Biofilms are associations of microorganisms embedded in a self-produced extracellular matrix. They create particular environments that confer bacterial tolerance and resistance to antibiotics by different mechanisms that depend upon factors such as biofilm composition, architecture, the stage of biofilm development, and growth conditions. The biofilm structure hinders the penetration of antibiotics and may prevent the accumulation of bactericidal concentrations throughout the entire biofilm. In addition, biofilm can induce a variety of physiological states involving different metabolism (aerobic, microaerobic, and fermentative) and growth rates (fast and slow growth, dormant cells, and persister cells). Thus, some bacteria are less vulnerable to antibiotics as a consequence of the inactivity of antibiotic targets, a phenomenon called “drug indifference”. Also, cells in biofilms enhance efflux-pump production that excretes antibiotics. Moreover, the biofilm environment enhance interbacterial communication, horizontal gene transfer and spontaneous mutations that ultimately increase resistance to antibiotics. Thus, the tolerance and resistance to antibiotics conferred by biofilms is multifactorial.
- biofilms
- antibiotic resistance,antibiotic tolerance
1. Basis of Biofilm-Mediated Antibiotic Survival
The recalcitrant nature of biofilms to antibiotics depends mostly on (i) the developmental stage of the biofilm, (ii) the extracellular matrix (ECM) composition, and (iii) the biofilm architecture.
1.1. Biogenesis of Biofilms
Biofilm formation is a dynamic process that takes place in a series of sequential steps. It is initiated by the interaction of the bacteria with a surface. Exposure of planktonic cells to stress, which may be provoked by antibiotics, starvation, or other adverse environmental conditions, can initiate biofilm formation by activating gene expression[1]. Additionally, molecules involved in cell-to-cell communication accumulate at high cell density. These molecules generally referred to as autoinducers, can activate and regulate the process[2] and allow for a coordinated response of the population members, which is known as quorum sensing (QS). The first step of biofilm formation consists of the adhesion of the bacteria to the substratum. This process is often mediated by long, proteinaceous, filamentous fibers that protrude from the bacterial cell surface, such as flagella, fimbriae, or pili. After initial interaction is established, shorter cell surface-exposed structures interact with the substratum, thereby increasing the contact between bacteria and the substratum[3]. Strains of Escherichia coli and Salmonella produce curli fimbrae that mediate both cell-to-substratum and cell-to-cell interactions[4]. Other proteins such as Bap-family proteins in Staphylococcus epidermidis[5], CdrA in Pseudomonas aeruginosa[6] or NHBA in Neisseria meningitidis[7] are proteins that interact with ECM components and the bacterial cell surface thereby strengthening the matrix. Autotransporters are proteins secreted through the Type V secretion system in many Gram-negative bacteria and often have demonstrated roles in interbacterial interactions[8]. Then, the bacteria secrete ECM components and proliferate to form a microcolony. ECM serves as a glue element that helps to stabilize interbacterial interactions. Bacteria within the microcolony communicate and organize spatially. Type IV pili act at the junction between cells to form microcolonies and can also contribute to the reorganization of bacteria within the biofilm[9]. Cell-to-cell communication, including QS [2] and also cell-contact-dependent communication systems[10], seem to be relevant for this process. At this stage, the expression of genes for the formation of the ECM, and biofilms become less vulnerable to antibiotics than earlier biofilm stages[11].
1.2. Composition of the ECM
The ECM consists of a conglomerate of different substances that together provide structural integrity to the biofilm. In general, the ECM can be composed of water, polysaccharides, proteins, lipids, surfactants, glycolipids, extracellular DNA (eDNA), extracellular RNA, membrane vesicles, and ions such as Ca2+. In many bacteria, extracellular polysaccharides and eDNA are prominent components of the ECM[12].
eDNA is constituted of chromosomal DNA that is released into the extracellular milieu through cell lysis, dedicated secretion systems, or membrane vesicles. eDNA is often involved in adhesion, particularly after the first interaction of the cell with the substratum. It mediates acid–base interactions and increases the hydrophobicity of bacterial cells which are favorable for the cell–substratum interaction[13][14]. Indeed, eDNA is used for initiation of biofilm formation in many pathogenic bacteria, including Gram-positive and Gram-negative bacteria and mycobacteria[15][7][16]. In addition, eDNA facilitates the interaction of the bacteria in the ECM. This is achieved by binding of positively charged segments of cell surface-exposed proteins with the negatively charged eDNA molecules[7]. Various proteins can be implicated in this interaction, such as autotransporters, lipoproteins or two-partner secretion protein A of Gram-negative bacteria, and cell wall-associated proteins in Gram-positive bacteria and fungi[17]. Thus, anchoring the eDNA to the cell surface by DNA-binding proteins is a widespread mechanism for biofilm formation that may also facilitate multispecies biofilms. eDNA can also mediate interactions with other ECM components such as polysaccharides[18]. Together, these interactions are relevant for the structural integrity of biofilms.
The composition of the polysaccharides present in the ECM varies between different bacterial species and even between different isolates of the same species. Most are long linear or branched molecules formed by one (homopolysaccharides) or several different (heteropolysaccharides) residues. They may contain substituents that greatly affect their biological properties. One of the most commonly studied polysaccharides is poly-β-1,6-N-acetyl-D-glucosamine, often named PGA or PNAG. It is synthetized by E. coli[19] and S. aureus[20], among others. In E. coli, PGA is required for initial cell-to-cell and cell-to-substratum attachment[19]. Another polysaccharide present in ECM is cellulose, a linear polymer of β-1,4 linked D-glucose. It is a major component of the ECM of some E. coli[21], Salmonella[22], and Pseudomonas strains[23]. Some E. coli strains produce a complex branched polysaccharide called colanic acid[24]. Additionally, P. aeruginosa can produce diverse exopolysaccharides. Mucoid P. aeruginosa strains produce alginate, a polymer of β-1-4-linked mannuronic acid and α-L-guluronate. Production of alginate confers a mucoid phenotype[25][26], typical of strains isolated from lungs of cystic fibrosis patients with Pseudomonas infections that underwent several rounds of antibiotic treatment. Therefore, secretion of alginate is related to pathogenic biofilms[27]. Alginate mediates the establishment of microcolonies at early stages of biofilm formation and provides stability to mature biofilms. Nonmucoid P. aeruginosa strains can produce other exopolysaccharides, e.g., Psl or Pel. Pel is a linear, cationic exopolysaccharide formed by 1®4 glycosidic linkages of N-acectylglucosamine and N-acetylgalactosamine. It has a critical role in maintaining cell-to-cell interactions and pellicle formation[28]. In contrast, Psl is composed of repeating pentasaccharide subunits of D-glucose, D-mannose, and L-rhamnose[29]. Psl mediates attachment to biotic surfaces such as mucin-coated epithelial surfaces and epithelial cells, indicating its relevance for the establishment of P. aeruginosa infection[30]. Additionally, P. aeruginosa strains can secrete cyclic and linear glucans[31][32] that are formed by β-1,3 linked glucose residues.
The proteinaceous content of the ECM includes proteins that are secreted through active secretion systems or released during cell lysis. The role of many of these proteins in the biofilm matrix is unknown, but some of them have been identified as important contributors to biofilm formation or restructuring in many pathogens. Various are extracellular enzymes. Their substrates can be polysaccharides, proteins, and nucleic acids, present in the ECM. They can function in remodeling of the ECM, detachment of cells from the biofilm, or degradation of polymers for nutrient acquisition.
ECM biogenesis and composition are dynamic and vary between strains of a given species and also depend on environmental conditions, such as nutrient availability and the presence of stressors, and on social crosstalk. Several functions have been attributed to the ECM based on its extraordinary capacity to establish intermolecular interactions between its components, and with surface-exposed structures of the cells, biotic and abiotic substrata, and many environmental molecules[12]. Thereby, the ECM immobilizes cells and keeps them in the biofilm community. By retaining the cells in close proximity, the ECM establishes the optimal conditions for interbacterial communication and exchange of genetic material, which is relevant, amongst others, for the dispersion of antibiotic-resistance genes. The ECM additionally retains water and thereby protects the cells against desiccation. Furthermore, the extracellular enzymes in that hydrated environment generate an external digestive system. In addition, ECM retains several other substances, for instance, nutrients, energy sources, antibiotics, antibiotic-degrading enzymes, and molecules released by cell lysis, thereby constituting a recycling unit [12]. In general, the ECM acts as a protective scaffold.
