During the selection process of probiotics for vaginal applications, twenty-five lactic acid bacteria (LAB) isolates from human vagina belonging to six different species were tested for antimicrobial resistance by a microdilution method. Gene-specific PCR amplifications proved the strains carry no acquired antibiotic resistance genes, except for a
tet(W) gene present in two tetracycline-susceptible Bifidobacterium bifidum strains. Genome analysis of a selected set of strains showed no other acquired resistance determinants. The tet(W) of B. bifidum was inactive by the insertion of two guanine residues in the middle of the gene. Surprisingly, the inactive gene became active and functional very easily, providing resistance to tetracycline and remaining stable afterward. LAB intended to be used in health applications must be free of acquired antimicrobial resistance genes; these could be spread and transferred to human pathogens.
- lactic acid bacteria
- antibiotic resistance
- vaginal microbiota
- genome analysis
- tet(W)
1. Isolation, Identification and Typing of Vaginal LAB
Twenty-five vaginal LAB isolates with clear acidification halos on MRS agar supplemented with 0.5% CaCO
3
Lactobacillus crispatus
Lactobacillus salivarius
Lactobacillus jensenii
Lactobacillus paracasei,
Lactobacillus reuteri,
Bifidobacterium bifidum
L. crispatus
L. salivarius
L. jensenii
L. paracasei
, L. reuteri
,
B. bifidum (two strains) (Supplementary Figure S1).
Table 1. Minimum inhibitory concentration (MIC) values of 16 antibiotics to the vaginal LAB species and strains of this study.
| Species | Strain | Antibiotic (MIC as µg mL | −1 | ) | |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GEN | KAN | STR | NEO | TET | ERY | CLI | CHL | AMP | PEN | VAN | QDA | LIN | TMP | CIP | RIF | ||||||||||||||||||||||||||||||||||
| L. crispatus | VA20-32AN | 2 | 16 | 2 | 16 | 2 | 0.06 | 0.25 | 4 | 1 | 0.5 | 0.5 | 1 | 4 | >64 | 16 | 1 | ||||||||||||||||||||||||||||||||
| VA27-7 | 4 | 32 | 64 | 8 | 1 | 1 | 2 | 8 | 2 | 2 | 1 | 1 | 4 | 32 | 64 | 4 | |||||||||||||||||||||||||||||||||
| VA27-9 | 1 | 16 | 2 | 2 | 2 | 0.03 | 0.5 | 4 | 2 | 0.5 | 0.5 | 1 | 4 | 64 | 32 | 2 | |||||||||||||||||||||||||||||||||
| VA28-12 | 1 | 16 | 2 | 2 | 2 | 0.06 | 0.5 | 4 | 2 | 0.5 | 0.5 | 2 | 4 | 64 | 32 | 2 | |||||||||||||||||||||||||||||||||
| VA32-17 | 2 | 64 | 2 | 8 | 4 | 0.03 | 0.5 | 2 | 1 | 1 | 0.5 | 1 | 2 | >64 | 64 | 8 | |||||||||||||||||||||||||||||||||
| VA32-17AN | 4 | 128 | 32 | 4 | 2 | 0.25 | 0.5 | 4 | 1 | 0.5 | 1 | 1 | 2 | 16 | 32 | 4 | |||||||||||||||||||||||||||||||||
| VA50-4AN | ≤0.5 | 32 | 1 | 2 | 4 | 0.12 | 0.12 | 4 | 4 | 1 | 0.5 | 1 | 4 | >64 | 32 | 4 | |||||||||||||||||||||||||||||||||
| L. jensenii | VA04-1AN | ≤0.5 | 4 | 2 | 1 | 0.25 | ≤0.016 | 0.12 | 4 | 0.25 | 0.12 | 1 | 0.5 | 1 | >64 | 8 | 0.25 | ||||||||||||||||||||||||||||||||
| VA04-2AN | ≤0.5 | 4 | 4 | 1 | 0.5 | 0.03 | 0.12 | 2 | 0.5 | 1 | 1 | 0.5 | 2 | >64 | 8 | 0.25 | |||||||||||||||||||||||||||||||||
