Kidney transplantation is the preferred treatment for end-stage kidney disease (ESKD). Compared to maintenance dialysis, kidney transplantation results in improved patient survival and quality of life. Kidneys from living donors perform best; however, many patients with ESKD depend on kidneys from deceased donors. After procurement, donor kidneys are placed in a cold-storage solution until a suitable recipient is located. Sadly, prolonged cold storage times are associated with inferior transplant outcomes; therefore, in most situations when considering donor kidneys, long cold-storage times are avoided. The identification of novel mechanisms of cold-storage-related renal damage will lead to the development of new therapeutic strategies for preserving donor kidneys; to date, these mechanisms remain poorly understood. In this review, we discuss the importance of mitochondrial function, protein homeostasis, and renal recovery during stress from cold storage plus transplantation. Additionally, we discuss novel targets for therapeutic intervention.
- cold storage
- transplantation
- proteasome function
- mitochondrial function
- therapetutics
1. Introduction
Acute kidney injury affects approximately 13–18% of hospitalized patients and has been shown to be associated with increased mortality. [1]. Additionally, acute kidney injury has also been linked to the development of chronic kidney disease and end-stage kidney disease (ESKD). ESKD affects 630,000 Americans and is the ninth-leading cause of death in the US (NIDDK, 2014). Kidney transplantation is the preferred treatment to increase longevity and quality of life for people with ESKD, but due to a shortage of transplantable kidneys, 7 of 10 ESKD patients will remain on dialysis (and many will die) while waiting for a kidney (95,268 waiting-list candidates vs. 19,848 transplants in 2017; http://www.unos.org).
Advances in tissue-type matching and immunosuppressive protocols have greatly reduced the incidence of acute transplant rejection and short-term graft dysfunction; however, optimizing long-term graft function continues to be a challenge, especially with kidneys from deceased donors. Kidneys from living donors have better long-term graft outcomes than those from deceased donors. One of the key variables is cold storage (CS) [2][3][4], which is the universal method for preserving kidneys from donors that allows time for identification of potential recipients, transportation of kidneys, tissue typing, and cross-matching. Kidneys from living donors generally are exposed to only a few hours of CS, while those from deceased donors undergo long hours of CS during transport, tissue typing, and cross-matching. Despite the increasing use of hypothermic machine perfusion (HMP) for the preservation of deceased donor kidneys, most transplant centers still rely on static CS techniques. Both CS techniques (HMP or static) lower the metabolic rate, allowing the organ to be stored until a recipient is located [5][6]. Acceptable CS times vary between transplant centers, ranging from 24 to 72 h. Unfortunately, each additional hour of CS increases the risk of graft failure [2][3][4] via poorly described mechanisms. Tragically, approximately 20% of donor kidneys that are retrieved each year are discarded or not transplanted (http://www.unos.org), partly due to prolonged CS [7][8]. A better understanding of injury-related pathways secondary to CS, and the identification of novel therapies aimed to mitigate damage would likely lead to improved deceased donor transplant rates and patient outcomes.
The kidney is a composite organ made up of many blood-filtering units called nephrons. Each nephron consists of anatomically and functionally discrete segments known as a renal corpuscle, proximal tubule, loop of Henle, distal tubule, and collecting duct system [9]. Each segment of the nephron is composed of multiple cell types of both epithelial and mesenchymal origin; all of which participate in various functions such as removing nitrogen waste and other waste products, controlling blood electrolytes and acid-base balance, and secreting hormones that regulate blood composition and blood pressure [9][10][11][12][13]. Like most organs, renal tissues consume a high level of oxygen [14][15][16], and have quality-control mechanisms that help fold nascent polypeptides, clear unfolded or misfolded proteins, respond to protein aggregates, and dispose of potentially toxic molecules. The balance of energy and the integrity of the proteome is extremely important for renal cell viability during stressful conditions, especially during renal cold storage plus transplantation (CS/Tx). One could reasonably expect to see damage within various compartments of renal tissue when the kidney undergoes a series of stressors, namely CS and blood reperfusion events during transplantation. Recovery from CS/Tx-induced stress is possible when the repair process remains in balance and overcomes the rate of CS/Tx-mediated injury. Unfortunately, very little is known about the mechanisms of CS/Tx-mediated damage that leads to renal dysfunction after CS/Tx. Here, we will discuss some of the literature that describes the triggers of damage during renal transplantation. Specifically, we will discuss the importance of mitochondrial and proteasomal function to maintain mitochondrial protein quality during renal CS/Tx and potential strategies to prevent organ damage and improve transplant outcomes.
