The mode of iron (Fe) uptake and transport in cereal species is distinguished from that in other plant species by the synthesis and utilization of phytosiderophores, which are a group of Fe chelators involved in Fe mobilization from the environment as well as within the plant body. In this entry, the overview of the molecular mechanisms behind the Fe uptake and transport is presented, highlighting the commonality and diversity among cereal species.
- Iron
- Strategy I
- Startegy II
- Nicotianamine
- Phytosiderophore
- YSL
1. Fe Uptake and Transport in Cereals
In this section, we provide a concise overview of the mechanisms underlying Fe homeostasis in cereal crops, highlighting their diversity among cereal species and recent insights. Readers interested in a detailed review on the mechanisms, especially with an extensive graphical illustration, are referred to preceding articles [1][2][3][4].
1.1. Fe Uptake System and Phytosiderophore Synthesis
In aerobic soil conditions, Fe is mostly present as sparingly soluble ferric oxides, which cannot be taken up by plants as such. So as to take up sufficient Fe from the soil, graminaceous species employ a mode of Fe acquisition called Strategy II [5][6]. In Strategy-II Fe uptake, plants secrete a group of Fe(III) chelators called mugineic-acid family phytosiderophores (PS) from the roots via TRANSPORTER OF MUGENEIC ACID 1 (TOM1) [7]. The resultant Fe(III)–PS complex are taken up by the cereal plants through YELLOW STRIPE 1 (YS1) or YS1-LIKE (YSL) transporters, such as maize (Zea mays) YS1 (ZmYS1), barley (Hordeum vulgare) YS1 (HvYS1), or rice YSL15 (OsYSL15) [8][9][10][11].
PS are produced from S-adenosyl methionine (SAM; AdoMet), which originates from the methionine cycle through a sequence of enzymatic reactions [12][13][14] (Figure 1), putatively in a intracellular vesicle derived from rough endoplasmic reticulum (rER) [15][16][17][18][19][20]. Firstly, nicotianamine (NA) is synthesized via trimerization of SAM by NA SYNTHASE (NAS) [21]. Then, deoxymugineic acid (DMA) is synthesized from NA in the reactions mediated by NA AMINOTRANSFERASE (NAAT) and DMA SYNTHASE (DMAS) [22][23] (Figure 1). DMA is the only type of PS produced by rice, wheat (Triticum aestivum), and maize [24][25], whereas barley and some other graminaceous species can further convert DMA into other members of PS, some of which have stronger Fe(III)-binding capability than DMA [26][27][28][29]. The amount of PS secreted from roots also varies among graminaceous species, positively correlating with the crops’ tolerance to Fe deficiency—barley is the most tolerant crop with the largest PS secretion, whereas rice is the least tolerant with the smallest PS secretion [30].
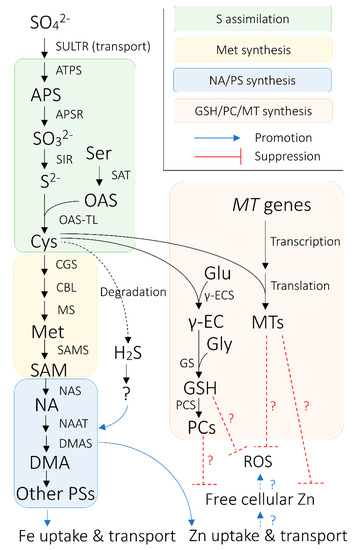
Figure 1. Overview of the connection between sulfur (S) nutrition and iron (Fe) nutrition in cereals plants. Black arrows indicate enzymatic synthetic reactions, unless otherwise indicated. Blue arrows signify promotional effects between biological processes, whereas red bars with flat ends indicate suppressive effects between biological agents and processes. Arrows and bars with dotted lines indicate relationships between biological processes/agents that are hypothetically pronounced in Fe-biofortified crops. Hydrogen sulfide (H2S) is hypothesized to regulate nicotianamine (NA)/phytosiderophores (PS) synthesis via unknown biological mechanisms or agents. Abbreviations are as follows: APS: ADENOSINE PHOSPHOSULFATE; APSR: APS REDUCTASE; ATPS: ATP SULFURYLASE; CBL: CYSTATHIONINE β-LYASE; CGS: CYSTATHIONINE γ-SYNTHASE; Cys: cysteine; DMA: deoxymugineic acid; DMAS: DMA SYNTHASE; Glu: glutamate; Gly: glycine; GS: GLUTATHIONE SYNTHETASE; GSH: glutathione; Met: methionine; MS: Met synthase; MT: METALLOTHIONEIN; NAAT: NA AMINOTRANSFERASE; NAS: NA SYNTHASE; OAS: O-acetylserine; OAS-TL: OAS (THIOL) LYASE; PC; phytochelatin; PCS: PC synthase; ROS: reactive oxygen species; S2−: sulfide; SAM: S-adenosyl methionine; SAMS: SAM SYNTHASE; Ser: serine; SO32−: sulfite; SO42−: sulfate; SULTR: SULFATE TRANSPORTER; Zn: zinc; γ-EC: glutamylcysteine; γ-ECS: γ-EC SYNTHETASE.
Relatively modest level of PS secretion from rice may be associated with the environment to which rice and its relatives have adapted [31][32]. Rice is commonly cultivated in water-logged paddy fields, where anaerobic soil conditions prevail. In deoxygenated soil, Fe is reduced to soluble Fe(II) and is fairly available for plant uptake [33]. Thus, in contrast to other cereal species [34], rice partially adopts Strategy-I Fe uptake, in which Fe(II) ions are directly taken up by IRON-REGULATED TRANSPORTERs (IRTs) [31][35]. Although a functional Strategy-II uptake system is present in rice, when Fe(II) ions are abundant in the rhizosphere, Fe acquisition is mainly achieved through Strategy-I uptake, and genes involved in Strategy-II Fe uptake are down-regulated [33][36].