| VA15-2AN | ≤0.5 | ≤2 | 1 | ≤0.5 | 1 | ≤0.016 | ≤0.03 | 2 | 0.06 | 0.06 | 0.5 | 0.5 | 0.5 | >64 | 8 | 0.25 | |||||||||||||||||||||||||||||||||
| VA16-11 | ≤0.5 | 8 | 1 | 2 | 4 | 0.06 | 0.25 | 4 | 0.06 | ≤0.03 | 2 | 0.5 | 2 | >64 | 16 | 0.5 | |||||||||||||||||||||||||||||||||
| Breakpoint (µg mL−1) | 16 | 16 | 16 | - | 4 | 1 | 4 | 4 | 2 | - | 2 | - | - | - | - | - | |||||||||||||||||||||||||||||||||
| L. salivarius | VA09-4 | 8 | 64 | 16 | 4 | 2 | 0.25 | 0.25 | 2 | 1 | 0.25 | 128 | 0.25 | 0.5 | ≤0.12 | 1 | 2 | ||||||||||||||||||||||||||||||||
| VA16-20 | ≤0.5 | 4 | 2 | 0.5 | 1 | 0.06 | 0.06 | 2 | 0.5 | 0.12 | >128 | 0.5 | 0.5 | 0.25 | 0.5 | 0.5 | |||||||||||||||||||||||||||||||||
| VA37-13 | ≤0.5 | 4 | ≤0.5 | ≤0.5 | 0.5 | 0.06 | 0.06 | 2 | 0.25 | 0.12 | >128 | 0.5 | 0.5 | 0.25 | ≤0.25 | 0.5 | |||||||||||||||||||||||||||||||||
| VA40-10 | 128 | >1024 | >256 | 256 | 2 | 1 | 1 | 4 | 1 | 0.25 | >128 | 1 | 1 | 1 | 4 | 0.5 | |||||||||||||||||||||||||||||||||
| VA40-12AN | 4 | 128 | 32 | 4 | 2 | 0.25 | 0.25 | 4 | 0.5 | 0.25 | >128 | 1 | 0.5 | 0.25 | 1 | 1 | |||||||||||||||||||||||||||||||||
| VA40-14AN | 4 | 128 | 32 | 4 | 2 | 0.25 | 0.5 | 4 | 0.5 | 0.25 | >128 | 1 | 0.5 | ≤0.12 | 1 | 1 | |||||||||||||||||||||||||||||||||
| Breakpoint (µg mL−1) | 16 | 64 | 64 | - | 8 | 1 | 4 | 4 | 4 | - | n.r. | - | - | - | - | - | |||||||||||||||||||||||||||||||||
| L. paracasei | VA02-1AN | ≤0.5 | 16 | 8 | 1 | 2 | 0.12 | 0.06 | 8 | 1 | 0.25 | >128 | 1 | 4 | 0.5 | 4 | 0.5 | ||||||||||||||||||||||||||||||||
| VA24-4 | 1 | 16 | 8 | 4 | 4 | 0.12 | 0.06 | 4 | 0.5 | 0.25 | >128 | 1 | 2 | 0.25 | 4 | 0.5 | |||||||||||||||||||||||||||||||||
| VA26-3 | ≤0.5 | 16 | 8 | 2 | 2 | 0.12 | 0.06 | 4 | 1 | 0.25 | >128 | 1 | 2 | 1 | 2 | 0.5 | |||||||||||||||||||||||||||||||||
| VA27-8 | 1 | 32 | 16 | 8 | 2 | 0.06 | 0.06 | 8 | 0.5 | 0.25 | >128 | 1 | 4 | 0.25 | 4 | 0.5 | |||||||||||||||||||||||||||||||||
| Breakpoint (µg mL−1) | 32 | 64 | 64 | - | 4 | 1 | 4 | 4 | 4 | - | n.r. | - | - | - | - | - | |||||||||||||||||||||||||||||||||
| L. reuteri | VA15-3 | ≤0.5 | 4 | 2 | ≤0.5 | 8 | 0.12 | ≤0.03 | 4 | 1 | 2 | >128 | 1 | 2 | >64 | 32 | 0.25 | ||||||||||||||||||||||||||||||||
| VA24-5 | ≤0.5 | 16 | 4 | ≤0.5 | 16 | 0.06 | ≤0.03 | 4 | 2 | 8 | >128 | 0.5 | 4 | >64 | 32 | 0.25 | |||||||||||||||||||||||||||||||||
| Breakpoint (µg mL−1) | 8 | 64 | 64 | - | 32 | 1 | 4 | 4 | 2 | - | n.r. | - | - | - | - | - | |||||||||||||||||||||||||||||||||
| B. bifidum | VA07-1AN | 8 | 64 | >256 | 16 | 1 | ≤0.016 | 0.06 | 1 | ≤0.03 | ≤0.03 | 0.5 | 0.5 | 0.5 | 16 | 8 | 2 | ||||||||||||||||||||||||||||||||
| VA07-2AN | 32 | 64 | >256 | 32 | 1 | ≤0.016 | ≤0.03 | 1 | ≤0.03 | ≤0.03 | 1 | 0.5 | 0.5 | 16 | 8 | 1 | |||||||||||||||||||||||||||||||||
| Breakpoint (µg mL−1) | 64 | - | 128 | - | 8 | 1 | 1 | 4 | 2 | - | 2 | - | - | - | - | - | |||||||||||||||||||||||||||||||||
2. Antibiotic Susceptibility
