlighted that the disabling of susceptibility genes may facilitate durable resistance since the pathogen needs to gain a new function to replace the lost host factor it was exploiting [43].In woody species, the investigation of S-genes has been performed only for MLO genes in rubber trees [32], poplar trees [33], apple trees, and grapevines [34][43]. Due to the absence of a
In woody species, the investigation of S-genes has been performed only for MLO genes in rubber trees [32], poplar trees [33], apple trees, and grapevines [34,44]. Due to the absence of aC. sativa
genome, highly similar S-genes were selected using theC. mollissima
v 1.1 genome. Based on the blastn survey, four loci with high similarity were identified in theC. mollissima genome and attributed to different subclasses of S-genes [31][44][45][46] due to the presence of specific domains:
genome and attributed to different subclasses of S-genes [31,45,46,47] due to the presence of specific domains:mlo1
,dmr6
,dnd1
, andpmr4
(, ). As previously observed [31], in the phylogenetic trees, monocot proteins formed a separate clade with respect to those of dicotyledonous species, supporting the hypothesis that an independent evolution occurred for these genes (). Quantitative PCR analysis has been carried out to identify the differential expression of candidate S-genes in response toP. cinnamomi
andC. parasitica
in the stems of a susceptible species,C. sativa
, and of a tolerant one,C. crenata
. Lesion analysis and DNA quantification of the pathogen (Figure 3 and
4 andFigure 4) confirmed the higher tolerance level of
6) confirmed the higher tolerance level ofC. crenata
in response to bothP. cinnamomi
andC. parasitica
infection. Our qPCR results highlighted the main upregulation ofpmr4
anddmr6
in response to infection by bothP. cinnamomi
andC. parasitica
. As expected, a greater increase in the transcription of these susceptibility genes was observed in the susceptible speciesC. sativa
. Remarkably,pmr4
was strongly expressed at early infection phases of both pathogens inC. sativa
; in the tolerantC. crenata
, significant upregulation was observed ( andFigure 6).
7).Pmr4
codifies for a callose synthase, which is necessary to create a physical barrier to avoid pathogen penetration and is also implicated in plant-triggered immunity suppression.Pmr4
is thus not only involved in the synthesis of callose, but it also acts as a negative regulator of the salicylic acid pathway [28].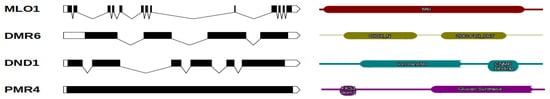
Figure 1. Chestnut S-genes and their protein structures. The graphical representations of gene exon/intron structures were generated using the http://wormweb.org/exonintron tool (accessed on 31 March 2021) and are shown in the left panel. The exons are indicated with black boxes, whereas introns are shown with lines. In the right panel, the protein structural domains are displayed.
Table 1. S-genes detected in the C. mollissima v1.1 genome and protein domain annotations.
| Gene Name | Scaffold | ORF Length (bp) | N° Exons | Size (aa) | Domains | PFAM DOMAINS |
|---|---|---|---|---|---|---|
| MLO1 | scaffold00101 | 1425 | 13 | 474 | Mlo | PF03094 |
| DMR6 | scaffold02358 | 1128 | 4 | 375 | 2OG-FeII_Oxy; DIOX_N | PF03171;PF14226 |
| DND1 | scaffold00410 | 1407 | 6 | 468 | cNMP_binding; Ion_trans | PF00027;PF00520 |
| PMR4 | scaffold00300 | 5346 | 1 | 1781 | FKS1_dom1; Glucan_synthase | PF14288;PF02364 |
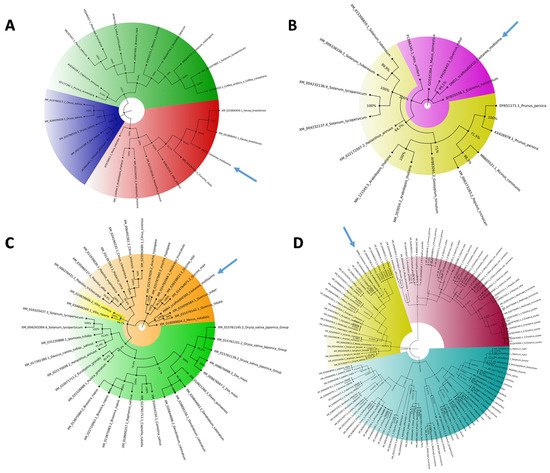
Figure 2. Phylogenetic analysis of the S-genes. The 4 phylogenetic trees of mlo1 (A), dnd1 (B), pmr4 (C), and dmr6 (D) were constructed using MEGAX software by aligning chestnut S-gene coding sequences with NCBI S-gene ortholog coding sequences. The colors indicate the main clades detected, and the arrows underline the location of C. mollissima.