1.3. Biofilm Architecture
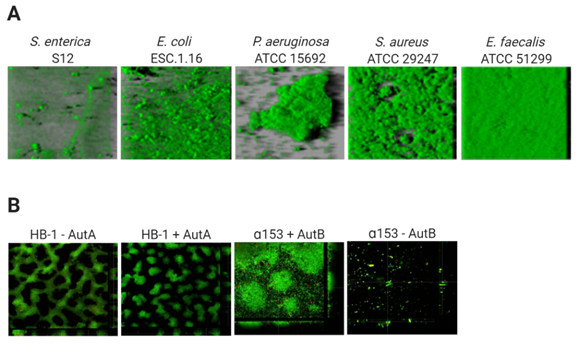
The architecture of biofilms is defined by the organization of the biomass and the spaces in between. The development of this structure depends on the composition of cell-surface structures mediating mutual interactions between cells and interactions of cells with ECM components and with the substratum[17]. The biofilm architecture is responsible for the generation of gradients of dispersion of substances within the biofilm. This will influence the accessibility of these substances to particular niches inside the biofilm, and determines, amongst others, the variation in antibiotic susceptibility of cells within biofilms. Figure 1 illustrates the biofilm architecture of different bacterial species. P. aeruginosa strain ATCC 15,692 forms complex biofilms with mushroom-like architectural features consisting of well-defined stalks and caps. Enterococcus faecalis ATCC 51,299 biofilms, however, are flat and compact[33], while Salmonella enterica strain S12 and E. coli strain ESC.1.16 form biofilms constituted of small cell clusters (Figure 1A)[33]. In contrast, biofilms of N. meningitidis strain HB-1 are constituted of cell aggregates of different sizes forming defined channel-like structures[7] (Figure 1B).
Figure 1. Variable architecture of biofilms. (A) Biofilms of five species (Salmonella enterica, Escherichia coli, Pseudomonas aeruginosa Staphylococcus aureus, Enterococcus faecalis) were formed under static conditions on abiotic surfaces during 24 h and were stained with Syto9, a green fluorescent nucleic acid marker. Reprinted from[33] with permission from Elsevier. (B) Strains of Neisseria meningitidis HB-1 and α153 and derivatives, which do or do not produce the autotransporters AutA and AutB (as indicated), formed biofilms under flow conditions during 14 h and were stained with the LIVE/DEAD Backlight bacterial viability stain (where red cells are dead and green cells are live). Reproduced from[34][35].
Thus, in general, based on their architecture, biofilms can be classified into (i) monolayer biofilms, formed by a compact layer with high surface coverage, or (ii) multilayer biofilms, formed by bacterial clusters of different morphology with a low surface interaction. The biofilm architecture can vary depending on different factors, for instance, the expression of surface-exposed proteins. Examples are the meningococcal autotransporters AutA and AutB, whose expression is phase variable and significantly alters the biofilm (Figure 1B)[34][35]. Additionally, the medium composition influences the biofilm architecture. P. aeruginosa PAO1 makes monolayer biofilms in the presence of citrate benzoate and casamino acids and multilayer biofilms in presence of glucose[36].
2. Biofilms mechanisms for Antibiotic Tolerance and Antibiotic Resistance
Biofilm recalcitrance comprises two independent phenomena: antibiotic resistance and antibiotic tolerance. Resistance refers to the capacity of a microorganism to survive and grow at increased antibiotic concentrations for long periods of time and is quantifiable by assessing the minimum inhibitory concentration (MIC)
[37]
. It involves mechanisms that prevent the binding of an antibiotic to its target, including enzymatic deactivation, active efflux of a drug once it is in the cytoplasm or the cytoplasmic membrane, or reduced influx, among others, and can be generated by HGT or mutations. Together, they preclude antibiotics from altering their target’s function and they prevent the production of toxic products that would end up damaging the cell. By contrast, antibiotic tolerance is the capacity of bacteria to survive a transient exposure to increased antibiotic concentrations, even those above the MIC. Tolerance is assessed by the minimum bactericidal concentration, that is, the minimum concentration of antibiotic required to kill 99.9% of the cells
[38]
. Unlike resistance, tolerance is only temporary and after longer exposure periods, the antibiotic will kill the bacteria. It is an adaptive phenomenon that implies a change in cellular behavior, from an active (growing) state to a quiescent (dormant) state
[39]
, and requires large metabolic rearrangements affecting, for example, energy production and nonessential functions. These changes are triggered during poor growth conditions or exposure to stress factors or antibiotics
. In this case, antibiotics can usually attach to the target molecules, but because their function is no longer essential, the microorganism survives. Tolerance in biofilms is also caused by entrapment of the antibiotics in the ECM, in this case, the antibiotic does not reach its target. In contrast to resistant cells, tolerant cells within the biofilm cannot grow in presence of a bactericidal antibiotic. Persistence is a special phenomenon of tolerance. Indeed, persistence is a phenomenon that increases the survival of a given population in the presence of bactericidal antibiotics without enhancing the MIC, but in contrast to tolerance, persistence only affects a subset of cells of the population called persisters
.
Persisters cells are tolerant cells that eventually can be killed at longer exposure times. There are two types of persisters, e.g., type I or triggered persistence, which is induced upon environmental signals, such as starvation, and type II or spontaneous persistence, where a subpopulation of growing bacteria converts into the persister state by a stochastic process
[41]
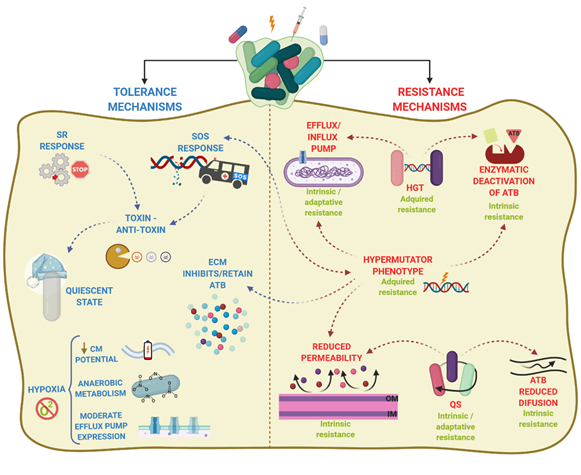
. Anyhow, persistence can be also referred to as heterotolerance, which is different than heteroresistance, as persisters can eventually be killed at longer exposure times. Figure 2 illustrates the mechanisms that govern antibiotic tolerance and antibiotic resistance of biofilms.
Figure 2.
Antibiotic resistance and antibiotic tolerance comprise a combination of mechanisms. Resistance mechanisms confer the ability to survive and grow at increased ATB concentrations for long periods and involve horizontal genetic transfer (HGT), hypermutation, and quorum sensing (QS), leading to transport of antibiotics via efflux pumps, reduced permeability of the outer membrane, or production of enzymes that inactivate ATB. The type of AMR is indicated in green. In contrast, tolerance mechanisms lead microorganisms to survive at increased ATB concentrations temporally, and involve activation of stress responses (SOS response, stringent response SR)) and hypoxia, leading to activation of a quiescent state, anaerobic metabolism, decrease of membrane potential, and moderate increase in efflux pump expression. Arrowhead lines indicate the interrelation between mechanisms. CM: cytoplasmic membrane.
Overall, antibiotic tolerance and antibiotic resistance do not depend on one unique mechanism but are a combination of both antibiotic tolerance and antibiotic resistance mechanisms. Such combination varies depending upon aspects such as the bacterial species or strain, the antimicrobial agent, the developmental stage of the biofilm, and the biofilm growth conditions.
3. Control of Biofilm Infections
As biofilms contribute to bacterial pathogenicity and recalcitrance, novel strategies and agents are required to deal with this issue. We have now clear evidence that the antibiotics used for the treatment of biofilm infections should be carefully selected, and such selection should consider the mechanisms of resistance and tolerance of biofilms. The use of cocktails of antibiotics would probably be more successful than a single antibiotic, but the antibiotic combination should also be thoroughly considered. Antibiotics should cover the heterogenic nature of biofilms. While one of the antibiotics in the combination should be active against persisters (e.g., colistin), others should target growing cells (e.g.
,
ciprofloxacin, tobramycin, or β-lactams). In addition, the selection of antibiotics will benefit from the characterization of ECM composition, particularly the sorption and charge of the matrix, as these properties are relevant contributors to AMR. Many alternatives to antibiotics have been proposed to inhibit and/or eradicate biofilms. Their nature and their mechanisms of action are ample. In general, they possess one or several activities as (i) biofilm inhibitors, (ii) biofilm dispersers, and (iii) antimicrobials. An overview of these substances is listed in Table 1.
Table 1.