1.2. Potential Diversity in the Manner of Fe Uptake and Radial Fe Transport in Roots
Among cereal species, there are also variations in root anatomy that possibly affect the site of Fe uptake in roots, as well as the fashion of subsequent within-root radial Fe transport required for xylem Fe loading.
The number of Casparian strips, which block apoplastic flow of water and nutrients across the root cell layers, is one of such anatomical variations. Almost all vascular plants have endodermis, a cell layer with a Casparian strip surrounding the root stele [37]. Moreover, the majority of the plants investigated possess exodermis, another cell layer with a Casparian strip beneath the epidermis [38]. Although rice and maize are commonly found to possess both endodermal and exodermal Casparian strips (exodermal species), wheat and barley tend to be void of exodermis (non-exodermal species), though these characteristics may vary depending on the type of roots, genotype, and stress conditions [38][39][40][41][42]. In exodermal species, nutrients should be taken up either by epidermal or exodermal cells because they are the only cell layers contacting the soil solution [37]. In non-exodermal species, on the other hand, cortical and endodermal cells can contribute to the uptake due to the absence of a barrier for the apoplastic flow of nutrients into the cortex from the external environment [37]. In line with this notion, the expression of Fe(III)-PS uptake transporter in maize, ZmYS1, is confined to epidermal cells [43].
Meanwhile, HvYS1 expression in barley can be found not only in epidermal cells but also in the cortex [8]. Furthermore, a Fe(III)-DMA transporter HvYSL2 is expressed in root endodermal cells [44]. These observations collectively suggest a potential variation in Fe uptake site in roots among different cereal species.
Subsequent to uptake from the rhizosphere, Fe should be radially transported to the root stele for xylem loading. The manner of such within-root Fe transport may also differ, especially between rice and other cereal species, owing to the difference in root anatomy. As a species well adapted to flooded conditions, rice has a highly developed aerenchyma between the endodermis and exodermis, through which air is supplied for the root cells [45]. Because only a small number of cells are alive in rice aerenchyma, there is little symplastic connection between the rice epi/exodermis and stele [46]. This distinct root internal structure in rice requires the nutrients to be apoplastically transported across the cortex before entering the stele, which is putatively mediated by a pair of efflux and influx transporters at the exodermis and endodermis, respectively [47][48]. Rice Fe transporters involved in this transport are currently unknown, but the fact that OsYSL15 is induced not only in epidermis but also in cortex upon Fe deficiency [11][49], even though rice is an exodermal species, suggests the potential involvement of OsYSL15 in radial Fe transport in roots.
1.3. Fe Mobilization for Long-Distance and Intercellular Transport
Owing to its high redox activity and low solubility, Fe has to be chelated by organic ligands for its solubilization and transport in plants [50]. The main chelators for Fe long-distance transport in cereals are considered to be citrate, NA, and PS [51][52][53][54][55][56][57][58]. In xylem Fe transport, citrate seems to be the major chelator [51][52][54][55], whereas DMA plays a supplementary role, especially under Fe deficiency (Figure 2) [52][53]. As citrate is considered to be loaded onto xylem via FERRIC REDUCTASE DEFECTIVE LIKE 1 (FRDL1) transporter in a form unbound to Fe, there can be a transporter that mediates unchelated Fe loading onto xylem, which is yet to be identified [55]. The loading of DMA and NA onto xylem may be undertaken by TOM2 and EFFLUX TRANSPORTER OF NA 1 (ENA1), respectively [59][60].
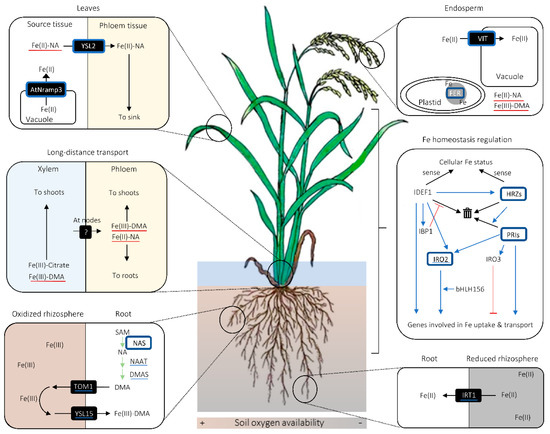
Figure 2. Graphical summary of important cereal iron (Fe) biofortification strategies. Proteins surrounded with blue squares are those whose modulation leads to increase in grain Fe concentration. Blue underlines indicate the molecules whose genes are up-regulated in rice lines overaccumulating nicotianamine (NA) or overexpressing NA SYNTHASE (NAS). Red underscores refer to the complexes whose amount is putatively increased in NAS-overexpressing lines. Black arrows indicate the movement of molecules unless otherwise noted, whereas green arrows signify synthetic enzymatic reactions. In the inset for Fe homeostasis regulation, blue arrows and red bars with flat ends indicate promotional and suppressive effects between molecular agents and/or processes, respectively. In the same inset, black arrows pointing to the bin indicate proteasomic degradation of proteins. Abbreviations are as follows: AtNRAMP3: NATURAL RESISTANCE-ASSOCIATED MACROPHAGE PROTEIN 3 from Arabidopsis thaliana; bHLH156: BASIC HELIX-LOOP-HELIX 156; DMA: deoxymugineic acid; DMAS: DMA SYNTHASE; FER: FERRITIN; HRZ: HEMERYTHRIN MOTIF-CONTAINING REALLY INTERESTING NEW GENE- AND ZINC-FINGER PROTEIN; IBP1: IDEF1-BINDING PROTEIN 1; IDEF1: IRON DEFICIENCY-RESPONSIVE ELEMENT-BINDING FACTOR 1; IRO2/3: IRON-RELATED TRANSCRIPTION FACTOR 2/3; IRT1: IRON-REGULATED TRANSPORTER 1; PRI: POSITIVE REGULATOR OF IRON DEFICIENCY RESPONSE; TOM1: TRANSPORTER OF MUGENEIC ACID 1; VIT: VACUOLAR IRON TRANSPORTER; YSL2/15: YELLOW STRIPE-1 LIKE 2/15.