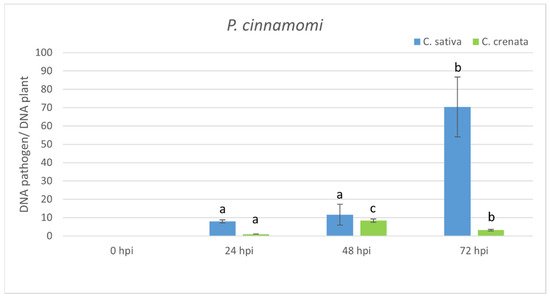
Figure 3. qRT-PCR pathogen DNA quantification after
P. cinnamomi inoculation. Data were quantified using the 2−ΔΔCt method based on the Ct values of pathogen genes (ypt and mf1) and actin-7 as a housekeeping gene. Data are the means of three biological replicates ± SE. C. sativa data are normalized with C. sativa 0 hpi control; C. crenata data are normalized with C. crenata 0 hpi control. Different letters associated with the set of means indicate a significant difference based on Tukey’s HSD test (p ≤ 0.05).
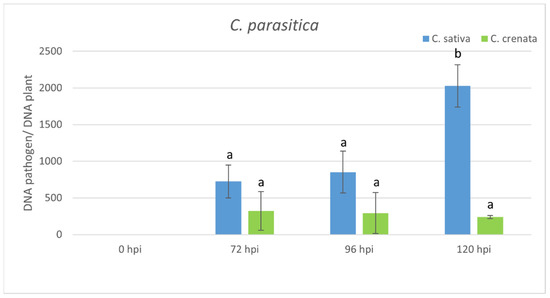
Figure 4. qRT-PCR pathogen DNA quantification after C. parasitica inoculation. Data were quantified using the 2−ΔΔCt method based on the Ct values of fungal genes (ypt and mf1) with actin-7 as a housekeeping gene. Data are the means of three biological replicates ± SE. C. sativa data are normalized with the C. sativa 0 hpi control; C. crenata data are normalized with C. crenata 0 hpi control. Different letters associated with the set of means indicate a significant difference based on Tukey’s HSD test (p ≤ 0.05).
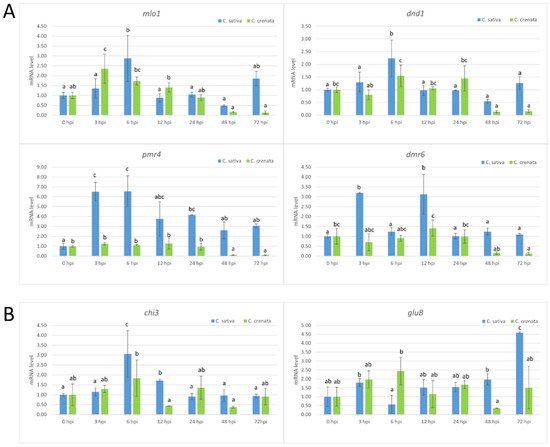
Figure 5. qRT-PCR-based transcription profiling after P. cinnamomi inoculation. (A) The S-gene transcription profiles in C. sativa (blue) and C. crenata (green) chestnut species. (B) The expression analysis of genes coding for several pathogenesis-related (PR) proteins in C. sativa (blue) and C. crenata (green) species. In all analyses, Cm7-actin was used as a housekeeping gene. Data are the means of three biological replicates ± SE. C. sativa data are normalized with C. sativa 0 hpi control; C. crenata data are normalized with C. crenata 0 hpi control. Different letters associated with the set of means indicate a significant difference based on Tukey’s HSD test (p ≤ 0.05).