2. The Proteasome
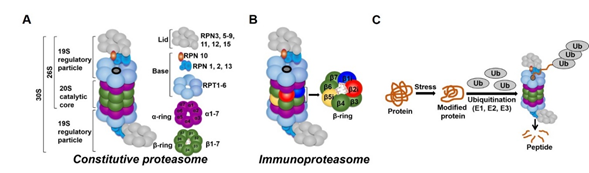
In all tissues, proteasomes are crucial for degrading modified, misfolded, and damaged proteins. The constitutive proteasome, selectively degrades ubiquitinated proteins (via the concerted actions of ubiquitinating enzymes) to small peptides (Figure 1A and [171][182][193][204]). The constitutive proteasome, a multi-subunit holoenzyme of ~2.5 MDa, is made up of two distinct sub-domains, namely, a 20S catalytic core particle and 1 or 2 19S regulatory particle(s) (Figure 1A). The 20S core particle is a barrel-shaped complex, composed of stacks of 2 β-rings (each ring made up of β1-7 subunits) in the center and 2 α-rings (each ring made up of α 1-7 subunits) on each end (Figure 1A). The α-rings (subunits) appear to have a regulatory function, allowing only unfolded substrates to enter into the 20S catalytic core. The β-ring of the 20S proteasome has 3 to 7 active sites (β-catalytic subunits) (Figure 1A) that hydrolyze peptide bonds in a chymotrypsin-like (β5 subunit), trypsin-like (β2 subunit), or caspase-like (β1 subunit) fashion (Figure 1C and [215]). The 19S regulatory particle recognizes and unfolds the ubiquitinated substrates before allowing the substrate to enter the 20S pore [226]. Functionally, the 19S particle is divided into a base and a lid. The base consists of an ATPase ring, made up of 6 AAA-ATPase subunits (Rpt1-6), and 3 non-ATPase subunits (Rpn1, 2, and 13) (Figure 1A). The ATPase subunits consume ATP to unfold the substrate and help translocate it to the pore of the 20S catalytic core. The lid, which is linked to the base by the Rpn10 subunit (Figure 1A), assists in the efficient degradation of the ubiquitinated substrates. The immunoproteasome is a proteasome variant that is normally found in immune cell compartments [237]. However, in response to inflammation, catalytic subunits (β1, β2, and β5) of the constitutive proteasome are exchanged for immunoproteasome subunits (β1i, β2i, and β5i) in most non-immune cells (Figure 1B and [237][248]).
Figure 1. Ubiquitin-proteasome system (UPS). (A) The constitutive proteasome (26S or 36S) is a barrel-shaped organelle that is made up of 20S catalytic core and one (26S) or two (30S) 19S particle(s). The 20S catalytic core is made up of stacks of two β rings (β1–β7) and two α rings (α1–α7). The 19S regulatory particle is made up of a lid and a base with multiple subunits. (B) The immunoproteasome is a proteasome variant that is normally found in immune cell compartments. However, in response to inflammation, constitutive proteasome subunits (β1, β2, and β5) are exchanged for the immunoproteasome subunits (β1i, β2i, and β5i) in most non-immune cells in the body. (C) Damaged or modified proteins are ubiquitinated (with the concerted action of ubiquitinating enzymes), which is then recognized by the constitutive proteasome for degradation. The constitutive proteasome selectively degrades ubiquitinated proteins to small peptides (A); it has 3 to 7 protease active sites (β-catalytic subunits) that hydrolyze peptide bonds in a chymotrypsin (β5 subunit)-, trypsin (β2 subunit)-, or caspase (β1 subunit)-like fashion. Following protein degradation, the peptides are released and recycled.