In phloem, NA and/or DMA are likely to be the primary chelators for Fe (Figure 2) [51][56][61]. A Fe(II)-NA transporter YSL2, and possibly a Fe(III)-DMA transporter YSL18, contribute to Fe transport via phloem in rice [61][62][63], which suggests that Fe is primarily loaded onto phloem in a chelated form. It remains to be examined whether or not there is a phloem loading of chelator-free Fe.
Fe transferred from roots to shoots is once accumulated in stem nodes and distributed to various aboveground tissues, probably via phloem [64]. This implies that there is a mechanism mediating xylem-to-phloem Fe transfer in stem nodes (Figure 2), an organ that plays a central role in intervascular nutrient transfer and nutrient delivery to various aboveground tissues in graminaceous species [65][66]. The transporters mediating this putative xylem-to-phloem Fe transfer at stem nodes are yet to be identified. Besides being a hub for nutrient distribution, stem nodes pool nutrients in their apoplastic regions [67][68][69]. The role of citrate and citrate transporter FRDL1 in remobilizing Fe deposited in the apoplastic regions of the nodes has been described [70].
Graminaceous plants can also accumulate Fe in apoplastic spaces in roots, which can serve as a reservoir for Fe, especially under Fe-limiting conditions [71][72][73][74][75]. Phenolic compounds can contribute to solubilizing Fe precipitated in the root apoplastic regions. In rice, PHENOLICS EFFLUX ZERO1 (PEZ1) is identified as a transporter responsible for mobilizing precipitated Fe in the root xylem by pumping in protocatechuic acid (PCA) in the xylem sap [74]. In addition, PEZ2 provides phenolics to solubilize Fe in the apoplasm in the roots [75].
Route for Fe loading onto the grains may differ among cereal species as well. In wheat and barley, xylem is discontinued towards the grain [76][77]. On the other hand, there is no discontinuity of xylem towards developing rice grains [78][79]. This implies that the relative contribution of phloem-mediated source-to-sink Fe translocation to grain Fe concentration may differ among cereals.
In cereal species including rice, there is no symplastic connection between the maternal tissue and the filial tissues in grains [76][80]. Therefore, Fe must be transported though a pair of unknown efflux and influx transporters in order to be loaded onto the grain. After grain loading, Fe tends to accumulate at a higher concentration in the aleurone layer than in the endosperm [81][82]. In aleurone layer, Fe is often associated with phytic acids, which makes Fe unavailable for humans [81][83]. Fe transfer between endosperm and embryo seems to be mediated by YSL9 transporter in rice [84].
1.4. Fe Intracellular Homeostasis
Given the reactive nature of Fe, surplus Fe has to be sequestered in vacuole in plant cells [85]. On the other hand, organelles such as mitochondria and chloroplasts should be supplied with ample Fe to fulfill their physiological roles involving many redox reactions [86][87]. To meet these somewhat dilemmatic needs, Fe transport and storage in these organelles are tightly controlled. In the last decade, roles of transporters and Fe-binding agents in the regulation of intracellular Fe homeostasis have been gradually revealed in cereals, especially in rice.
MITOCHONDRIAL IRON TRANSPORTER (MIT) assumes a critical role in providing Fe for mitochondria [88]. As Fe importers for chloroplasts, Fe DEFICIENCY-RELATED 3 (FDR3) and FDR4 transporters were identified in maize [89][90]. Fe storage in plastids, particularly under sufficient or excess Fe conditions, is likely to be mediated by Fe storage protein FERRITIN (FER), whose genes encode transit peptides for plastid localization at its N-terminus [91][92][93]. In addition to FER, VACUOLAR IRON TRANSPORTERs (VITs) on the tonoplast promote compartmentalization of surplus Fe into vacuoles [94][95]. In rice, knockdown of a tonoplast-localized DMA influx transporter VACUOLAR MUGINEIC ACID TRANSPORTER (VMT) leads to higher root cell sap Fe concentration as well as lower xylem Fe concentration [96]. This implies that Fe once sequestered in vacuoles is stored as Fe(III) and can be exported again as Fe(III)-DMA. Moreover, it has been postulated that FERRIC REDUCTASE OXIDASE 1 (FRO1) localized on rice tonoplast can contribute to increasing the Fe availability for cytoplasm by reducing Fe(III) to Fe(II) in vacuoles [97]. Therefore, there may also be an unknown Fe(II) export mechanism from the vacuoles, in addition to the Fe(III)-DMA export machinery.
1.5. Fe Homeostasis Regulation
Genes involved in Fe uptake are up-regulated in response to Fe deficiency in graminaceous species, whereas they are down-regulated under Fe sufficiency [33][98][99][100][101]. An elaborate network of regulatory factors underlying such an adaptive response has been delineated recently in rice, which may be transferrable to other cereal crops. The factors that assume a pivotal role in the regulatory network in rice are IRON DEFICIENCY-RESPONSIVE ELEMENT-BINDING FACTOR 1 (IDEF1) and HEMERYTHRIN MOTIF-CONTAINING REALLY INTERESTING NEW GENE- AND ZINC-FINGER PROTEINs (HRZs), which are classified into Fe sensors by virtue of their putative capacity to alter their function to regulate Fe homeostasis by directly sensing the Fe availability in the cells [102].