Proposed alternatives to antibiotics with antimicrobial or antibiofilm activities. The substance, the mechanism of action (including anti-biofilm activity) and the target bacterial species are indicated for each agent.
|
Substance(s) |
Mechanism of Action |
Targets |
References |
|
Antimicrobial Peptides |
|||
|
|
|||
|
Melittin |
Formation of short-lived pores in the membrane and increase of permeability of OM |
P. aeruginosa, S. aureus, E. coli, K. pneumoniae, A. baumannii |
|
|
Japonicin-2LF |
Detergent-like activity against components of biofilm matrix; higher activity in inhibiting than in eradicating biofilms |
S. aureus, MRSA, E. coli |
[47] |
|
Magainin 2 |
Destabilizes the bacterial membrane and intracellular processes |
A. baumannii, P. aeruginosa, E. coli |
|
|
LL-37 |
Membrane disruption; inhibits twitching and QS; interferes in bacterial attachment; downregulates rhlA and rhlB genes |
P. aeruginosa, A. baumanni, S. aureus |
|
|
Temporin 1Tb |
Disruption of cell membrane integrity; capable of penetrating biofilm and killing bacteria; hemolytic activity |
S. epidermidis, S. aureus, K. pneumoniae, P. aeruginosa, E. faecium |
|
|
Synthetic Antimicrobial Peptides |
|||
|
1037 |
Downregulates genes of biofilm development; reduces swimming and swarming motilities |
P. aeruginosa, L. monocytogenes, Burkolderia cenocepacia |
[56] |
|
Esculentin (1–21) |
Biofilm eradication |
P. aeruginosa |
[57] |
|
1018 |
Binds (p)ppGpp and inhibits SR; inhibits attachment, QS, and twitching motility |
E. coli, S. aureus, MRSA, P. aeruginosa, A. baumannii, K. pneumoniae, A. baumannii, S. Typhimurium, E. faecium |
|
|
STAMP G10KHc |
Disrupts and permeabilizes OM and IM |
P. aeruginosa |
[60] |
|
F2,5,12W |
Reduces initial adhesion of bacteria; eliminates mature biofilms; suppresses biofilm formation |
S. epidermidis |
[61] |
|
Combined Therapies |
|||
|
1018 + antibiotics (e.g., ciprofloxacin) |
Inhibition of (p)ppGpp activation; downregulation of genes that interfere with antibiotic resistance and biofilm formation |
E. coli, MRSA, P. aeruginosa, K. pneumoniae, A. baumannii, S. enterica |
[58] |
|
Esculentin (1–21) + AuNPs (AuNPsEsc(1–21)) |
Disruption of membrane forming clusters |
P. aeruginosa |
[62] |
|
Temporin 1Tb + EDTA |
Mature biofilm eradication |
S. epidermidis |
[54] |
|
lin-SB056-1 + EDTA |
Perturbation of membrane; eradication biofilm; chelation of divalent metal ions |
P. aeruginosa |
[63] |
|
Bacteriophages |
|||
|
Phages |
|||
|
EFDG1 |
Mature biofilm eradication |
E. faecium, E. faecalis |
[64] |
|
vB_EfaH_EF1TV |
Mature biofilm eradication |
E. faecalis |
[65] |
|
vB_PaeM_LS1 |
Disrupts and avoids dispersion of biofilms; inhibits biofilm growth |
P. aeruginosa |
[66] |
|
vB_SauM_philPLA-RODI |
Penetrates biofilms; inhibits biofilm formation |
S. aureus S. epidermidis |
[67] |
|
Phage-derived Enzymes |
|||
|
LysAB3 |
Degradation of bacterial wall peptidoglycan, biofilm eradication |
A. baumannii |
[68] |
|
Dpo48 |
Degrades exopolysaccharide and eradicates biofilm |
A. baumannii |
[69] |
|
Combined Phage Therapy |
|||
|
Phage + amoxicillin |
Biofilm eradication |
K. pneumoniae |
[70] |
|
SAP-26 + rifampicin |
Hydrolysis of bacterial wall; mature biofilm eradication; reduction of biofilm growth |
S. aureus |
[71] |
|
Phage K + DRA88 |
Inhibits biofilm formation; disperses biofilms |
S. aureus |
[72] |
|
Phage K + its derivatives (e.g., K.MS811) |
Biofilm eradication |
S. aureus |
|
|
Phage M4 + E2005-24-39 + E2005-40-16 + W2005-24-39 + W2005-37-18-03 |
Biofilm eradication |
P. aeruginosa |
[75] |
|
DL52 + DL54 + DL60 + DL62 + DL64 + DL68 |
Attachment to cell by binding to lipopolysaccharide; biofilm eradication |
P. aeruginosa |
[76] |
|
Plant-Derived Natural Products |
|||
|
Essential Oils or Principal Active Compounds |
|||
|
Cinnamon (cinnamaldehyde) |
Inhibits QS mechanism: regulates production of rhamnolipids, proteases, and alginate and swarming activity; disrupts synthesis of DNA, RNA, proteins, lipids, and polysaccharides; alters expression of genes related to biofilm formation (e.g., icaA) |
E. coli, P. aeruginosa, K. pneumoniae, A. baumannii, S. epidermidis, S. aureus, MRSA, S. enteridis, S. Typhimurium |
|
|
Clove |
Disrupts QS communication: biofilm dispersal, inhibits AHL synthesis; downregulates relA gene |
E. coli, P. aeruginosa, K. pneumoniae, A. baumannii, S. aureus |
|
|
Thyme (thymol) |
Downregulates sarA gene; increases membrane permeability; penetrates polysaccharide matrix: eradicates biofilms |
E. coli, P. aeruginosa, K. pneumoniae, A. baumannii, S. aureus, S. enteridis |
|
|
Tea tree oil |
Alters expression of multiple genes related to biofilm formation (e.g., sarA, cidA, igrA, ifrB) |
S. aureus |
[84] |
|
Oregano (carvacrol) |
Increases membrane permeability; penetrates polysaccharide matrix; eradicates biofilms |
K. pneumoniae, P. aeruginosa, A. baumannii |
[82] |
|
Halogenated furanones |
QS inhibition; antagonist of LuxR |
E. coli P. aeruginosa |
|
|
Flavonoids (e.g., quercetin) |
Represses exopolysaccharides production; inhibits rpoS gene expression; decreases swimming motility |
S. aureus, E. coli, P. aeruginosa, E. faecalis |
|
|
Combined Therapy |
|||
|
Carvacrol + eugenol |
Increases membrane permeability |
K. pneumoniae, P. aeruginosa, A. baumannii, S. aureus |
|
|
Cinnamaldehyde + eugenol |
Membrane permeabilization |
S. epidermidis |
[92] |
|
Curcumin + antibiotics (e.g., ciprofloxacin) |
QS inhibition |
E. coli, K. pneumoniae, P. aeruginosa, S. aureus, E. faecalis |
[93] |
|
Enzymes |
|||
|
Dispersin B |
Hydrolyses PNAG |
S. epidermidis, S. aureus, E. coli, A. pleuropneumoniae |
|
|
DNases |
Hydrolyses DNA |
A. baumannii, K. pneumoniae, E. coli, P. aeruginosa, S. aureus |
|
|
Alginate lyase |
Degrades alginate |
P. aeruginosa |
[99] |
|
Lysozyme |
Hydrolytic activity |
S. pneumoniae, Gardnerella vaginalis, S. aureus, P. aeruginosa |
|
|
Lysostaphin |
Degrades cell wall |
S. aureus, S. epidermidis |
[103] |
|
Proteases (e.g., SpeB) |
Degrades cell wall |
Streptococcus spp. P. aeruginosa, S. aureus |
|
|
Paraoxonases (e.g., acylase I) |
Inhibits QS |
A. hydrophila, P. putida, P. aeruginosa |
|
|
Lactonase |
Inhibits QS |
P. aeruginosa |
|
|
Small molecules |
|||
|
Small molecules (e.g., LP 3134, LP 3145, LP 4010) |
Inhibition of diguanylate cyclase |
P. aeruginosa, A. baumannii |
[111] |
|
Pilicides (FN075, BibC6, Ec240) |
Blocks synthesis of and Type I pili, and inhibits chaperone-usher pathway for pili biogenesis |
E. coli |
|
|
Mannosides |
Inhibits FimH of type I pili |
E. coli |
[114] |
|
Ethyl pyruvate |
Inhibits enzymes of the glycolytic pathway |
E. coli |
[115] |
|
Polysaccharides |
|||
|
Psl, Pel |
Disperses biofilm |
S. epidermidis |
[116] |
4. Concluding remarks
The capacity of microorganisms to evolve and adapt to environmental cues has led to a health crisis as they became resistant to most, or almost all, commercial antibiotics. Biofilm formation is an ancient form of bacterial adaptation that contributes substantially to the problem because of their recalcitrance to treatment. Indeed, biofilms are the origin of significant morbidity and mortality. As discussed here, biofilm recalcitrance integrates many mechanisms, including metabolic heterogeneity, stress responses, efflux pump regulation, entrapment and inactivation of antibiotics in the ECM, interbacterial communication increased mutability, and exchange of genetic material.