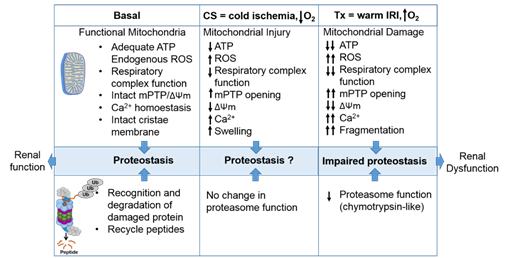
The proteasome maintains functional protein homeostasis, also known as proteostasis, by monitoring misfolded and damaged proteins; however, this is a challenge in the context of renal CS/Tx, especially in kidneys that have undergone prolonged CS (Figure 2). Given that prolonged CS followed by warm IRI produces ROS [259][2610], and that ROS modulate the constitutive proteasome function [2711][2812][2913], we can postulate that CS/Tx-mediated ROS could trigger denaturation of intracellular proteins and modulation of the constitutive proteasome function. Indeed, a recent report demonstrated that the chymotrypsin-like activity of the proteasome was compromised after renal CS/Tx [3014], and that this correlated with severe renal dysfunction [3115][3216]. A study performed by using pharmacological inhibition of chymotrypsin-like activity of the proteasome during warm IRI showed aggravated renal damage [3317]. Genetic or pharmacologic modulation of proteasome function (chymotrypsin-like) inhibition, achieved by siRNA or bortezomib treatment, respectively, in rat proximal tubular cells showed increase of ROS production [3418]. At this point, the mechanisms of proteasome dysfunction during CS/Tx are not known. One of the possible mechanisms could be CS/Tx-mediated post-translational modification of the proteasome subunits because this mechanism has been described to modulate proteasome function and assembly in various experimental models [3519][3620].
Figure 2. The intricate relationship between the mitochondria and proteasome. Schematic summary depicting mitochondrial and proteasomal changes during cold storage (CS) and transplantation. During CS, mitochondrial respiration function, ATP level, and mitochondrial membrane potential (ΔΨm) decreases, whereas ROS and calcium levels increase leading to an increase of mitochondrial permeability transition pore (mPTP) opening. This leads to increased swelling and decreased function of mitochondria. These changes are further exacerbated and sustained following transplantation leading to mitochondrial fragmentation and bioenergetic crisis. Proteasome function remains unchanged during renal CS, whereas the chymotrypsin-like proteasome function is decreased following transplantation. This leads to alteration of mitochondrial protein homeostasis and acute tubular necrosis after transplantation and significantly decreases renal function.
It is well-accepted that CS/Tx produces inflammation and releases inflammatory cytokines [3721]. Immunoproteasomes are induced and activated in response to the inflammatory cytokines, and help in processing donor-derived antigen effectively. Unlike the constitutive proteasome, the immunoproteasome is resistant to oxidative stress and can function in an ATP-independent manner [3822][3923]. For example, interferon-induced ROS activate the immunoproteasome [3923]. There are a handful of transplant studies showing a negative correlation of immunoproteasome activity with organ function. In this context, a recent report by Li et al. indicates that pharmacological inhibition of the immunoproteasome with ONX 0914 (a reversible β5i inhibitor) reduces donor-specific antibody production in a rat model of renal CS/Tx [4024]. Thus, it is worth investigating whether ONX 0914 should be included in the CS solution. This is particularly exciting because some studies indicate that ONX 0914 protects against cardiac and neuronal IRI [4125][4226]. However, one caveat of the Li et al. study is that the authors used a very short CS time (~35 min) that is likely not clinically relevant. Future studies with longer CS times are needed to verify whether blunting the immunoproteasome with ONX 0914 during CS confers protection after transplantation. The specific mechanisms underlying compromised proteasome function and exacerbated immunoproteasome activity during renal CS/Tx are not understood and should be addressed.