Table 1 shows the MIC values of the 16 tested antibiotics for the 25 vaginal LAB isolates. All isolates were phenotypically susceptible to tetracycline, erythromycin, clindamycin, penicillin, quinupristin-dalfopristin, linezolid, and rifampicin. The distribution of neomycin MICs covered more than nine 2-fold dilutions, ranging from ≤0.5 to 256 µg mL
−1
−1
−1
−1
−1
−1
−1
B. bifidum
−1
L. salivarius VA40-10 was highly resistant to all four aminoglycosides tested (gentamicin, kanamycin, streptomycin, and neomycin).
3. Detection of AR Genes by PCR
The presence of genes coding for the commonest AR genes spread among LAB was investigated by PCR. No genes involved in resistance to chloramphenicol (
cat
bla
aac(6′)-aph(2″)
aad
erm
erm
erm
erm
mef
tet
tet
tet
tet
tet
lsaA
vanA
B. bifidum
tet
tet(W) gene highly homologous to those present in many Gram-positive and Gram-negative bacteria.
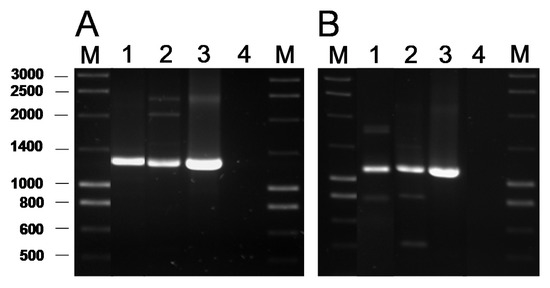
Figure 1.
A
tet
B
B. bifidum
B. bifidum
Leuconostoc mesenteroides
mesenteroides LbE16 (positive control) [1]; line 4, blank (no template DNA). M, molecular weight marker.
4. Genome Analysis for AR Genes
Based on the phenotype and genotype results (Table 1; Supplementary Figure S1), six strains were subjected to genome sequencing:
L. crispatus
L. jensenii
L. salivarius
L. paracasei
L. reuteri
B. bifidum VA07-1AN (resistant to streptomycin). Supplementary Table S2 shows the general features of their genomes. Their size was, in all cases, around 2.2 Mbp but the number of contigs obtained after assembly ranged from 17 to 300. Supplementary Table S3 summarizes some of the key genetic features of the genomes of the sequenced strains. Genes coding for penicillin binding proteins (PBP) were found in all the genomes, although with different numbers and types for the distinct species. Mutations in PBPs encoding-genes known to confer AR were not identified. One gene coding for a D-alanine-D-alanine ligase (Ddl) was detected in each of the strains. In several LAB species, the presence of phenylalanine at the enzyme active site in Ddl is correlated with intrinsic resistance to vancomycin [2]. In addition, in each of the strains, 9-32 genes were classified by the RAST server as belonging to the category “Virulence, Disease, and Defence”, subcategory “Resistance to Antibiotic and Toxic Compounds”. The majority of these genes encoded components dedicated to homeostasis or resistance to heavy metals, such as copper, mercury, and the cobalt-zinc-cadmium triad. Genes encoding elongation factors, efflux pumps, DNA gyrases, and topoisomerases were also included by RAST in this subcategory.
By comparing the genome sequences against the databases CARD, ResFinder, and ARG-ANNOT, no genes known to be involved in AR in
L. jensenii
L. paracasei
L. reuteri
Leuconostoc mesenteroides enzyme [2], in the deduced sequence of all vancomycin-resistant (Vm
r
s) strains were characterized by the presence of a tyrosine (Y) residue at this position (Figure 2).