Many of these factors have been discovered particularly in strains of
P. aeruginosa
. However, the specificity and multifaceted nature of the described mechanisms indicate the necessity of studying them also in other bacteria. Even more challenging, but necessary, will be to study biofilms in natural infections, where heterogeneous bacterial populations are common, and many environmental factors, including host defenses or diffusion of antibiotics in tissues, are present.
The understanding of the mechanisms that mediate recalcitrance will definitely guide therapeutic strategies to successfully deal with biofilm infections. These should be accompanied with methodologies for rapid diagnosis of biofilm infections and characterization of the biofilm biology and composition in vivo. Additionally, the availability of a panel of substances to inhibit and disperse biofilms will contribute to the selection of adequate therapeutic strategies to deal with particular biofilm infections.
References
- Daniel López; Hera Vlamakis; Roberto Kolter; Biofilms. Cold Spring Harbor Perspectives in Biology 2010, 2, a000398-a000398, 10.1101/cshperspect.a000398.
- David G. Davies; Matthew R. Parsek; James P. Pearson; Barbara H. Iglewski; J. W. Costerton; E. P. Greenberg; The Involvement of Cell-to-Cell Signals in the Development of a Bacterial Biofilm. Science 1998, 280, 295-298, 10.1126/science.280.5361.295.
- Muhsin Jamal; Wisal Ahmad; Saadia Andleeb; Fazal Jalil; Muhammad Imran; Muhammad Asif Nawaz; Tahir Hussain; Muhammad Ali; Muhammad Rafiq; Muhammad Atif Kamil; et al. Bacterial biofilm and associated infections. Journal of the Chinese Medical Association 2018, 81, 7-11, 10.1016/j.jcma.2017.07.012.
- Claire Prigent‐Combaret; Gerard Prensier; Thanh Thuy Le Thi; Olivier Vidal; Philippe Lejeune; Corinne Dorel; Developmental pathway for biofilm formation in curli-producing Escherichia coli strains: role of flagella, curli and colanic acid. Environmental Microbiology 2000, 2, 450-464, 10.1046/j.1462-2920.2000.00128.x.
- María Ángeles Tormo-Mas; Erwin Knecht; Friedrich Götz; Iñigo Lasa; José R. Penadés; Bap-dependent biofilm formation by pathogenic species of Staphylococcus: evidence of horizontal gene transfer?. Microbiology 2005, 151, 2465-2475, 10.1099/mic.0.27865-0.
- Bradley R. Borlee; Aaron D. Goldman; Keiji Murakami; Ram Samudrala; Daniel J. Wozniak; Matthew R. Parsek; Pseudomonas aeruginosauses a cyclic-di-GMP-regulated adhesin to reinforce the biofilm extracellular matrix. Molecular Microbiology 2010, 75, 827-842, 10.1111/j.1365-2958.2009.06991.x.
- Jesús Arenas; Reindert Nijland; Francisco Javier Rodríguez; Tom N. P. Bosma; Jan Tommassen; Involvement of three meningococcal surface-exposed proteins, the heparin-binding protein NhbA, the α-peptide of IgA protease and the autotransporter protease NalP, in initiation of biofilm formation. Molecular Microbiology 2012, 87, 254-268, 10.1111/mmi.12097.
- Jan Grijpstra; Jesús Arenas; Lucy Rutten; Jan Tommassen; Autotransporter secretion: varying on a theme. Research in Microbiology 2013, 164, 562-582, 10.1016/j.resmic.2013.03.010.
- George A. O'toole; Roberto Kolter; Flagellar and twitching motility are necessary forPseudomonas aeruginosabiofilm development. Molecular Microbiology 1998, 30, 295-304, 10.1046/j.1365-2958.1998.01062.x.
- Jan Tommassen; Jesús Arenas; Biological Functions of the Secretome of Neisseria meningitidis. Frontiers in Cellular and Infection Microbiology 2017, 7, 256-256, 10.3389/fcimb.2017.00256.
- María-Carmen Muñoz-Egea; María García-Pedrazuela; Ignacio Mahillo-Fernandez; Jaime Esteband; Effect of Antibiotics and Antibiofilm Agents in the Ultrastructure and Development of Biofilms Developed by Nonpigmented Rapidly Growing Mycobacteria. Microbial Drug Resistance 2016, 22, 1-6, 10.1089/mdr.2015.0124.
- Hans-Curt Flemming; Jost Wingender; The biofilm matrix. Nature Reviews Microbiology 2010, 8, 623-633, 10.1038/nrmicro2415.
- Theerthankar Das; Prashant K. Sharma; Henk J. Busscher; Henny C. Van Der Mei; Bastiaan Philip Krom; Role of Extracellular DNA in Initial Bacterial Adhesion and Surface Aggregation. Applied and Environmental Microbiology 2010, 76, 3405-3408, 10.1128/aem.03119-09.
- Theerthankar Das; Prashant K. Sharma; Bastiaan P. Krom; Henny C. Van Der Mei; Henk J. Busscher; Role of eDNA on the Adhesion Forces betweenStreptococcus mutansand Substratum Surfaces: Influence of Ionic Strength and Substratum Hydrophobicity. Langmuir 2011, 27, 10113-10118, 10.1021/la202013m.
- Zhiqiang Qin; Yuanzhu Ou; Liang Yang; Yuli Zhu; Tim Tolker-Nielsen; Soeren Molin; Di Qu; Role of autolysin-mediated DNA release in biofilm formation of Staphylococcus epidermidis. Microbiology 2007, 153, 2083-2092, 10.1099/mic.0.2007/006031-0.
- Cynthia B. Whitchurch; Tim Tolker-Nielsen; Paula C. Ragas; John S. Mattick; Extracellular DNA Required for Bacterial Biofilm Formation. Science 2002, 295, 1487-1487, 10.1126/science.295.5559.1487.
- Jesús Arenas; Jan Tommassen; Meningococcal Biofilm Formation: Let's Stick Together. Trends in Microbiology 2017, 25, 113-124, 10.1016/j.tim.2016.09.005.
- Midian C. Castillo Pedraza; Tatiana F. Novais; Roberta C. Faustoferri; Robert G. Quivey Jr; Anton Terekhov; Bruce R. Hamaker; Marlise I. Klein; Extracellular DNA and lipoteichoic acids interact with exopolysaccharides in the extracellular matrix of Streptococcus mutans biofilms. Biofouling 2017, 33, 722-740, 10.1080/08927014.2017.1361412.
- Xin Wang; James F. Preston; Tony Romeo; The pgaABCD Locus of Escherichia coli Promotes the Synthesis of a Polysaccharide Adhesin Required for Biofilm Formation. Journal of Bacteriology 2004, 186, 2724-2734, 10.1128/jb.186.9.2724-2734.2004.
- Era A. Izano; Matthew A. Amarante; William B. Kher; Jeffrey B. Kaplan; Differential Roles of Poly-N-Acetylglucosamine Surface Polysaccharide and Extracellular DNA in Staphylococcus aureus and Staphylococcus epidermidis Biofilms. Applied and Environmental Microbiology 2007, 74, 470-476, 10.1128/aem.02073-07.
- Sandra Da Re; Jean-Marc Ghigo; A CsgD-Independent Pathway for Cellulose Production and Biofilm Formation in Escherichia coli. Journal of Bacteriology 2006, 188, 3073-87, 10.1128/jb.188.8.3073-3087.2006.
- Cristina Solano; Begoña García; Jaione Valle; Carmen Berasain; Jean-Marc Ghigo; Carlos Gamazo; Iñigo Lasa; Genetic analysis ofSalmonella enteritidisbiofilm formation: critical role of cellulose. Molecular Microbiology 2002, 43, 793-808, 10.1046/j.1365-2958.2002.02802.x.
- Susanne Ude; Dawn L. Arnold; Christina D. Moon; Tracey Timms-Wilson; Andrew J. Spiers; Biofilm formation and cellulose expression among diverse environmental Pseudomonas isolates. Environmental Microbiology 2006, 8, 1997-2011, 10.1111/j.1462-2920.2006.01080.x.
- G Stevenson; K Andrianopoulos; M Hobbs; P R Reeves; Organization of the Escherichia coli K-12 gene cluster responsible for production of the extracellular polysaccharide colanic acid.. Journal of Bacteriology 1996, 178, 4885-4893, 10.1128/jb.178.16.4885-4893.1996.