Although the proteasome manages protein turnover, aberrations in the expression and function of the constitutive proteasome and immunoproteasome are implicated in the pathogenesis of several human diseases, including cancer, autoimmune disorders, and inflammatory diseases [237][248][4327][4428][4529][4630][142,143,158–161]. In the context of CS/Tx, it is clear that the initial damage occurs within the epithelial or vascular compartments within the kidneys (during cold ischemia) that eventually triggers an immune response following blood reperfusion (warm IRI) after CS/Tx. It is expected that prolonged CS followed by transplantation disrupts protein homeostasis, which overwhelms the immune response, and this could directly impact long-term graft and patient outcomes.
References
- Sawhney, S.; Mitchell, M.; Marks, A.; Fluck, N.; Black, C. Long-term prognosis after acute kidney injury (AKI): What is the role of baseline kidney function and recovery? A systematic review. BMJ Open 2015, 5.Ciechanover, A.; Schwartz, A.L. The ubiquitin-dependent proteolytic pathway: Specificity of recognition of the proteolytic substrates. Revis. Sobre Boil. Cel. RBC 1989, 20, 217–234.
- Debout, A.; Foucher, Y.; Trébern-Launay, K.; Legendre, C.; Kreis, H.; Mourad, G.; Garrigue, V.; Morelon, E.; Buron, F.; Rostaing, L.; et al. Each additional hour of cold ischemia time significantly increases the risk of graft failure and mortality following renal transplantation. Kidney Int. 2015, 87, 343–349.Glickman, M.; Ciechanover, A. The Ubiquitin-Proteasome Proteolytic Pathway: Destruction for the Sake of Construction. Physiol. Rev. 2002, 82, 373–428.
- Hosgood, S.A.; Bagul, A.; Nicholson, M.L. Minimising cold ischaemic injury in an experimental model of kidney transplantation. Eur. J. Clin. Investig. 2010, 41, 233–240.Zwickl, P.; Voges, D.; Baumeister, W. The proteasome: A macromolecular assembly designed for controlled proteolysis. Philos. Trans. R. Soc. B: Boil. Sci. 1999, 354, 1501–1511.
- Salahudeen, A.K.; May, W. Reduction in Cold Ischemia Time of Renal Allografts in the United States Over the Last Decade. Transplant. Proc. 2008, 40, 1285–1289.Hershko, A.; Ciechanover, A. The Ubiquitin Pathway for the Degradation of Intracellular Proteins. Prog. Nucl. Acid Res. Mol. Biol. 1986, 33, 19–56.
- Kay, M.D.; Hosgood, S.A.; Bagul, A.; Nicholson, M. Comparison of preservation solutions in an experimental model of organ cooling in kidney transplantation. BJS 2009, 96, 1215–1221.Jung, T.; Grune, T. Structure of the Proteasome. Prog. Mol. Biol. Transl. Sci. 2012, 109, 1–39.
- O’Callaghan, J.; Morgan, R.D.; Knight, S.; Morris, P.J. Systematic review and meta-analysis of hypothermic machine perfusion versus static cold storage of kidney allografts on transplant outcomes. BJS 2013, 100, 991–1001.Lander, G.C.; Estrin, E.; Matyskiela, M.E.; Bashore, C.; Nogales, E.; Martin, A. Complete subunit architecture of the proteasome regulatory particle. Nature 2012, 482, 186–191.