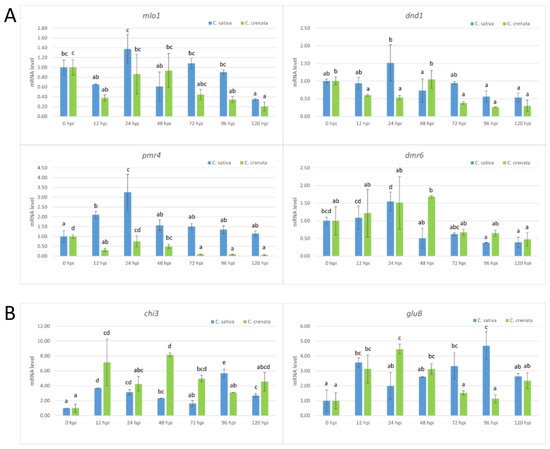
Figure 6. qRT-PCR-based transcription profiling after C. parasitica inoculation. (A) The S-gene transcription profile in C. sativa (blue) and C. crenata (green) chestnut species. (B) The expression analysis of genes coding for several pathogenesis-related (PR) proteins of C. sativa (blue) and C. crenata (green) species. In all the analyses, Cm7-actin was used as the housekeeping gene. The data are the means of three biological replicates ± SE. C. sativa data are normalized with C. sativa 0 hpi control; C. crenata data are normalized with C. crenata 0 hpi control. Different letters associated with the set of means indicate a significant difference based on Tukey’s HSD test (p ≤ 0.05).
Huibers et al. [47] demonstrated that resistance due to the silencing of Pmr4 is associated with salicylic acid (SA) accumulation rather than with the callose deposition absence. Salicylic acid signaling plays a key role protecting against biotrophic pathogens through the establishment of a hypersensitive response (HR). Saiz-Fernandez et al. [48] revealed the increment of SA levels in
is associated with salicylic acid (SA) accumulation rather than with the callose deposition absence. Salicylic acid signaling plays a key role protecting against biotrophic pathogens through the establishment of a hypersensitive response (HR). Saiz-Fernandez et al. [49] revealed the increment of SA levels inP. cinnamomi
inoculated stems, indicating thatP. cinnamomi
activates a defense response similar to that triggered by biotrophic pathogens. Inoculation with both virulent and hypovirulent strains ofC. parasitica led to SA accumulation in European chestnut plantlets that were grown in vitro [49]. Transcriptome analyses carried out in both
led to SA accumulation in European chestnut plantlets that were grown in vitro [50]. Transcriptome analyses carried out in bothC. dentata
andC. mollissima highlighted activation of salicylic-acid-related genes in canker tissues [36].
highlighted activation of salicylic-acid-related genes in canker tissues [37]. In chestnut trees, callose deposition aroundP. cinnamomi hyphae was detected early in the infection process; however, it does not seem to play a key role in the associated interactions since the pathogen can reach the vascular cylinder in both susceptible and resistant plant genotypes [50]. This result was validated by transcriptomes analyses of
hyphae was detected early in the infection process; however, it does not seem to play a key role in the associated interactions since the pathogen can reach the vascular cylinder in both susceptible and resistant plant genotypes [51]. This result was validated by transcriptomes analyses ofC. sativa
andC. crenata
, in which no overexpression ofCallose synthases
afterP. cinnamomi infection was observed [37].