- J Lam; R Chan; K Lam; J W Costerton; Production of mucoid microcolonies by Pseudomonas aeruginosa within infected lungs in cystic fibrosis.. Infection and Immunity 1980, 28, 546-556.
- Morten Hentzer; Gail M. Teitzel; Grant J. Balzer; Arne Heydorn; Søren Molin; Michael Givskov; Matthew R. Parsek; Alginate Overproduction Affects Pseudomonas aeruginosa Biofilm Structure and Function. Journal of Bacteriology 2001, 183, 5395-5401, 10.1128/jb.183.18.5395-5401.2001.
- Giovanna Pulcrano; Dora Vita Iula; Valeria Raia; Fabio Rossano; Maria Rosaria Catania; Different mutations in mucA gene of Pseudomonas aeruginosa mucoid strains in cystic fibrosis patients and their effect on algU gene expression.. The new microbiologica 2012, 35, 295.
- Laura K. Jennings; Kelly M. Storek; Hannah E. Ledvina; Charlène Coulon; Lindsey S. Marmont; Irina Sadovskaya; Patrick R. Secor; Boo Shan Tseng; Michele Scian; Alain Filloux; et al.Daniel J. WozniakP. Lynne HowellMatthew R. Parsek Pel is a cationic exopolysaccharide that cross-links extracellular DNA in thePseudomonas aeruginosabiofilm matrix. Proceedings of the National Academy of Sciences 2015, 112, 11353-11358, 10.1073/pnas.1503058112.
- Matthew S. Byrd; Irina Sadovskaya; Evgueny Vinogradov; Haiping Lu; April B. Sprinkle; Stephen H. Richardson; Luyan Ma; Brad Ralston; Matthew R. Parsek; Erin M. Anderson; et al.Joseph S. LamDaniel J. Wozniak Genetic and biochemical analyses of thePseudomonas aeruginosaPsl exopolysaccharide reveal overlapping roles for polysaccharide synthesis enzymes in Psl and LPS production. Molecular Microbiology 2009, 73, 622-638, 10.1111/j.1365-2958.2009.06795.x.
- Luyan Ma; Kara D. Jackson; Rebecca M. Landry; Matthew R. Parsek; Daniel J. Wozniak; Analysis of Pseudomonas aeruginosa Conditional Psl Variants Reveals Roles for the Psl Polysaccharide in Adhesion and Maintaining Biofilm Structure Postattachment. Journal of Bacteriology 2006, 188, 8213-8221, 10.1128/jb.01202-06.
- Yannick Lequette; Eglantine Rollet; Aurélie Delangle; E. Peter Greenberg; Jean-Pierre Bohin; Linear osmoregulated periplasmic glucans are encoded by the opgGH locus of Pseudomonas aeruginosa. Microbiology 2007, 153, 3255-3263, 10.1099/mic.0.2007/008953-0.
- Charlène Coulon; Evgeny Vinogradov; Alain Filloux; Irina Sadovskaya; Chemical Analysis of Cellular and Extracellular Carbohydrates of a Biofilm-Forming Strain Pseudomonas aeruginosa PA14. PLOS ONE 2010, 5, e14220, 10.1371/journal.pone.0014220.
- Arnaud Bridier; Florence Dubois-Brissonnet; A. Boubetra; V. Thomas; Romain Briandet; The biofilm architecture of sixty opportunistic pathogens deciphered using a high throughput CLSM method. Journal of Microbiological Methods 2010, 82, 64-70, 10.1016/j.mimet.2010.04.006.
- Jesús Arenas; Sara Cano; Reindert Nijland; Vérène Van Dongen; Lucy Rutten; Arie Van Der Ende; Jan Tommassen; The meningococcal autotransporter AutA is implicated in autoaggregation and biofilm formation. Environmental Microbiology 2014, 17, 1321-1337, 10.1111/1462-2920.12581.
- Jesús Arenas; Fernanda L. Paganelli; Patricia Rodríguez-Castaño; Sara Cano-Crespo; Arie Van Der Ende; Jos P. M. Van Putten; Jan Tommassen; Expression of the Gene for Autotransporter AutB of Neisseria meningitidis Affects Biofilm Formation and Epithelial Transmigration. Frontiers in Cellular and Infection Microbiology 2016, 6, 162, 10.3389/fcimb.2016.00162.
- P S Stewart; B M Peyton; W J Drury; R Murga; Quantitative observations of heterogeneities in Pseudomonas aeruginosa biofilms.. Applied and Environmental Microbiology 1993, 59, 327-329, 10.1128/aem.59.1.327-329.1993.
- Asher Brauner; Ofer Fridman; Orit Gefen; Nathalie Q. Balaban; Distinguishing between resistance, tolerance and persistence to antibiotic treatment. Nature Reviews Microbiology 2016, 14, 320-330, 10.1038/nrmicro.2016.34.
- Clayton W. Hall; Thien-Fah Mah; Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiology Reviews 2017, 41, 276-301, 10.1093/femsre/fux010.
- Mohsen Arzanlou; Wern Chern Chai; Henrietta Venter; Intrinsic, adaptive and acquired antimicrobial resistance in Gram-negative bacteria. Essays in Biochemistry 2017, 61, 49-59, 10.1042/ebc20160063.
- Nathalie Q. Balaban; Sophie Helaine; Kim Lewis; Martin Ackermann; Bree Aldridge; Dan I. Andersson; Mark P. Brynildsen; Dirk Bumann; Andrew Camilli; James J. Collins; et al.Christoph DehioSarah M. FortuneJean-Marc GhigoWolf-Dietrich HardtAlexander HarmsMatthias HeinemannDeborah T. HungUrs JenalBruce R. LevinJan MichielsGisela StorzMan-Wah TanTanel TensonLaurence Van MelderenAnnelies Zinkernagel Definitions and guidelines for research on antibiotic persistence. Nature Reviews Microbiology 2019, 17, 441-448, 10.1038/s41579-019-0196-3.
- Dorien Wilmaerts; Etthel M. Windels; Natalie Verstraeten; Jan Michiels; General Mechanisms Leading to Persister Formation and Awakening. Trends in Genetics 2019, 35, 401-411, 10.1016/j.tig.2019.03.007.
- Mohd Adil; Kunal Singh; Praveen K. Verma; Asad Ullah Khan; Eugenol-induced suppression of biofilm-forming genes in Streptococcus mutans: An approach to inhibit biofilms. Journal of Global Antimicrobial Resistance 2014, 2, 286-292, 10.1016/j.jgar.2014.05.006.
- Ramamourthy Gopal; Jun Ho Lee; Young Gwon Kim; Myeong-Sun Kim; Chang Ho Seo; Yoonkyung Park; Anti-Microbial, Anti-Biofilm Activities and Cell Selectivity of the NRC-16 Peptide Derived from Witch Flounder, Glyptocephalus cynoglossus. Marine Drugs 2013, 11, 1836-1852, 10.3390/md11061836.
- Sibel Dosler; Elif Karaaslan; A. Alev Gerceker; Antibacterial and anti-biofilm activities of melittin and colistin, alone and in combination with antibiotics against Gram-negative bacteria. Journal of Chemotherapy 2016, 28, 95-103, 10.1179/1973947815y.0000000004.
- T. Picoli; C.M. Peter; João Luíz Zani; Stefanie Bressan Waller; Matheus Gomes Lopes; Kamilla Neutzling Boesche; Gilberto Davila Vargas; Silvia De Oliveira Hübner; G. Fischer; Melittin and its potential in the destruction and inhibition of the biofilm formation by Staphylococcus aureus , Escherichia coli and Pseudomonas aeruginosa isolated from bovine milk. Microbial Pathogenesis 2017, 112, 57-62, 10.1016/j.micpath.2017.09.046.
- Ali Mohammadi Bardbari; Mohammad Reza Arabestani; Manoochehr Karami; Fariba Keramat; Hossein Aghazadeh; Mohammad Yousef Alikhani; Kamran Pooshang Bagheri; Highly synergistic activity of melittin with imipenem and colistin in biofilm inhibition against multidrug-resistant strong biofilm producer strains of Acinetobacter baumannii. European Journal of Clinical Microbiology & Infectious Diseases 2018, 37, 443-454, 10.1007/s10096-018-3189-7.
- Yipeng Yuan; Yu Zai; Xinping Xi; Chengbang Ma; Lei Wang; Mei Zhou; Chris Shaw; Tianbao Chen; A novel membrane-disruptive antimicrobial peptide from frog skin secretion against cystic fibrosis isolates and evaluation of anti-MRSA effect using Galleria mellonella model. Biochimica et Biophysica Acta (BBA) - General Subjects 2019, 1863, 849-856, 10.1016/j.bbagen.2019.02.013.