- Mohan, S.; Foley, K.F.; Chiles, M.C.; Dube, G.K.; Patzer, R.E.; Pastan, S.O.; Crew, R.J.; Cohen, D.J.; Ratner, L.E. The weekend effect alters the procurement and discard rates of deceased donor kidneys in the United States. Kidney Int. 2016, 90, 157–163.Basler, M.; Mundt, S.; Bitzer, A.; Schmidt, C.; Groettrup, M. The immunoproteasome: A novel drug target for autoimmune diseases. Clin. Exp. Rheumatol. 2015, 33.
- Stewart, D.; Garcia, V.C.; Rosendale, J.D.; Klassen, D.; Carrico, B.J. Diagnosing the Decades-Long Rise in the Deceased Donor Kidney Discard Rate in the United States. Transplantation 2017, 101, 575–587.Kaur, G.; Batra, S. Emerging role of immunoproteasomes in pathophysiology. Immunol. Cell Boil. 2016, 94, 812–820.
- Oxburgh, L. Kidney Nephron Determination. Ann. Rew. Cell Dev. Biol. 2018, 34, 427–450.Salahudeen, A.K. Free radicals in kidney disease and transplantation. Saudi J. Kidney Dis. Transplant. 1999, 10, 137–143.
- M’Donnell, R. Observations on the Anatomy and Physiology of the Kidney. Glas. Med. J. 2009, 101, 285–295.Johnson, K.J.; Weinberg, J.M. Postischemic renal injury due to oxygen radicals. Curr. Opin. Nephrol. Hypertens. 1993, 2, 625–635.
- Kriz, W.; Sanchez-Lozada, L.G.; Bulger, R.E.; Burg, M.B.; Goncharevskaya, O.A.; Imai, M.; Kaissling, B.; Maunsbach, A.B.; Moffat, D.B.; Morgan, T.O.; et al. A standard nomenclature for structures of the kidney. Pflüger, Archiv für die Gesammte Physiologie des Menschen und der Thiere 1988, 411, 113–120.Andersson, M.; Sjöstrand, J.; Karlsson, J.-O. Differential Inhibition of Three Peptidase Activities of the Proteasome in Human Lens Epithelium by Heat and Oxidation. Exp. Eye Res. 1999, 69, 129–138.
- Kumaran, G.K.; Hanukoglu, I. Identification and classification of epithelial cells in nephron segments by actin cytoskeleton patterns. FEBS J. 2019, 287, 1176–1194.Reinheckel, T.; Sitte, N.; Ullrich, O.; Kuckelkorn, U.; Davies, K.E.; Grune, T. Comparative resistance of the 20S and 26S proteasome to oxidative stress. Biochem. J. 1998, 335, 637–642.
- Wallace, M.A. Anatomy, and physiology of the kidney. AORN J. 1998, 68, 799–820.Reinheckel, T.; Ullrich, O.; Sitte, N.; Grune, T. Differential Impairment of 20S and 26S Proteasome Activities in Human Hematopoietic K562 Cells during Oxidative Stress. Arch. Biochem. Biophys. 2000, 377, 65–68.
- Hansell, P.; Welch, W.J.; Blantz, R.C.; Palm, F. Determinants of kidney oxygen consumption and their relationship to tissue oxygen tension in diabetes and hypertension. Clin. Exp. Pharmacol. Physiol. 2013, 40, 123–137.Lo, S.; MacMillan-Crow, L.A.; Parajuli, N. Renal cold storage followed by transplantation impairs proteasome function and mitochondrial protein homeostasis. Am. J. Physiol. 2019, 316, 42–53.
- Persson, P.B.; Ehmke, H.; Kirchheim, H.R.; Janssen, B.; E Baumann, J.; Just, A.; Nafz, B. Autoregulation and non-homeostatic behaviour of renal blood flow in conscious dogs. J. Physiol. 1993, 462, 261–273.Parajuli, N.; Shrum, S.; Tobacyk, J.; Harb, A.; Arthur, J.; MacMillan-Crow, L.A. Renal cold storage followed by transplantation impairs expression of key mitochondrial fission and fusion proteins. PLoS ONE 2017, 12, e0185542.