IDEF1 is a positive regulator of Fe uptake-related genes, which can sense the Fe availability in cells by binding to Fe and other metal ions (Figure 2) [103][104][105]. IDEF1 interacts with a cis-element IDE1 in the promoter regions of Fe deficiency-responsive genes to up-regulate them [103][106]. As there is a diversity in amino acid sequence in the metal-binding region of IDEF1 among different graminaceous species, there may also be an inter-species functional diversity for IDEF1 [105]. IDEF1 gene is expressed irrespective of the Fe status of the plant, but its protein is prone to 26S proteasome-dependent degradation under Fe-sufficient conditions (Figure 2) [107][108]. In Fe-deficient conditions, its degradation is prevented by IDEF1-BINDING PROTEINs (IBPs), and as a result, genes involved in Fe deficiency response are induced (Figure 2) [107]. IBP genes have many IDE1 motifs in their promoter regions and are positively regulated by IDEF1, thereby constituting a positive feedback loop between IDEF1 and IBP1 for Fe deficiency response (Figure 2) [107]. In rice, IDEF1 is known to positively regulate another transcription factor governing Fe deficiency response, called IRON-RELATED TRANSCRIPTION FACTOR 2 (IRO2) (Figure 2) [103][109]. IRO2 can induce genes involved in Strategy-II Fe uptake upon Fe deficiency by binding to their promoter regions [110]. It has been recently revealed that IRO2 requires another transcription factor BASIC HELIX-LOOP-HELIX 156 (bHLH156) to localize in the nucleus and regulate gene expression (Figure 2) [111].
In contrast to IDEF1, HRZ1 and HRZ2 are identified as negative regulators of Strategy-II Fe uptake in rice, especially when there is sufficient external Fe available [112]. Supposedly, HRZs sense Fe availability also by binding with Fe and other metal ions (Figure 2) [112]. Like IDEF1, HRZs are as well susceptible to 26S proteasome-dependent degradation in roots, but contrary to IDEF1, they are so regardless of the Fe status (Figure 2) [112]. HRZs negatively regulate Fe uptake and translocation by contributing to the proteasomic degradation of POSITIVE REGULATORS OF IRON HOMEOSTASIS (PRIs) through their ubiquitination (Figure 2) [113][114]. PRIs positively regulate the expression of IRO2 and also YSL2, which is not regulated by IRO2 [114][115].
Interestingly, HRZs are positively regulated by IDEF1 (Figure 2) [112]. Moreover, PRIs, which are deemed overall as positive regulators of Fe uptake and translocation, induce the expression of IRO3, which codes for a negative transcription factor for Fe deficiency response in rice (Figure 2) [113][114][116]. These interrelations between antagonistic regulatory factors could be part of a sophisticated machinery to prevent excessive Fe deficiency response.
There are several other characterized regulatory factors involved in Fe homeostasis regulation in rice, whose association with the abovementioned regulation network, with IDEF1 and HRZs at its core, is absent or unknown. In parallel to IDE1 and IDEF1, a pair of cis-element and trans-factor, namely, IDE2 and IDEF2, were found to positively regulate Fe deficiency response [106][117]. Moreover, rice bHLH133 is a transcription factor that contributes to Fe retention in roots under Fe-deficient conditions [118]. Rice homologues of IRON MANs (IMAs), which are regulatory factors identified in Arabidopsis (Arabidopsis thaliana) as a positive regulator of systemic Fe deficiency signaling, are also likely to have similar roles in rice [119].
One rare exception of a transcription factor related to Fe transport identified first in wheat is NO APICAL MERISTEM B-1 (NAM B-1) [120]. Wheat NAM B-1 promotes the senescence of vegetative tissues, which leads to increased translocation of Fe from leaves to grains along with other nutrients [120]. Although NAM B-1 homologue is identified in rice, it does not have an identical role as the wheat counterpart [121].
In addition to transcription factors, signal transduction through phytohormones such as auxin or jasmonates are suggested as being important components of Fe deficiency response [122][123]. NA is also speculated as being a possible Fe deficiency signaling molecule [19][102][124], given that the genes involved in Fe acquisition and translocation are up-regulated in rice lines accumulating increased NA [18][125][126].
References
- Yuta Kawakami; Navreet K. Bhullar; Molecular processes in iron and zinc homeostasis and their modulation for biofortification in rice.. Journal of Integrative Plant Biology 2018, 60, 1181-1198, 10.1111/jipb.12751.
- Takanori Kobayashi; Naoko K. Nishizawa; Iron Uptake, Translocation, and Regulation in Higher Plants. Annual Review of Plant Biology 2012, 63, 131-152, 10.1146/annurev-arplant-042811-105522.
- Khurram Bashir; Tomoko Nozoye; Yasuhiro Ishimaru; Hiromi Nakanishi; Naoko K. Nishizawa; Exploiting new tools for iron bio-fortification of rice. Biotechnology Advances 2013, 31, 1624-1633, 10.1016/j.biotechadv.2013.08.012.
- James Connorton; Janneke Balk; Jorge Rodriguez-Celma; Iron homeostasis in plants – a brief overview. Metallomics 2017, 9, 813-823, 10.1039/c7mt00136c.
- Volker Römheld; Horst Marschner; Evidence for a Specific Uptake System for Iron Phytosiderophores in Roots of Grasses 1. Plant Physiology 1986, 80, 175-180.
- Sei-Ichi Takagi; Naturally occurring iron-chelating compounds in oat- and rice-root washings. Soil Science and Plant Nutrition 1976, 22, 423-433, 10.1080/00380768.1976.10433004.