- Emre Yüksel; Ayşe Karakeçili; Antibacterial activity on electrospun poly(lactide-co-glycolide) based membranes via Magainin II grafting. Materials Science and Engineering: C 2014, 45, 510-518, 10.1016/j.msec.2014.10.004.
- Min Kyung Kim; Na Hee Kang; Su Jin Ko; Jonggwan Park; Eunji Park; Dong Won Shin; Seo Hyun Kim; Seung Hyun Lee; Ji In Lee; Eun Gi Ha; et al.Seung Hun JeonYoonkyung Park Antibacterial and Antibiofilm Activity and Mode of Action of Magainin 2 against Drug-Resistant Acinetobacter baumannii. International Journal of Molecular Sciences 2018, 19, 3041, 10.3390/ijms19103041.
- Nitzan Shtreimer Kandiyote; Gunasekaran Mohanraj; Canwei Mao; Roni Kasher; Christopher J. Arnusch; Synergy on Surfaces: Anti-Biofouling Interfaces Using Surface-Attached Antimicrobial Peptides PGLa and Magainin-2. Langmuir 2018, 34, 11147-11155, 10.1021/acs.langmuir.8b01617.
- J. Overhage; A. Campisano; M. Bains; E. C. W. Torfs; B. H. A. Rehm; R. E. W. Hancock; Human Host Defense Peptide LL-37 Prevents Bacterial Biofilm Formation. Infection and Immunity 2008, 76, 4176-4182, 10.1128/IAI.00318-08 IAI.00318-08 [pii].
- Xiaorong Feng; Karthik Sambanthamoorthy; Thomas Palys; Chrysanthi Paranavitana; The human antimicrobial peptide LL-37 and its fragments possess both antimicrobial and antibiofilm activities against multidrug-resistant Acinetobacter baumannii. Peptides 2013, 49, 131-137, 10.1016/j.peptides.2013.09.007.
- Jason Kang; Matthew J. Dietz; Bingyun Lie; Antimicrobial peptide LL-37 is bactericidal against Staphylococcus aureus biofilms. PLOS ONE 2019, 14, e0216676, 10.1371/journal.pone.0216676.
- Giuseppantonio Maisetta; Lucia Grassi; Mariagrazia Di Luca; Silvia Bombardelli; Chiara Medici; Franca Lisa Brancatisano; Semih Esin; Giovanna Batoni; Anti-biofilm properties of the antimicrobial peptide temporin 1Tb and its ability, in combination with EDTA, to eradicate Staphylococcus epidermidis biofilms on silicone catheters. Biofouling 2016, 32, 787-800, 10.1080/08927014.2016.1194401.
- Lucia Grassi; Giuseppantonio Maisetta; Giuseppe Maccari; Semih Esin; Giovanna Batoni; Analogs of the Frog-skin Antimicrobial Peptide Temporin 1Tb Exhibit a Wider Spectrum of Activity and a Stronger Antibiofilm Potential as Compared to the Parental Peptide. Frontiers in Chemistry 2017, 5, 24, 10.3389/fchem.2017.00024.
- César De La Fuente-Núñez; Fany Reffuveille; Evan F. Haney; Suzana K. Straus; Robert E. W. Hancock; Broad-Spectrum Anti-biofilm Peptide That Targets a Cellular Stress Response. PLOS Pathogens 2014, 10, e1004152, 10.1371/journal.ppat.1004152.
- Vincenzo Luca; Annarita Stringaro; Marisa Colone; Alessandro Pini; Maria Luisa Mangoni; Esculentin(1-21), an amphibian skin membrane-active peptide with potent activity on both planktonic and biofilm cells of the bacterial pathogen Pseudomonas aeruginosa. Cellular and Molecular Life Sciences 2013, 70, 2773-2786, 10.1007/s00018-013-1291-7.
- Fany Reffuveille; César De La Fuente-Núñez; Sarah Mansour; Robert E.W. Hancock; A Broad-Spectrum Antibiofilm Peptide Enhances Antibiotic Action against Bacterial Biofilms. Antimicrobial Agents and Chemotherapy 2014, 58, 5363-5371, 10.1128/aac.03163-14.
- Jelena Beljantseva; Pavel Kudrin; Steffi Jimmy; Marcel Ehn; Radek Pohl; Vallo Varik; Yuzuru Tozawa; Victoria Shingler; Tanel Tenson; Dominik Rejman; et al.Vasili Hauryliuk Molecular mutagenesis of ppGpp: turning a RelA activator into an inhibitor. Scientific Reports 2017, 7, srep41839, 10.1038/srep41839.
- Randal Eckert; Keith M. Brady; E. Peter Greenberg; Fengxia Qi; Daniel K. Yarbrough; Jian He; Ian McHardy; Maxwell H. Anderson; Wenyuan Shi; Enhancement of Antimicrobial Activity against Pseudomonas aeruginosa by Coadministration of G10KHc and Tobramycin. Antimicrobial Agents and Chemotherapy 2006, 50, 3833-3838, 10.1128/aac.00509-06.
- E. Margo Molhoek; Albert Van Dijk; Edwin J.A. Veldhuizen; Henk P Haagsman; Floris J. Bikker; A cathelicidin-2-derived peptide effectively impairs Staphylococcus epidermidis biofilms. International Journal of Antimicrobial Agents 2011, 37, 476-479, 10.1016/j.ijantimicag.2010.12.020.
- Bruno Casciaro; Maria Moros; Sara Rivera-Fernández; Andrea Bellelli; Jesús M. De La Fuente; Maria Luisa Mangoni; Gold-nanoparticles coated with the antimicrobial peptide esculentin-1a(1-21)NH 2 as a reliable strategy for antipseudomonal drugs. Acta Biomaterialia 2017, 47, 170-181, 10.1016/j.actbio.2016.09.041.
- Giuseppantonio Maisetta; Lucia Grassi; Semih Esin; Ilaria Serra; M. Andrea Scorciapino; Andrea C. Rinaldi; Giovanna Batoni; The Semi-Synthetic Peptide Lin-SB056-1 in Combination with EDTA Exerts Strong Antimicrobial and Antibiofilm Activity against Pseudomonas aeruginosa in Conditions Mimicking Cystic Fibrosis Sputum. International Journal of Molecular Sciences 2017, 18, 1994, 10.3390/ijms18091994.
- Leron Khalifa; Yair Brosh; Daniel Gelman; Shunit Coppenhagen-Glazer; Shaul Beyth; Ronit Poradosu-Cohen; Yok-Ai Que; Nurit Beyth; Ronen Hazan; Targeting Enterococcus faecalis Biofilms with Phage Therapy. Applied and Environmental Microbiology 2015, 81, 2696-2705, 10.1128/aem.00096-15.
- Marco Maria D’Andrea; Domenico Frezza; Elena Romano; Pasquale Marmo; Lucia Henrici De Angelis; Nicoletta Perini; Maria Cristina Thaller; Gustavo Di Lallo; The lytic bacteriophage vB_EfaH_EF1TV, a new member of the Herelleviridae family, disrupts biofilm produced by Enterococcus faecalis clinical strains. Journal of Global Antimicrobial Resistance 2020, 21, 68-75, 10.1016/j.jgar.2019.10.019.
- Yuyu Yuan; Kunli Qu; Demeng Tan; Xiaoyu Li; Lili Wang; Cong Cong; Zhilong Xiu; Yongping Xu; Isolation and characterization of a bacteriophage and its potential to disrupt multi-drug resistant Pseudomonas aeruginosa biofilms. Microbial Pathogenesis 2019, 128, 329-336, 10.1016/j.micpath.2019.01.032.
- Diana Gutiérrez; Dieter Vandenheuvel; Beatriz Martínez; Ana Rodríguez; Rob Lavigne; Pilar García; Two Phages, phiIPLA-RODI and phiIPLA-C1C, Lyse Mono- and Dual-Species Staphylococcal Biofilms. Applied and Environmental Microbiology 2015, 81, 3336-3348, 10.1128/aem.03560-14.
- Jie Zhang; Lu-Lu Xu; Dan Gan; Xingping Zhang; In Vitro Study of Bacteriophage AB3 Endolysin LysAB3 Activity Against Acinetobacter baumannii Biofilm and Biofilm-Bound A. baumannii. Clinical Laboratory 2018, 64, 1021-1030, 10.7754/clin.lab.2018.180342.