- Levy, M.N. Effect of variations of blood flow on renal oxygen extraction. Am. J. Physiol. Content 1960, 199, 13–18.Shrum, S.; MacMillan-Crow, L.A.; Parajuli, N. Cold Storage Exacerbates Renal and Mitochondrial Dysfunction Following Transplantation. J. Kidney 2016, 2.
- Ciechanover, A.; Schwartz, A.L. The ubiquitin-dependent proteolytic pathway: Specificity of recognition of the proteolytic substrates. Revis. Sobre Boil. Cel. RBC 1989, 20, 217–234.Huber, J.M.; Tagwerker, A.; Heininger, D.; Mayer, G.; Rosenkranz, A.R.; Eller, K. The proteasome inhibitor Bortezomib aggravates renal ischemia-reperfusion injury. Am. J. Physiol. 2009, 297, 451–460.
- Glickman, M.; Ciechanover, A. The Ubiquitin-Proteasome Proteolytic Pathway: Destruction for the Sake of Construction. Physiol. Rev. 2002, 82, 373–428.Parajuli, N. A Cycle of Altered Proteasome and Reactive Oxygen Species Production in Renal Proximal Tubular Cells. Toxicol. Forensic Med. Open J. 2019, 4, 13–17.
- Zwickl, P.; Voges, D.; Baumeister, W. The proteasome: A macromolecular assembly designed for controlled proteolysis. Philos. Trans. R. Soc. B: Boil. Sci. 1999, 354, 1501–1511.Scruggs, S.B.; Zong, N.C.; Wang, D.; Stefani, E.; Ping, P. Post-translational modification of cardiac proteasomes: Functional delineation enabled by proteomics. Am. J. Physiol. Circ. Physiol. 2012, 303, 9–18.
- Hershko, A.; Ciechanover, A. The Ubiquitin Pathway for the Degradation of Intracellular Proteins. Prog. Nucl. Acid Res. Mol. Biol. 1986, 33, 19–56.Kors, S.; Geijtenbeek, K.; Reits, E.; Schipper-Krom, S. Regulation of Proteasome Activity by (Post-)transcriptional Mechanisms. Front. Mol. Biosci. 2019, 6, 48.
- Jung, T.; Grune, T. Structure of the Proteasome. Prog. Mol. Biol. Transl. Sci. 2012, 109, 1–39.Cherukuri, A.; Mehta, R.; Sood, P.; Hariharan, S. Early allograft inflammation and scarring associate with graft dysfunction and poor outcomes in renal transplant recipients with delayed graft function: A prospective single center cohort study. Transpl. Int. 2018, 31, 1369–1379.
- Lander, G.C.; Estrin, E.; Matyskiela, M.E.; Bashore, C.; Nogales, E.; Martin, A. Complete subunit architecture of the proteasome regulatory particle. Nature 2012, 482, 186–191.Aiken, C.T.; Kaake, R.M.; Wang, X.; Huang, L. Oxidative stress-mediated regulation of proteasome complexes. Mol. Cell. Proteom. MCP 2011, 10.
- Basler, M.; Mundt, S.; Bitzer, A.; Schmidt, C.; Groettrup, M. The immunoproteasome: A novel drug target for autoimmune diseases. Clin. Exp. Rheumatol. 2015, 33.Seifert, U.; Bialy, L.; Ebstein, F.; Bech-Otschir, D.; Voigt, A.; Schröter, F.; Prozorovski, T.; Lange, N.; Steffen, J.; Rieger, M.; et al. Immunoproteasomes Preserve Protein Homeostasis upon Interferon-Induced Oxidative Stress. Cell 2010, 142, 613–624.