infection was observed [38]. Based on our results and the literature, we can hypothesize that callose accumulation due to thepmr4
upregulation in inoculatedC. sativa
lines may not be responsible for controllingP. cinnamomi
colonization. We suggest that the upregulation ofpmr4
could lead to a negative regulation of the SA pathway that in turn provokes the susceptibility ofC. sativa
to bothP. cinnamomi
andC. parasitica
. A clear link with SA pathway has emerged even with the other chestnut gene candidatedmr6
(downy mildew resistance 6). The mutation ofArabidopsis dmr6
gene, associated with salicylic acid (SA) homeostasis [31], results in the generation of plants that are resistant to bacteria and oomycetes.Dmr6
is involved in the conversion of salicylic acid (SA) to 2,3-dihydroxybenzoic acid (2,3-DHBA) and negatively regulates the expression of defense genes (PR-1, PR-2, and PR-5) [30]. The expression trend of theDmr6
gene in response toP. cinnamomi
infection turned out to be similar to the profile ofpmr4
. Indeed,dmr6
was strongly expressed at early infection phases ofP. cinnamomi
inC. sativa
; inC. crenata
no significant upregulation was detected (). No upregulation ofdmr6
in response toC. parasitica
was highlighted in both plant species (Figure 6). We can thus hypothesize that
7). We can thus hypothesize thatdmr6
upregulation observed inC. sativa
could negatively regulate defense gene expression, leading to susceptibility toP. cinnamomi
.Plants produce a variety of hydrolytic defense enzymes against pathogens, including chitinases, proteases, and also glucanases [51]. The genes coding for several pathogenesis-related (PR) proteins,
Plants produce a variety of hydrolytic defense enzymes against pathogens, including chitinases, proteases, and also glucanases [52]. The genes coding for several pathogenesis-related (PR) proteins,Acidic 26 kDa endochitinase
gene (chi3
) andGlucan endo-1,3-beta-glucosidase B
gene (gluB), were selected in our analysis because they are considered as responsive to SA-dependent signaling [52][53].
), were selected in our analysis because they are considered as responsive to SA-dependent signaling [53,54].Chi3
andgluB are enzymes that cause the lysis of hyphae of various pathogens, resulting in growth inhibition [54][55][56].
are enzymes that cause the lysis of hyphae of various pathogens, resulting in growth inhibition [55,56,57]. In bothC. sativa
andC. crenata
plants inoculated withC. parasitica
,chi3
andgluB
were significantly upregulated. The transcription ofchi3
was higher inC. crenata
than inC. sativa
, presumably as a consequence of the improved defense mechanism againstC. parasitica. Our results are in agreement with Shain et al. [57], who demonstrated the involvement of
. Our results are in agreement with Shain et al. [58], who demonstrated the involvement ofb-1,3-glucanase
andchitinase
in chestnut species affected byC. parasitica
. Studies on the role ofchitinase
in blight infection mostly involvedC. sativa as a model system [49][58]. In both
as a model system [50,59]. In bothC. dentata
andC. mollissima
, transcripts of several compounds expressingchitinase accumulated more in canker tissues than healthy stem tissues [36]. In order to obtain chestnut plants with potentially increased resistance/tolerance to chestnut blight, the endogenous Ch3gene encoding a chitinase-like protein was over-expressed in the European chestnut through Agrobacterium-mediated transformation [59].
accumulated more in canker tissues than healthy stem tissues [37]. In order to obtain chestnut plants with potentially increased resistance/tolerance to chestnut blight, the endogenous Ch3gene encoding a chitinase-like protein was over-expressed in the European chestnut through Agrobacterium-mediated transformation [60].The emergent CRISPR/Cas9 technology is expected to play a key role in future crop breeding as it allows highly efficient gene editing and generates genetic changes indistinguishable from those arising spontaneously in nature or through conventional breeding [60]. Several examples of edited plants resistant to fungal pathogens have been described [61][62]. For example, genome editing was successfully applied to knock out
The emergent CRISPR/Cas9 technology is expected to play a key role in future crop breeding as it allows highly efficient gene editing and generates genetic changes indistinguishable from those arising spontaneously in nature or through conventional breeding [61]. Several examples of edited plants resistant to fungal pathogens have been described [62,63]. For example, genome editing was successfully applied to knock outmlo S-genes, leading to Powdery mildew (PM) resistance [43][63][64][65].
S-genes, leading to Powdery mildew (PM) resistance [44,64,65,66].Pmr4
anddmr6 loss-of-function through CRISPR/Cas reduced the susceptibility to PM in tomato plants [28][66]. In our laboratory, we are setting up a CRISPR/Cas9 transformation protocol in
loss-of-function through CRISPR/Cas reduced the susceptibility to PM in tomato plants [28,67]. In our laboratory, we are setting up a CRISPR/Cas9 transformation protocol inCastanea sativa
. Our future goal will be to perform the functional characterization using the CRISPR/Cas9 approach of the two candidate genes (dmr6
andpmr4
), while checking if the two genes may also play a role in the interaction betweenC. sativa
and the emergent nut rot and canker agentGnomoniopsis castaneae [67].
[68].