Figure 2.
Lactobacillus spp. strains sequenced. Strains with phenylalanine (F) at the enzyme active site (green) show a vancomycin-resistant phenotype, while those having a tyrosine (Y) (pale blue) display a vancomycin-susceptible phenotype.
Genome analysis of
L. crispatus
L. salivarius
B. bifidum
L. crispatus
L. salivarius
L. crispatus
B. bifidum
L. salivarius VA40-10 showed six exclusive amino acid changes at positions 12 (G→E), 67 (D→N), 186 (N→D), 199 (I→V), 208 (Q→K), and 209 (V→I). However, by comparing RsmG sequences from resistant and susceptible strains, none of the changes considered could be associated with streptomycin resistance.
As expected, the genome analysis confirmed the presence of
tet
B. bifidum
tet
B. bifidum
tet
tet
Bifidobacterium longum
tet
B. bifidum
Lachnospiraceae
Ruminococcus
Cutibacterium acnes
tet
B. longum
tet
B. bifidum
tet
B. longum
B. bifidum in the downstream region (Supplementary Table S4).
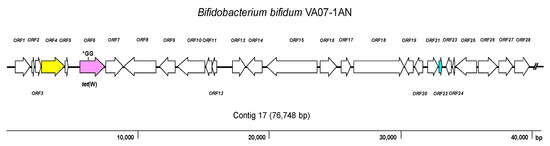
Figure 3.
tet
Bifidobacterium bifidum
tet(W) gene (the position of the GG insertion disrupting the ORF is indicated); yellow, conjugation-associated gene; pale blue, gene encoding a transcription regulator; white, genes involved in other processes. The broken line symbol indicates the contig extends beyond this point.
The CARD database further identified in the genome of
B. bifidum
rpsL gene (encoding the ribosomal S12 protein), a variation causing an amino acid substitution (K→R) at position 43 of the protein (Figure 4). This amino acid change has been associated with streptomycin resistance in many species [3].
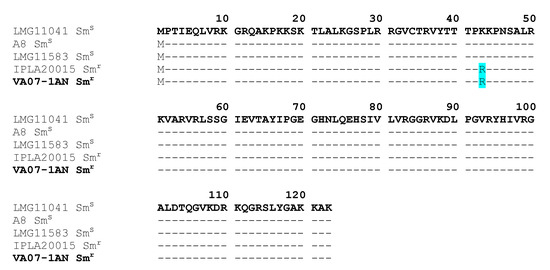

Figure 4.
rpsL
r
s
Bifidobacterium bifidum strains. The amino acid replacement K→R at position 43 in the resistant strains is highlighted in pale blue. In bold, the strain of this study (VA07-1AN).
5. Restoration of the Tetracycline Resistance Phenotype in B. Bifidum VA07-1AN
When the susceptibility of
B. bifidum
tet
tet
−1. In contrast, growing the antibiotic-resistant variants in the absence of tetracycline for about 80-100 generations showed no tetracycline-susceptible revertants, demonstrating high stability of the mutations that restored the resistant phenotype.
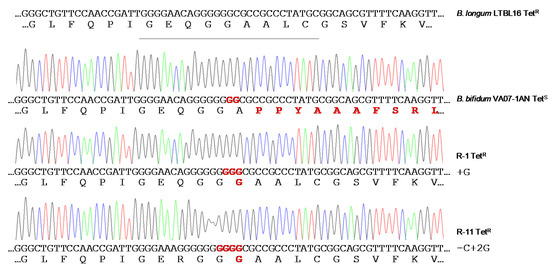
Figure 5.
tet
B. bifidum
tet
Bifidobacterium longum LTBL16 strain (on top of the figure) are highlighted in red.
6. Discussion
LAB contribute to the maintenance of vaginal health via the production of substances (mainly organic acids) that acidify the environment and inhibit the development of pathogens [4]. However, there is an increasing concern that LAB may act as reservoirs of AR determinants, from which they could ultimately be transferred to pathogens [5][6]. Indeed, the existence of lactobacilli and bifidobacteria strains resistant to several antibiotics, by either acquiring mutations or exogenous genes, has been repeatedly reported [7][8][9][10][11]. Therefore, during the selection of probiotics, the susceptibility of lactobacilli and bifidobacteria to antibiotics has to be assessed and the absence in the selected strains of transferable AR genes should be assured [12].