- Kaur, G.; Batra, S. Emerging role of immunoproteasomes in pathophysiology. Immunol. Cell Boil. 2016, 94, 812–820.Li, J.; Basler, M.; Alvarez, G.; Brunner, T.; Kirk, C.J.; Groettrup, M. Immunoproteasome inhibition prevents chronic antibody-mediated allograft rejection in renal transplantation. Kidney Int. 2018, 93, 670–680.
- Reinheckel, T.; Ullrich, O.; Sitte, N.; Grune, T. Differential Impairment of 20S and 26S Proteasome Activities in Human Hematopoietic K562 Cells during Oxidative Stress. Arch. Biochem. Biophys. 2000, 377, 65–68.Chen, X.; Zhang, X.; Wang, Y.; Lei, H.; Su, H.; Zeng, J.; Pei, Z.; Huang, R. Inhibition of immunoproteasome reduces infarction volume and attenuates inflammatory reaction in a rat model of ischemic stroke. Cell Death Dis. 2015, 6, 1626.
- Lo, S.; MacMillan-Crow, L.A.; Parajuli, N. Renal cold storage followed by transplantation impairs proteasome function and mitochondrial protein homeostasis. Am. J. Physiol. 2019, 316, 42–53.Karreci, E.S.; Fan, H.; Uehara, M.; Mihali, A.B.; Singh, P.K.; Kurdi, A.T.; Solhjou, Z.; Riella, L.V.; Ghobrial, I.; Laragione, T.; et al. Brief treatment with a highly selective immunoproteasome inhibitor promotes long-term cardiac allograft acceptance in mice. Proc. Nat. Acad. Sci. USA 2016, 113, 8425–8432.
- Salahudeen, A.K. Free radicals in kidney disease and transplantation. Saudi J. Kidney Dis. Transplant. 1999, 10, 137–143.Bellavista, E.; Andreoli, F.; Parenti, M.D.; Martucci, M.; Santoro, A.; Salvioli, S.; Capri, M.; Baruzzi, A.; Del Rio, A.; Franceschi, C.; et al. Immunoproteasome in Cancer and Neuropathologies: A New Therapeutic Target? Curr. Pharm. Des. 2013, 19, 702–718.
- Johnson, K.J.; Weinberg, J.M. Postischemic renal injury due to oxygen radicals. Curr. Opin. Nephrol. Hypertens. 1993, 2, 625–635.Miller, Z.; Ao, L.; Kim, K.B.; Lee, W. Inhibitors of the immunoproteasome: Current status and future directions. Curr. Pharm. Des. 2013, 19, 4140–4151.
- Andersson, M.; Sjöstrand, J.; Karlsson, J.-O. Differential Inhibition of Three Peptidase Activities of the Proteasome in Human Lens Epithelium by Heat and Oxidation. Exp. Eye Res. 1999, 69, 129–138.Paul, S. Dysfunction of the ubiquitin-proteasome system in multiple disease conditions: Therapeutic approaches. BioEssays 2008, 30, 1172–1184.
- Reinheckel, T.; Sitte, N.; Ullrich, O.; Kuckelkorn, U.; Davies, K.E.; Grune, T. Comparative resistance of the 20S and 26S proteasome to oxidative stress. Biochem. J. 1998, 335, 637–642.Schmidt, M.; Finley, D. Regulation of proteasome activity in health and disease. Biochim. et Biophys. Acta Bioenerg. 2013, 1843, 13–25.
- Parajuli, N.; Shrum, S.; Tobacyk, J.; Harb, A.; Arthur, J.; MacMillan-Crow, L.A. Renal cold storage followed by transplantation impairs expression of key mitochondrial fission and fusion proteins. PLoS ONE 2017, 12, e0185542.
- Shrum, S.; MacMillan-Crow, L.A.; Parajuli, N. Cold Storage Exacerbates Renal and Mitochondrial Dysfunction Following Transplantation. J. Kidney 2016, 2.