- Tomoko Nozoye; Seiji Nagasaka; Takanori Kobayashi; Michiko Takahashi; Yuki Sato; Yoko Sato; Nobuyuki Uozumi; Hiromi Nakanishi; Naoko K. Nishizawa; Phytosiderophore Efflux Transporters Are Crucial for Iron Acquisition in Graminaceous Plants. Journal of Biological Chemistry 2010, 286, 5446-5454, 10.1074/jbc.M110.180026.
- Murata, Y.; Ma, J.F.; Yamaji, N.; Ueno, D.; Nomoto, K.; Iwashita, T; A specific transporter for iron(III)-phytosiderophore in barley roots. Plant J. 2006, 46, 563–572.
- Catherine Curie; Zivile Panaviene; Clarisse Loulergue; Stephen L. Dellaporta; Jean-Francois Briat; Elsbeth L. Walker; Maize yellow stripe1 encodes a membrane protein directly involved in Fe(III) uptake. Nature 2001, 409, 346-349, 10.1038/35053080.
- Inoue, H.; Kobayashi, T.; Nozoye, T.; Takahashi, M.; Kakei, Y.; Suzuki, K.; Nakazono, M.; Nakanishi, H.; Mori, S.; Nishizawa, N.K. Rice OsYSL15 is an iron-regulated iron (III)-deoxymugineic acid transporter expressed in the roots and is essential for iron uptake in early growth of the seedlings. J. Biol. Chem. 2009, 284, 3470–3479.
- Sichul Lee; Jeff C. Chiecko; Sun A. Kim; Elsbeth L. Walker; Youngsook Lee; Mary Lou Guerinot; Gynheung An; Disruption of OsYSL15 Leads to Iron Inefficiency in Rice Plants1[C][W][OA]. Plant Physiology 2009, 150, 786-800, 10.1104/pp.109.135418.
- Satoshi Mori; Naoko Nishizawa; Methionine as a Dominant Precursor of Phytosiderophores in Graminaceae Plants. Plant And Cell Physiology 1987, 28, 1081–1092, 10.1093/oxfordjournals.pcp.a077388.
- Shojima, S.; Nishizawa, N.-K.; Fushiya, S.; Nozoe, S.; Irifune, T.; Mori, S; Biosynthesis of phytosiderophores. Plant Physiol. 1990, 93, 1497–1503.
- J. F. Ma; Tetsuro Shinada; Chitose Matsuda; Kyosuke Nomoto; Biosynthesis of Phytosiderophores, Mugineic Acids, Associated with Methionine Cycling. Journal of Biological Chemistry 1995, 270, 16549-16554, 10.1074/jbc.270.28.16549.
- N. Nishizawa; S. Mori; The particular vesicle appearing in barley root cells and its relation to mugineic acid secretion. Journal of Plant Nutrition 1987, 10, 1013-1020, 10.1080/01904168709363629.
- Takashi Negishi; Hiromi Nakanishi; Junshi Yazaki; Naoki Kishimoto; Fumiko Fujii; Kanako Shimbo; Kimiko Yamamoto; Katsumi Sakata; Takuji Sasaki; Shoshi Kikuchi; Satoshi Mori; Naoko K. Nishizawa; cDNA microarray analysis of gene expression during Fe-deficiency stress in barley suggests that polar transport of vesicles is implicated in phytosiderophore secretion in Fe-deficient barley roots.. The Plant Journal 2002, 30, 83-94, 10.1046/j.1365-313x.2002.01270.x.
- Daichi Mizuno; Kyoko Higuchi; Tatsuya Sakamoto; Hiromi Nakanishi; Satoshi Mori; Naoko K. Nishizawa; Three Nicotianamine Synthase Genes Isolated from Maize Are Differentially Regulated by Iron Nutritional Status. Plant Physiology 2003, 132, 1989-1997, 10.1104/pp.102.019869.
- Nozoye, T.; Nagasaka, S.; Bashir, K.; Takahashi, M.; Kobayashi, T.; Nakanishi, H.; Nishizawa, N.K. Nicotianamine synthase 2 localizes to the vesicles of iron-deficient rice roots, and its mutation in the YXXφ or LL motif causes the disruption of vesicle formation or movement in rice. Plant J. 2014, 77, 246–260.
- Tomoko Nozoye; Kyoko Tsunoda; Seiji Nagasaka; Khurram Bashir; Michiko Takahashi; Takanori Kobayashi; Hiromi Nakanishi; Naoko K. Nishizawa; Rice nicotianamine synthase localizes to particular vesicles for proper function. Plant Signaling & Behavior 2014, 9, e28660.
- Luqing Zheng; Miho Fujii; Naoki Yamaji; Akimasa Sasaki; Miki Yamane; Isamu Sakurai; Kazuhiro Sato; Jian Feng Ma; Isolation and Characterization of a Barley Yellow Stripe-Like Gene, HvYSL5. Plant And Cell Physiology 2011, 52, 765-774, 10.1093/pcp/pcr009.
- K. Higuchi; Cloning of Nicotianamine Synthase Genes, Novel Genes Involved in the Biosynthesis of Phytosiderophores. Plant Physiology 1999, 119, 471-480, 10.1104/pp.119.2.471.
- M. Takahashi; Cloning Two Genes for Nicotianamine Aminotransferase, a Critical Enzyme in Iron Acquisition (Strategy II) in Graminaceous Plants. Plant Physiology 1999, 121, 947-956, 10.1104/pp.121.3.947.
- Khurram Bashir; Haruhiko Inoue; Seiji Nagasaka; Michiko Takahashi; Hiromi Nakanishi; Satoshi Mori; Naoko K. Nishizawa; Cloning and Characterization of Deoxymugineic Acid Synthase Genes from Graminaceous Plants. Journal of Biological Chemistry 2006, 281, 32395-32402, 10.1074/jbc.m604133200.