Studies reporting lactobacilli and bifidobacteria to be generally susceptible to tetracycline, erythromycin, chloramphenicol, penicillin, ampicillin, clindamycin, quinupristin-dalfopristin, linezolid, and rifampicin have been published over the last 15 years [10][13][14]. In agreement, the phenotypic analysis of the present isolates showed them to be susceptible to these antibiotics, with the exception of ampicillin and chloramphenicol—to which one and three isolates, respectively, were associated with MIC values higher than EFSA’s cut-offs [15]. In contrast, nine isolates showed resistance to one or more aminoglycosides (gentamicin, kanamycin, streptomycin, and neomycin). Resistance to aminoglycosides may occur based on several mechanisms, which include (i) enzymatic modification and inactivation of the antibiotics mediated by aminoglycoside acetyltransferases, nucleotidyltransferases, or phosphotransferases, (ii) increased efflux, (iii) decreased permeability, and (iv) modifications of the 30S ribosomal subunit interfering with the binding of this class of antibiotics [16]. However, most of the MIC values recorded in this study were just one dilution higher than the corresponding cut-off. These small MIC differences might be explained by the normal variation associated with the microdilution assay [17]. Accordingly, none of the aminoglycoside resistance genes searched for by PCR, including the widespread
aac
aph
aad(E) genes [18], were found in any of the isolates. The genome analysis further discarded the presence of acquired resistances in the sequenced strains, comprising genes and well-characterized mutations involved in aminoglycoside resistance. Given the lack of cytochrome-mediated drug transport, aminoglycoside resistance has been claimed as an intrinsic feature of LAB and other anaerobic bacteria [19]. However, large differences in the MIC values for aminoglycosides even in strains from the same species have been reported in the literature [10][13][14]. The cooperation of other non-specific mechanisms, such as increased membrane impermeability, enhanced activity of unspecific efflux pumps and multi-drug transporters, or the presence of defective cell wall autolytic systems, may further account for differences in MICs between different species and strains [20].
Resistance to the aminoglycoside streptomycin has largely been associated with mutations in chromosomal genes, for example, in
rpsL that codes for the ribosomal protein S12 [21], or in
rsmG that codes for the 16S rRNA guanine(527)-N(7)-methyltranferase (RsmG) [22]. Comparison of the deduced proteins from streptomycin-susceptible and -resistant strains of the different lactobacilli species analyzed revealed random differences between the RsmG sequences. However, none of them could be consistently associated with streptomycin resistance. In contrast, a mutation in
rpsL
B. bifidum VA07-1AN. The same amino acid replacement has been reported in other streptomycin-resistant strains of bifidobacteria [23] and many other species [3], suggesting this to be the most likely explanation for the high resistance to streptomycin shown by VA07-1AN.
Strong resistance to vancomycin is an intrinsic feature in certain
Lactobacillus
Leuconostoc spp. [24] caused by an amino acid replacement in the active site of the DdlA ligase (F261Y), as it has been experimentally demonstrated for
Leuconostoc mesenteroides [2] and
L. reuteri [25].
Although cut-offs for trimethoprim and ciprofloxacin in LAB and bifidobacteria have yet to be defined, strains of most of the present species were associated with quite high MICs. In fact, the resistance of most
Lactobacillus species to these antibiotics has been repeatedly reported [10][14]. Folate auxotrophy in lactobacilli is generally accepted as the intrinsic cause of resistance to trimethoprim [26]. Similarly, the reduced affinity of DNA gyrase (GyrA) and topoisomerase IV (ParC) variants for ciprofloxacin and other fluoroquinolones saw in some LAB species has been determined responsible for their insensitivity to this class of antibiotics [27]. Further, the presence of active multidrug efflux systems could contribute to an increase in the MIC for ciprofloxacin in some strains [7]. Since no genes coding for β-lactamases has ever been detected in LAB, non-specific mechanisms, as already discussed for the aminoglycosides, might contribute to the increased MIC of ampicillin in
L. crispatus
L. reuteri [28].
The vaginal lactobacilli in the present study were very susceptible to tetracycline—even though many LAB strains are resistant to it [9][18]. Unexpectedly, PCR analysis detected the presence of
tet
B. bifidum strains. This gene has been found to be disseminated among gut-dwelling bacteria of different species from humans and animals [29]. The genome analysis of
B. bifidum VA07-1AN showed the gene to contain an insertion of two Gs bases at its center, shifting the ORF and rendering a shorter non-functional peptide. The presence of silent tetracycline resistance genes in bifidobacteria has been reported elsewhere [30][31]. The reactivation of a silent tetracycline resistance phenotype has also been reported for
Bifidobacterium animalis
lactis Bb12 [31]. Silent AR genes could, therefore, represent a hazard, even more so when they can be easily reactivated and the restored gene remains stable afterward. Therefore, the use of strains harboring such genes in food and feed systems should be avoided.