- Huber, J.M.; Tagwerker, A.; Heininger, D.; Mayer, G.; Rosenkranz, A.R.; Eller, K. The proteasome inhibitor Bortezomib aggravates renal ischemia-reperfusion injury. Am. J. Physiol. 2009, 297, 451–460.
- Parajuli, N. A Cycle of Altered Proteasome and Reactive Oxygen Species Production in Renal Proximal Tubular Cells. Toxicol. Forensic Med. Open J. 2019, 4, 13–17.
- Scruggs, S.B.; Zong, N.C.; Wang, D.; Stefani, E.; Ping, P. Post-translational modification of cardiac proteasomes: Functional delineation enabled by proteomics. Am. J. Physiol. Circ. Physiol. 2012, 303, 9–18.
- Kors, S.; Geijtenbeek, K.; Reits, E.; Schipper-Krom, S. Regulation of Proteasome Activity by (Post-)transcriptional Mechanisms. Front. Mol. Biosci. 2019, 6, 48.
- Cherukuri, A.; Mehta, R.; Sood, P.; Hariharan, S. Early allograft inflammation and scarring associate with graft dysfunction and poor outcomes in renal transplant recipients with delayed graft function: A prospective single center cohort study. Transpl. Int. 2018, 31, 1369–1379.
- Aiken, C.T.; Kaake, R.M.; Wang, X.; Huang, L. Oxidative stress-mediated regulation of proteasome complexes. Mol. Cell. Proteom. MCP 2011, 10.
- Seifert, U.; Bialy, L.; Ebstein, F.; Bech-Otschir, D.; Voigt, A.; Schröter, F.; Prozorovski, T.; Lange, N.; Steffen, J.; Rieger, M.; et al. Immunoproteasomes Preserve Protein Homeostasis upon Interferon-Induced Oxidative Stress. Cell 2010, 142, 613–624.
- Li, J.; Basler, M.; Alvarez, G.; Brunner, T.; Kirk, C.J.; Groettrup, M. Immunoproteasome inhibition prevents chronic antibody-mediated allograft rejection in renal transplantation. Kidney Int. 2018, 93, 670–680.
- Chen, X.; Zhang, X.; Wang, Y.; Lei, H.; Su, H.; Zeng, J.; Pei, Z.; Huang, R. Inhibition of immunoproteasome reduces infarction volume and attenuates inflammatory reaction in a rat model of ischemic stroke. Cell Death Dis. 2015, 6, 1626.
- Karreci, E.S.; Fan, H.; Uehara, M.; Mihali, A.B.; Singh, P.K.; Kurdi, A.T.; Solhjou, Z.; Riella, L.V.; Ghobrial, I.; Laragione, T.; et al. Brief treatment with a highly selective immunoproteasome inhibitor promotes long-term cardiac allograft acceptance in mice. Proc. Nat. Acad. Sci. USA 2016, 113, 8425–8432.
- Bellavista, E.; Andreoli, F.; Parenti, M.D.; Martucci, M.; Santoro, A.; Salvioli, S.; Capri, M.; Baruzzi, A.; Del Rio, A.; Franceschi, C.; et al. Immunoproteasome in Cancer and Neuropathologies: A New Therapeutic Target? Curr. Pharm. Des. 2013, 19, 702–718.
- Miller, Z.; Ao, L.; Kim, K.B.; Lee, W. Inhibitors of the immunoproteasome: Current status and future directions. Curr. Pharm. Des. 2013, 19, 4140–4151.
- Paul, S. Dysfunction of the ubiquitin-proteasome system in multiple disease conditions: Therapeutic approaches. BioEssays 2008, 30, 1172–1184.
- Schmidt, M.; Finley, D. Regulation of proteasome activity in health and disease. Biochim. et Biophys. Acta Bioenerg. 2013, 1843, 13–25.