- Shigenao Kawai; Sei‐Ichi Takagi; Yoshimasa Sato; Mugineic acid?family phytosiderophores in root?secretions of barley, corn and sorghum varieties. Journal of Plant Nutrition 1988, 11, 633-642, 10.1080/01904168809363829.
- Jian Feng Ma; Shin Taketa; Yi-Chieh Chang; Kazuyoshi Takeda; Hideaki Matsumoto; Biosynthesis of phytosiderophores in several Triticeae species with different genomes. Journal of Experimental Botany 1999, 50, 723-726, 10.1093/jxb/50.334.723.
- Hiromi Nakanishi; Hirotaka Yamaguchi; Tetsuo Sasakuma; Naoko K. Nishizawa; Satoshi Mori; Two dioxygenase genes, Ids3 and Ids2, from Hordeum vulgare are involved in the biosynthesis of mugineic acid family phytosiderophores.. Plant Molecular Biology 2000, 44, 199-207, 10.1023/a:1006491521586.
- von Wirén, N.; Khodr, H.; Hider, R.C. Hydroxylated phytosiderophore species possess an enhanced chelate stability and affinity for Iron(III). Plant Physiol. 2002, 124, 1149–1158.
- D. Ueno; A. D. Rombolà; T. Iwashita; K. Nomoto; J. F. Ma; Identification of two novel phytosiderophores secreted by perennial grasses. New Phytologist 2007, 174, 304-310, 10.1111/j.1469-8137.2007.02056.x.
- Tomoko Nozoye; May Sann Aung; Hiroshi Masuda; Hiromi Nakanishi; Naoko K. Nishizawa; Bioenergy grass [Erianthus ravennae(L.) Beauv.] secretes two members of mugineic acid family phytosiderophores which involved in their tolerance to Fe deficiency. Soil Science and Plant Nutrition 2017, 63, 543-552, 10.1080/00380768.2017.1394168.
- Takagi, S.-I. Production of phytosiderophores. In Iron Chelation in Plants and Soil Microorganisms; Barton, L., Hemming, B.C., Eds.; Academic Press: San Diego, CA, USA, 1993; pp. 111–131.
- Yasuhiro Ishimaru; Motofumi Suzuki; Takashi Tsukamoto; Kazumasa Suzuki; Mikio Nakazono; Takanori Kobayashi; Yasuaki Wada; Satoshi Watanabe; Shinpei Matsuhashi; Michiko Takahashi; Hiromi Nakanishi; Satoshi Mori; Naoko K. Nishizawa; Rice plants take up iron as an Fe3+-phytosiderophore and as Fe2+. The Plant Journal 2006, 45, 335-346, 10.1111/j.1365-313x.2005.02624.x.
- Wairich, A.; de Oliveira, B.H.N.; Arend, E.B.; Duarte, G.L.; Ponte, L.R.; Sperotto, R.A.; Ricachenevsky, F.K.; Fett, J.P. The Combined Strategy for iron uptake is not exclusive to domesticated rice (Oryza sativa). Sci. Rep. 2019, 9, 1–17.
- Peitong Wang; Naoki Yamaji; Komaki Inoue; Keiich Mochida; Jian Feng Ma; Plastic transport systems of rice for mineral elements in response to diverse soil environmental changes. New Phytologist 2019, 226, 156-169, 10.1111/nph.16335.
- Tatiana Zaharieva; Volker Römheld; Specific Fe2+uptake system in strategy I plants inducible under Fe deficiency. Journal of Plant Nutrition 2000, 23, 1733-1744, 10.1080/01904160009382137.
- Naimatullah Bughio; Hirotaka Yamaguchi; Naoko K. Nishizawa; Hiromi Nakanishi; Satoshi Mori; Cloning an iron-regulated metal transporter from rice.. Journal of Experimental Botany 2002, 53, 1677-1682, 10.1093/jxb/erf004.
- Chengshuai Liu; Ting Gao; Yuhui Liu; Jingyu Liu; Fangbai Li; Zhenwu Chen; YongZhu Li; Yiwen Lv; Zhiyi Song; John R. Reinfelder; Weilin Huang; Isotopic fingerprints indicate distinct strategies of Fe uptake in rice. Chemical Geology 2019, 524, 323-328, 10.1016/j.chemgeo.2019.07.002.
- Daryl E. Enstone; Carol A. Peterson; Fengshan Ma; Root Endodermis and Exodermis: Structure, Function, and Responses to the Environment. Journal of Plant Growth Regulation 2002, 21, 335-351, 10.1007/s00344-003-0002-2.
- Christopher J. Perumalla; Jerry G. Chmielewski; Carol A. Peterson; A survey of angiosperm species to detect hypodermal Casparian bands. III. Rhizomes. Botanical Journal of the Linnean Society 1990, 103, 127-132, 10.1111/j.1095-8339.1990.tb00178.x.
- Carol A. Peterson; Mary E. Emanuel; Christine Wilson; Identification of a Casparian band in the hypodermis of onion and corn roots. Canadian Journal of Botany 1982, 60, 1529-1535, 10.1139/b82-195.
- Clark, L.H.; Harris, W.H. Observations on the root anatomy of rice (Oryza sativa L.). Am. J. Bot. 1981, 68, 154–161.
- Edita Tylová; Eva Pecková; Zuzana Blascheová; Aleš Soukup; Casparian bands and suberin lamellae in exodermis of lateral roots: an important trait of roots system response to abiotic stress factors. Annals of Botany 2017, 120, 71-85, 10.1093/aob/mcx047.
- Tino Kreszies; Stella Eggels; Victoria Kreszies; Alina Osthoff; Nandhini Shellakkutti; Jutta A. Baldauf; Viktoria V. Zeisler‐Diehl; Frank Hochholdinger; Kosala Ranathunge; Lukas Schreiber; et al. Seminal roots of wild and cultivated barley differentially respond to osmotic stress in gene expression, suberization, and hydraulic conductivity. Plant, Cell & Environment 2020, 43, 344-357, 10.1111/pce.13675.