- Munita, J.M.; Arias, C.A. Mechanisms of antibiotic resistance. Microbiol. Spectr. 2016, 4, 1–37. [Google Scholar] [CrossRef]
- Berendonk, T.U.; Manaia, C.M.; Merlin, C.; Fatta-Kassinos, D.; Cytryn, E.; Walsh, F.; Bürgmann, H.; Sørum, H.; Norström, M.; Pons, M.N.; et al. Tackling antibiotic resistance: The environmental framework. Nat. Rev. Microbiol. 2015, 13, 310–317. [Google Scholar] [CrossRef]
- Ammor, M.S.; Flórez, A.B.; Mayo, B. Antibiotic resistance in non-enterococcal lactic acid bacteria and bifidobacteria. Food Microbiol. 2007, 24, 559–570. [Google Scholar] [CrossRef]
- Duranti, S.; Lugli, G.A.; Mancabelli, L.; Turroni, F.; Milani, C.; Mangifesta, M.; Ferrario, C.; Anzalone, R.; Viappiani, A.; van Sinderen, D.; et al. Prevalence of antibiotic resistance genes among human gut-derived bifidobacteria. Appl. Environ. Microbiol. 2017, 83, e02894-16. [Google Scholar] [CrossRef]
- Štšepetova, J.; Taelma, H.; Smidt, I.; Hutt, P.; Lapp, E.; Aotäht, E.; Mändar, R. Assessment of phenotypic and genotypic antibiotic susceptibility of vaginal Lactobacillus sp. J. Appl. Microbiol. 2017, 123, 524–534. [Google Scholar] [CrossRef] [PubMed]
- Campedelli, I.; Mathur, H.; Salvetti, E.; Clarke, S.; Rea, M.C.; Torriani, S.; Ross, R.P.; Hill, C.; O’Toole, P.W. Genus-wide assessment of antibiotic resistance in Lactobacillus spp. Appl. Environ. Microbiol. 2018, 85, e01738-18. [Google Scholar] [CrossRef] [PubMed]
- Rozman, V.; Lorbeg, P.M.; Accetto, T.; Matijašić, B.B. Characterization of antimicrobial resistance in lactobacilli and bifidobacteria used as probiotics or starter cultures based on integration of phenotypic and in silico data. Int. J. Food. Microbiol. 2020, 314, 108388. [Google Scholar] [CrossRef] [PubMed]
- Salvetti, E.; O’Toole, P.W. When regulation challenges innovation: The case of the genus Lactobacillus. Trends Food Sci. Technol. 2017, 66, 187–194. [Google Scholar] [CrossRef]
- Delgado, S.; Flórez, A.B.; Mayo, B. Antibiotic susceptibility of Lactobacillus and Bifidobacterium species from the human gastrointestinal tract. Curr. Microbiol. 2005, 50, 202–207. [Google Scholar] [CrossRef] [PubMed]
- Abriouel, H.; Casado Munoz, M.D.C.; Lavilla Lerma, L.; Perez Montoro, B.; Bockelmann, W.; Pichner, R.; Kabisch, J.; Cho, G.S.; Franz, C.M.A.P.; Gálvez, A.; et al. New insights in antibiotic resistance of Lactobacillus species from fermented foods. Food Res. Int. 2015, 78, 465–481. [Google Scholar] [CrossRef]
- Doi, Y.; Wachino, J.I.; Arakawa, Y. Aminoglycoside resistance: The emergence of acquired 16S ribosomal RNA methyltransferases. Infect. Dis. Clin. N. Am. 2016, 30, 523–537. [Google Scholar] [CrossRef]
- Huys, G.; D’Haene, K.; Cnockaert, M.; Tosi, L.; Danielsen, M.; Flórez, A.B.; Mättö, J.; Axelsson, L.; Korhonen, J.; Mayrhofer, S.; et al. Intra- and interlaboratory performances of two commercial antimicrobial susceptibility testing methods for bifidobacteria and nonenterococcal lactic acid bacteria. Antimicrob. Agents Chemother. 2010, 54, 2567–2574. [Google Scholar] [CrossRef]