- Yannan Liu; Zhiqiang Mi; Liyuan Mi; Yong Huang; Puyuan Li; Huiying Liu; Xin Yuan; Wenkai Niu; Ning Jiang; Changqing Bai; et al.Zhan-Cheng Gao Identification and characterization of capsule depolymerase Dpo48 from Acinetobacter baumannii phage IME200. PeerJ 2019, 7, e6173, 10.7717/peerj.6173.
- Manmeet Sakshi Bedi; Vivek Verma; Sanjay Chhibber; Amoxicillin and specific bacteriophage can be used together for eradication of biofilm of Klebsiella pneumoniae B5055. World Journal of Microbiology and Biotechnology 2009, 25, 1145-1151, 10.1007/s11274-009-9991-8.
- Marzia Rahman; Shukho Kim; Sung Min Kim; Sung Yong Seol; Jungmin Kim; Characterization of inducedStaphylococcus aureusbacteriophage SAP-26 and its anti-biofilm activity with rifampicin. Biofouling 2011, 27, 1087-1093, 10.1080/08927014.2011.631169.
- D. R. Alves; A. Gaudion; J. E. Bean; P. Perez Esteban; T. C. Arnot; D. R. Harper; W. Kot; L. H. Hansen; M. C. Enright; A. Toby A. Jenkins; et al. Combined Use of Bacteriophage K and a Novel Bacteriophage To Reduce Staphylococcus aureus Biofilm Formation. Applied and Environmental Microbiology 2014, 80, 6694-6703, 10.1128/aem.01789-14.
- David Kelly; Olivia McAuliffe; Jim O’Mahony; A. Coffey; Development of a broad-host-range phage cocktail for biocontrol. Bioengineered Bugs 2011, 2, 31-37, 10.4161/bbug.2.1.13657.
- D. Kelly; O. McAuliffe; R.P. Ross; A. Coffey; Prevention of Staphylococcus aureus biofilm formation and reduction in established biofilm density using a combination of phage K and modified derivatives. Letters in Applied Microbiology 2012, 54, 286-291, 10.1111/j.1472-765x.2012.03205.x.
- Weiling Fu; Terri Forster; Oren Mayer; John J. Curtin; Susan M. Lehman; Rodney M. Donlan; Bacteriophage Cocktail for the Prevention of Biofilm Formation by Pseudomonas aeruginosa on Catheters in an In Vitro Model System. Antimicrobial Agents and Chemotherapy 2009, 54, 397-404, 10.1128/aac.00669-09.
- Diana R. Alves; P. Perez-Esteban; W. Kot; J.E. Bean; T. Arnot; L.H. Hansen; Mark C. Enright; A. Tobias A. Jenkins; A novel bacteriophage cocktail reduces and dispersesPseudomonas aeruginosabiofilms under static and flow conditions. Microbial Biotechnology 2015, 9, 61-74, 10.1111/1751-7915.12316.
- M.S.A. Khan; M. Zahin; S. Hasan; F.M. Husain; I. Ahmad; Inhibition of quorum sensing regulated bacterial functions by plant essential oils with special reference to clove oil. Letters in Applied Microbiology 2009, 49, 354-360, 10.1111/j.1472-765x.2009.02666.x.
- Titik Nuryastuti; Henny C. Van Der Mei; Henk J. Busscher; Susi Iravati; Abu T. Aman; Bastiaan Philip Krom; Effect of Cinnamon Oil on icaA Expression and Biofilm Formation by Staphylococcus epidermidis. Applied and Environmental Microbiology 2009, 75, 6850-6855, 10.1128/aem.00875-09.
- Natan De Jesus Pimentel-Filho; Mayra Carla De Freitas Martins; Guilherme Bicalho Nogueira; Hilário Cuquetto Mantovani; Maria Cristina Dantas Vanetti; Bovicin HC5 and nisin reduce Staphylococcus aureus adhesion to polystyrene and change the hydrophobicity profile and Gibbs free energy of adhesion. International Journal of Food Microbiology 2014, 190, 1-8, 10.1016/j.ijfoodmicro.2014.08.004.
- Manmohit Kalia; Vivek Kumar Yadav; Pradeep Kumar Singh; Deepmala Sharma; Himanshu Pandey; Shahid Suhail Narvi; Vishnu Agarwal; Effect of Cinnamon Oil on Quorum Sensing-Controlled Virulence Factors and Biofilm Formation in Pseudomonas aeruginosa. PLOS ONE 2015, 10, e0135495, 10.1371/journal.pone.0135495.
- Sana Alibi; Walid Ben Selma; José Ramos-Vivas; Mohamed Ali Smach; Ridha Touati; Jalel Boukadida; Jésus Navas; Hedi Ben Mansour; Anti-oxidant, antibacterial, anti-biofilm, and anti-quorum sensing activities of four essential oils against multidrug-resistant bacterial clinical isolates. Current Research in Translational Medicine 2020, 68, 59-66, 10.1016/j.retram.2020.01.001.
- P. Raei; Tala Pourlak; M. Y. Memar; N. Alizadeh; M. Aghamali; E. Zeinalzadeh; M. Asgharzadeh; Hossein Samadi Kafil; Thymol and carvacrol strongly inhibit biofilm formation and growth of carbapenemase-producing Gram negative bacilli. Cellular and Molecular Biology 2017, 63, 108-112, 10.14715/cmb/2017.63.5.20.
- Alaguvel Valliammai; Anthonymuthu Selvaraj; Udayakumar Yuvashree; Chairmandurai Aravindraja; Shunmugiah Karutha Pandian; sarA-Dependent Antibiofilm Activity of Thymol Enhances the Antibacterial Efficacy of Rifampicin Against Staphylococcus aureus. Frontiers in Microbiology 2020, 11, 1744, 10.3389/fmicb.2020.01744.
- Xingchen Zhao; Zonghui Liu; Zuojia Liu; Rizeng Meng; Ce Shi; Xiangrong Chen; Xiujuan Bu; Na Guo; Phenotype and RNA-seq-Based transcriptome profiling of Staphylococcus aureus biofilms in response to tea tree oil. Microbial Pathogenesis 2018, 123, 304-313, 10.1016/j.micpath.2018.07.027.
- Morten Hentzer; Kathrin Riedel; Thomas B. Rasmussen; Arne Heydorn; Jens Bo Andersen; Matthew R. Parsek; Scott A. Rice; Leo Eberl; Søren Molin; Niels Høiby; et al.Staffan KjellebergMichael Givskov Inhibition of quorum sensing in Pseudomonas aeruginosa biofilm bacteria by a halogenated furanone compound. Microbiology 2002, 148, 87-102, 10.1099/00221287-148-1-87.
- Michael Manefield; Thomas Bovbjerg Rasmussen; Morten Henzter; Jens Bo Andersen; Peter Steinberg; Staffan Kjelleberg; Michael Givskov; Halogenated furanones inhibit quorum sensing through accelerated LuxR turnover. Microbiology 2002, 148, 1119-1127, 10.1099/00221287-148-4-1119.
- Zoya Samoilova; Nadezda Muzyka; Elena Lepekhina; Oleg Oktyabrsky; Galina V. Smirnova; Medicinal plant extracts can variously modify biofilm formation in Escherichia coli. Antonie van Leeuwenhoek 2014, 105, 709-722, 10.1007/s10482-014-0126-3.
- Sérgio Dias Da Costa Júnior; João Victor De Oliveira Santos; Luís André De Almeida Campos; Marcela Araújo Pereira; Nereide Stela Santos Magalhães; Isabella Macário Ferro Cavalcanti; Antibacterial and antibiofilm activities of quercetin against clinical isolates of Staphyloccocus aureus and Staphylococcus saprophyticus with resistance profile. International Journal of Environment, Agriculture and Biotechnology 2018, 3, 1948-1958, 10.22161/ijeab/3.5.50.
- J. Ouyang; F. Sun; W. Feng; Y. Sun; X. Qiu; L. Xiong; Y. Liu; Y. Chen; Quercetin is an effective inhibitor of quorum sensing, biofilm formation and virulence factors in Pseudomonas aeruginosa. Journal of Applied Microbiology 2016, 120, 966-974, 10.1111/jam.13073.
- R.J.W. Lambert; P.N. Skandamis; P.J. Coote; G.-J.E. Nychas; A study of the minimum inhibitory concentration and mode of action of oregano essential oil, thymol and carvacrol. Journal of Applied Microbiology 2001, 91, 453-462, 10.1046/j.1365-2672.2001.01428.x.
- Rashin Namivandi-Zangeneh; Yiling Yang; Sihao Xu; Edgar H. H. Wong; Cyrille Boyer; Antibiofilm Platform based on the Combination of Antimicrobial Polymers and Essential Oils. Biomacromolecules 2019, 21, 262-272, 10.1021/acs.biomac.9b01278.