- Daisei Ueno; Naoki Yamaji; Jian Feng Ma; Further characterization of ferric-phytosiderophore transporters ZmYS1 and HvYS1 in maize and barley.. Journal of Experimental Botany 2009, 60, 3513-20, 10.1093/jxb/erp191.
- Ryoichi Araki; Jun Murata; Yoshiko Murata; A Novel Barley Yellow Stripe 1-Like Transporter (HvYSL2) Localized to the Root Endodermis Transports Metal-Phytosiderophore Complexes. Plant And Cell Physiology 2011, 52, 1931-1940, 10.1093/pcp/pcr126.
- M. B. Jackson; W. Armstrong; Formation of Aerenchyma and the Processes of Plant Ventilation in Relation to Soil Flooding and Submergence. Plant Biology 1999, 1, 274-287, 10.1055/s-2007-978516.
- Maki Kawai-Yamada; P. K. Samarajeewa; R. A. Barrero; M. Nishiguchi; H. Uchimiya; Cellular dissection of the degradation pattern of cortical cell death during aerenchyma formation of rice roots. Planta 1998, 204, 277-287, 10.1007/s004250050257.
- Akimasa Sasaki; Naoki Yamaji; Jian Feng Ma; Transporters involved in mineral nutrient uptake in rice. Journal of Experimental Botany 2016, 67, 3645-3653, 10.1093/jxb/erw060.
- Jing Che; Naoki Yamaji; Jian Feng Ma; Efficient and flexible uptake system for mineral elements in plants. New Phytologist 2018, 219, 513-517, 10.1111/nph.15140.
- Yuko Ogo; Yusuke Kakei; Reiko Nakanishi Itai; Takanori Kobayashi; Hiromi Nakanishi; Hirokazu Takahashi; Mikio Nakazono; Naoko K. Nishizawa; Spatial transcriptomes of iron-deficient and cadmium-stressed rice. New Phytologist 2014, 201, 781-794, 10.1111/nph.12577.
- Ruediger Hell; U. W. Stephan; Iron uptake, trafficking and homeostasis in plants. Planta 2003, 216, 541-551, 10.1007/s00425-002-0920-4.
- Nicolaus Von Wirén; Sukhbinder Klair; Suhkibar Bansal; Jean-Francois Briat; Hicham Khodr; Takayuki Shioiri; Roger A. Leigh; Robert C. Hider; Nicotianamine Chelates Both FeIII and FeII. Implications for Metal Transport in Plants1. Plant Physiology 1999, 119, 1107-1114.
- Tomoko Ariga; Kenji Hazama; Shuichi Yanagisawa; Tadakatsu Yoneyama; Chemical forms of iron in xylem sap from graminaceous and non-graminaceous plants. Soil Science and Plant Nutrition 2014, 60, 460-469, 10.1080/00380768.2014.922406.
- Yusuke Kakei; Isomaro Yamaguchi; Takanori Kobayashi; Michiko Takahashi; Hiromi Nakanishi; Takashi Yamakawa; Naoko K. Nishizawa; A highly sensitive, quick and simple quantification method for nicotianamine and 2'-deoxymugineic acid from minimum samples using LC/ESI-TOF-MS achieves functional analysis of these components in plants. Plant And Cell Physiology 2009, 50, 1988-93, 10.1093/pcp/pcp141.
- Rubén Rellán-Álvarez; Anunciación Abadía; Ana Álvarez-Fernández; Formation of metal-nicotianamine complexes as affected by pH, ligand exchange with citrate and metal exchange. A study by electrospray ionization time-of-flight mass spectrometry. Rapid Communications in Mass Spectrometry 2008, 22, 1553-1562, 10.1002/rcm.3523.
- Kengo Yokosho; Naoki Yamaji; Daisei Ueno; Namiki Mitani; Jian Feng Ma; OsFRDL1 Is a Citrate Transporter Required for Efficient Translocation of Iron in Rice1[OA]. Plant Physiology 2009, 149, 297-305, 10.1104/pp.108.128132.
- Nishiyama, R.; Kato, M.; Nagata, S.; Yanagisawa, S.; Yoneyama, T. Identification of Zn-nicotianamine and Fe-2’-deoxymugineic acid in the phloem sap from rice plants (Oryza sativa L.). Plant Cell Physiol. 2012, 53, 381–390.
- Shah Alam; Shigeru Kamei; Shigenao Kawai; Effect of iron deficiency on the chemical composition of the xylem sap of barley. Soil Science and Plant Nutrition 2001, 47, 643-649, 10.1080/00380768.2001.10408428.
- Mori, S.; Nishizawa, N.; Hayashi, H.; Chino, M.; Yoshimura, E.; Ishihara, J. Why are young rice plants highly susceptible to iron deficiency? Plant Soil 1991, 130, 143–156.
- Tomoko Nozoye; Seiji Nagasaka; Takanori Kobayashi; Yuki Sato; Nobuyuki Uozumi; Hiromi Nakanishi; Naoko K. Nishizawa; The Phytosiderophore Efflux Transporter TOM2 Is Involved in Metal Transport in Rice*. Journal of Biological Chemistry 2015, 290, 27688-27699, 10.1074/jbc.M114.635193.
- Tomoko Nozoye; Nicolaus Von Wirén; Yoshikatsu Sato; Tetsuya Higashiyama; Hiromi Nakanishi; Naoko K. Nishizawa; Characterization of the Nicotianamine Exporter ENA1 in Rice. Frontiers in Plant Science 2019, 10, 502, 10.3389/fpls.2019.00502.