- Ammor, M.S.; Flórez, A.B.; van Hoek, A.H.; de Los Reyes-Gavilán, C.G.; Aarts, H.J.; Margolles, A.; Mayo, B. Molecular characterization of intrinsic and acquired antibiotic resistance in lactic acid bacteria and bifidobacteria. J. Mol. Microbiol. Biotechnol. 2008, 14, 6–15. [Google Scholar] [CrossRef]
- Condon, S. Aerobic metabolism of lactic acid bacteria. Irish J. Food Sci. Technol. 1983, 7, 15–25. [Google Scholar]
- Elkins, C.A.; Mullis, L.B. Bile-mediated aminoglycoside sensitivity in Lactobacillus species likely results from increased membrane permeability attributable to cholic acid. Appl. Environ. Microbiol. 2004, 70, 7200–7209. [Google Scholar] [CrossRef] [PubMed]
- Carter, A.P.; Clemons, W.M.; Brodersen, D.E.; Morgan-Warren, R.J.; Wimberly, B.T.; Ramakrishnan, V. Functional insights from the structure of the 30S ribosomal subunit and its interactions with antibiotics. Nature 2000, 407, 340–348. [Google Scholar] [CrossRef] [PubMed]
- Demirci, H.; Murphy, F.V.t.; Murphy, E.L.; Connetti, J.L.; Dahlberg, A.E.; Jogl, G.; Gregory, S.T. Structural analysis of base substitutions in Thermus thermophilus 16S rRNA conferring streptomycin resistance. Antimicrob. Agents Chemother. 2014, 58, 4308–4317. [Google Scholar] [CrossRef] [PubMed]
- Kiwaki, M.; Sato, T. Antimicrobial susceptibility of Bifidobacterium breve strains and genetic analysis of streptomycin resistance of probiotic B. breve strain Yakult. Int. J. Food Microbiol. 2009, 134, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, E.J.; Tyrrell, K.L.; Citron, D.M. Lactobacillus species: Taxonomic complexity and controversial susceptibilities. Clin. Infect. Dis. 2015, 60, S98–S107. [Google Scholar] [CrossRef] [PubMed]
- van Pijkeren, J.P.; Britton, R.A. High efficiency recombineering in lactic acid bacteria. Nucleic Acids Res. 2012, 40, e76. [Google Scholar] [CrossRef]
- Katla, A.K.; Kruse, H.; Johnsen, G.; Herikstad, H. Antimicrobial susceptibility of starter culture bacteria used in Norwegian dairy products. Int. J. Food Microbiol. 2001, 67, 147–152. [Google Scholar] [CrossRef]
- Castro, W.; Navarro, M.; Biot, C. Medicinal potential of ciprofloxacin and its derivatives. Future Med. Chem. 2013, 5, 81–96. [Google Scholar] [CrossRef]
- Rosander, A.; Connolly, E.; Roos, S. Removal of antibiotic resistance gene-carrying plasmids from Lactobacillus reuteri ATCC 55730 and characterization of the resulting daughter strain, L. reuteri DSM 17938. Appl. Environ. Microbiol. 2008, 74, 6032–6040. [Google Scholar] [CrossRef]
- Thaker, M.; Spanogiannopoulos, P.; Wright, G.D. The tetracycline resistome. Cell. Mol. Life Sci. 2010, 67, 419–431. [Google Scholar] [CrossRef]
- Ammor, M.S.; Flórez, A.B.; Alvarez-Martín, P.; Margolles, A.; Mayo, B. Analysis of tetracycline resistance tet(W) genes and their flanking sequences in intestinal Bifidobacterium species. J. Antimicrob. Chemother. 2008, 62, 688–693. [Google Scholar] [CrossRef] [PubMed]
- Gueimonde, M.; Flórez, A.B.; van Hoek, A.H.; Stuer-Lauridsen, B.; Strøman, P.; de los Reyes-Gavilán, C.G.; Margolles, A. Genetic basis of tetracycline resistance in Bifidobacterium animalis subsp. lactis. Appl. Environ. Microbiol. 2010, 76, 3364–3369. [Google Scholar] [CrossRef] [PubMed]