- Garima Sharma; Kiran Raturi; Shweta Dang; Sanjay Gupta; Reema Gabrani; Inhibitory effect of cinnamaldehyde alone and in combination with thymol, eugenol and thymoquinone against Staphylococcus epidermidis. Journal of Herbal Medicine 2017, 9, 68-73, 10.1016/j.hermed.2016.11.001.
- Arunava Kali; Devaraj Bhuvaneshwar; Pravin M. V. Charles; Kunigal Srinivasaiah Seetha; Antibacterial synergy of curcumin with antibiotics against biofilm producing clinical bacterial isolates. Journal of Basic and Clinical Pharmacy 2016, 7, 93-6, 10.4103/0976-0105.183265.
- E. Burton; N. Yakandawala; K. LoVetri; Srinivasa Madhyastha; A microplate spectrofluorometric assay for bacterial biofilms. Journal of Industrial Microbiology & Biotechnology 2006, 34, 1-4, 10.1007/s10295-006-0086-3.
- Era A. Izano; Irina Sadovskaya; Hailin Wang; Evgeny Vinogradov; Chandran Ragunath; Narayanan Ramasubbu; Saïd Jabbouri; Malcolm B. Perry; Jeffrey B. Kaplan; Poly-N-acetylglucosamine mediates biofilm formation and detergent resistance in Aggregatibacter actinomycetemcomitans. Microbial Pathogenesis 2008, 44, 52-60, 10.1016/j.micpath.2007.08.004.
- Kati Seidl; Christiane Goerke; Christiane Wolz; Dietrich Mack; Brigitte Berger-Bächi; Markus Bischoff; Staphylococcus aureus CcpA Affects Biofilm Formation. Infection and Immunity 2008, 76, 2044-2050, 10.1128/iai.00035-08.
- Victor Tetz; Natalia K. Artemenko; Victor V. Tetz; Effect of DNase and Antibiotics on Biofilm Characteristics. Antimicrobial Agents and Chemotherapy 2008, 53, 1204-1209, 10.1128/aac.00471-08.
- Quinn M. Parks; Robert L. Young; Katie R. Poch; Kenneth C. Malcolm; Michael L. Vasil; Jerry A. Nick; Neutrophil enhancement of Pseudomonas aeruginosa biofilm development: human F-actin and DNA as targets for therapy. Journal of Medical Microbiology 2009, 58, 492-502, 10.1099/jmm.0.005728-0.
- Deborah M. Ramsey; Daniel J. Wozniak; Understanding the control of Pseudomonas aeruginosa alginate synthesis and the prospects for management of chronic infections in cystic fibrosis. Molecular Microbiology 2005, 56, 309-322, 10.1111/j.1365-2958.2005.04552.x.
- Waldemar Vollmer; Alexander Tomasz; ThepgdAGene Encodes for a PeptidoglycanN-Acetylglucosamine Deacetylase inStreptococcus pneumoniae. Journal of Biological Chemistry 2000, 275, 20496-20501, 10.1074/jbc.m910189199.
- Mirsada Hukić; Dzenita Seljmo; Amra Ramovic; Monia Avdić Ibrišimović; Serkan Dogan; Jasna Hukic; Elma Feric Bojic; The Effect of Lysozyme on Reducing Biofilms byStaphylococcus aureus,Pseudomonas aeruginosa, andGardnerella vaginalis: AnIn VitroExamination. Microbial Drug Resistance 2018, 24, 353-358, 10.1089/mdr.2016.0303.
- Mohamed ElAdawy; Mohammed El-Mowafy; Mohamed Mohamed Adel El-Sokkary; Rasha Barwa; Effects of Lysozyme, Proteinase K, and Cephalosporins on Biofilm Formation by Clinical Isolates of Pseudomonas aeruginosa.. Interdisciplinary Perspectives on Infectious Diseases 2020, 2020, 6156720-9, 10.1155/2020/6156720.
- Maria Do Carmo De Freire Bastos; Bruna Goncalves Coutinho; Marcus Lívio Varella Coelho; Lysostaphin: A Staphylococcal Bacteriolysin with Potential Clinical Applications. Pharmaceuticals 2010, 3, 1139-1161, 10.3390/ph3041139.
- Kristie L. Connolly; Amity L. Roberts; Robert C. Holder; Sean D. Reid; Dispersal of Group A Streptococcal Biofilms by the Cysteine Protease SpeB Leads to Increased Disease Severity in a Murine Model. PLOS ONE 2011, 6, e18984, 10.1371/journal.pone.0018984.
- Joo-Hyeon Park; Jin-Hyung Lee; Moo Hwan Cho; Moshe Herzberg; Jintae Lee; Acceleration of protease effect on Staphylococcus aureus biofilm dispersal. FEMS Microbiology Letters 2012, 335, 31-38, 10.1111/j.1574-6968.2012.02635.x.
- Vipin C. Kalia; Sanjay K.S. Patel; Yun Chan Kang; Jung-Kul Lee; Quorum sensing inhibitors as antipathogens: biotechnological applications. Biotechnology Advances 2019, 37, 68-90, 10.1016/j.biotechadv.2018.11.006.
- Fan Yang; Lian-Hui Wang; Jing Wang; Yi-Hu Dong; Jiang Yong Hu; Lian-Hui Zhang; Quorum quenching enzyme activity is widely conserved in the sera of mammalian species. FEBS Letters 2005, 579, 3713-3717, 10.1016/j.febslet.2005.05.060.
- Diby Paul; Young Sam Kim; Kannan Ponnusamy; Jihyang Kweon; Application of Quorum Quenching to Inhibit Biofilm Formation. Environmental Engineering Science 2009, 26, 1319-1324, 10.1089/ees.2008.0392.
- Benjamin Rémy; Sonia Mion; Laure Plener; Mikael Elias; Eric Chabrière; David Daudé; Interference in Bacterial Quorum Sensing: A Biopharmaceutical Perspective. Frontiers in Pharmacology 2018, 9, 203, 10.3389/fphar.2018.00203.
- Parul Gupta; Sanjay Chhibber; Kusum Harjai; Efficacy of purified lactonase and ciprofloxacin in preventing systemic spread of Pseudomonas aeruginosa in murine burn wound model. Burns 2015, 41, 153-162, 10.1016/j.burns.2014.06.009.
- Karthik Sambanthamoorthy; Chunyuan Luo; Nagarajan Pattabiraman; Xiarong Feng; Benjamin Koestler; Christopher M. Waters; Thomas J. Palys; Identification of small molecules inhibiting diguanylate cyclases to control bacterial biofilm development. Biofouling 2013, 30, 17-28, 10.1080/08927014.2013.832224.
- Lynette Cegelski; Jerome S Pinkner; Neal D Hammer; Corinne K Cusumano; Chia S Hung; Erik Chorell; Veronica Åberg; Jennifer N Walker; Patrick C Seed; Fredrik Almqvist; et al.Matthew ChapmanScott J Hultgren Small-molecule inhibitors target Escherichia coli amyloid biogenesis and biofilm formation. Nature Chemical Biology 2009, 5, 913-919, 10.1038/nchembio.242.
- Sarah E. Greene; Jerome S. Pinkner; Erik Chorell; Karen W. Dodson; Carrie L. Shaffer; Matt S. Conover; Jonathan Livny; Maria Hadjifrangiskou; Fredrik Almqvist; Scott J Hultgren; et al. Pilicide ec240 Disrupts Virulence Circuits in Uropathogenic Escherichia coli. mBio 2014, 5, e02038-14, 10.1128/mbio.02038-14.
- Corinne K. Cusumano; Jerome S. Pinkner; Zhenfu Han; Sarah E. Greene; Bradley A. Ford; Jan R. Crowley; Jeffrey P. Henderson; James W. Janetka; Scott J Hultgren; Treatment and Prevention of Urinary Tract Infection with Orally Active FimH Inhibitors. Science Translational Medicine 2011, 3, 109ra115-109ra115, 10.1126/scitranslmed.3003021.
- Tewodros Debebe; Monika Krüger; Klaus Huse; Johannes Kacza; Katja Mühlberg; Brigitte König; Gerd Birkenmeier; Ethyl Pyruvate: An Anti-Microbial Agent that Selectively Targets Pathobionts and Biofilms. PLOS ONE 2016, 11, e0162919, 10.1371/journal.pone.0162919.
- Zhiqiang Qin; Liang Yang; D. Qu; S. Molin; Tim Tolker-Nielsen; Pseudomonas aeruginosa extracellular products inhibit staphylococcal growth, and disrupt established biofilms produced by Staphylococcus epidermidis. Microbiology 2009, 155, 2148-2156, 10.1099/mic.0.028001-0.