- Shintaro Koike; Haruhiko Inoue; Daichi Mizuno; Michiko Takahashi; Hiromi Nakanishi; Satoshi Mori; Naoko K. Nishizawa; OsYSL2 is a rice metal-nicotianamine transporter that is regulated by iron and expressed in the phloem. The Plant Journal 2004, 39, 415-424, 10.1111/j.1365-313x.2004.02146.x.
- Yasuhiro Ishimaru; Hiroshi Masuda; Khurram Bashir; Haruhiko Inoue; Takashi Tsukamoto; Michiko Takahashi; Hiromi Nakanishi; N Aoki; Tatsuro Hirose; Ryu Ohsugi; Naoko K. Nishizawa; Rice metal-nicotianamine transporter, OsYSL2, is required for the long-distance transport of iron and manganese. The Plant Journal 2010, 62, 379-390, 10.1111/j.1365-313x.2010.04158.x.
- Aoyama, T.; Kobayashi, T.; Takahashi, M.; Nagasaka, S.; Usuda, K.; Kakei, Y.; Ishimaru, Y.; Nakanishi, H.; Mori, S.; Nishizawa, N.K. OsYSL18 is a rice iron(III)-deoxymugineic acid transporter specifically expressed in reproductive organs and phloem of lamina joints. Plant Mol. Biol. 2009, 70, 681–692.
- Tsukamoto, T.; Nakanishi, H.; Uchida, H.; Watanabe, S.; Matsuhashi, S.; Mori, S.; Nishizawa, N.K. 52Fe translocation in barley as monitored by a positron-emitting tracer imaging system (PETIS): Evidence for the direct translocation of Fe from roots to young leaves via phloem. Plant Cell Physiol. 2009, 50, 48–57.
- Naoki Yamaji; Jian Feng Ma; The node, a hub for mineral nutrient distribution in graminaceous plants. Trends in Plant Science 2014, 19, 556-563, 10.1016/j.tplants.2014.05.007.
- Naoki Yamaji; Jian Feng Ma; Node-controlled allocation of mineral elements in Poaceae. Current Opinion in Plant Biology 2017, 39, 18-24, 10.1016/j.pbi.2017.05.002.
- Noriko Yamaguchi; Satoru Ishikawa; Tadashi Abe; Koji Baba; Tomohito Arao; Yasuko Terada; Role of the node in controlling traffic of cadmium, zinc, and manganese in rice. Journal of Experimental Botany 2012, 63, 2729-37, 10.1093/jxb/err455.
- Katie Moore; Yi Chen; Allison M. L. Van De Meene; Louise Hughes; Wenju Liu; Tina Geraki; Fred Mosselmans; Stephen McGrath; Chris Grovenor; Fang-Jie Zhao; et al. Combined NanoSIMS and synchrotron X-ray fluorescence reveal distinct cellular and subcellular distribution patterns of trace elements in rice tissues. New Phytologist 2014, 201, 104-115, 10.1111/nph.12497.
- Naoki Yamaji; Jian Feng Ma; Bioimaging of multiple elements by high-resolution LA-ICP-MS reveals altered distribution of mineral elements in the nodes of rice mutants. The Plant Journal 2019, 99, 1254-1263, 10.1111/tpj.14410.
- Kengo Yokosho; Naoki Yamaji; Jian Feng Ma; OsFRDL1 expressed in nodes is required for distribution of iron to grains in rice. Journal of Experimental Botany 2016, 67, 5485-5494, 10.1093/jxb/erw314.
- Bienfait, H.F.; van den Briel, W.; Mesland-Mul, N.T. Free space iron pools in roots: Generation and mobilization. Plant Physiol. 1985, 78, 596–600.
- Fu-Suo Zhang; Volker Römheld; Horst Marschner; Role of the Root Apoplasm for Iron Acquisition by Wheat Plants. Plant Physiology 1991, 97, 1302-1305, 10.1104/pp.97.4.1302.
- Rongli Shi; Michael Melzer; Shao Jian Zheng; Andreas Benke; Benjamin Stich; Nicolaus Von Wirén; Iron Retention in Root Hemicelluloses Causes Genotypic Variability in the Tolerance to Iron Deficiency-Induced Chlorosis in Maize. Frontiers in Plant Science 2018, 9, 557, 10.3389/fpls.2018.00557.
- Yasuhiro Ishimaru; Yusuke Kakei; Hugo Shimo; Khurram Bashir; Yutaka Sato; Yuki Sato; Nobuyuki Uozumi; Hiromi Nakanishi; Naoko K. Nishizawa; A Rice Phenolic Efflux Transporter Is Essential for Solubilizing Precipitated Apoplasmic Iron in the Plant Stele*. Journal of Biological Chemistry 2011, 286, 24649-24655, 10.1074/jbc.M111.221168.
- Khurram Bashir; Yasuhiro Ishimaru; Hugo Shimo; Yusuke Kakei; Takeshi Senoura; Ryuichi Takahashi; Yutaka Sato; Yuki Sato; Nobuyuki Uozumi; Hiromi Nakanishi; Naoko K. Nishizawa; Rice phenolics efflux transporter 2 (PEZ2) plays an important role in solubilizing apoplasmic iron. Soil Science and Plant Nutrition 2011, 57, 803-812, 10.1080/00380768.2011.637305.
- Palmgren, M.G.; Clemens, S.; Williams, L.E.; Krämer, U.; Borg, S.; Schjørring, J.K.; Sanders, D; Zinc biofortification of cereals: Problems and solutions. Trends Plant Sci. 2008, 13, 464–473.
- Sy Zee; Tp O'brien; A Special Type of Tracheary Element Associated with "Xylem Discontinuity" in the Floral Axis of Wheat. Australian Journal of Biological Sciences 1970, 23, 783, 10.1071/bi9700783.
- Tjeerd‐Jan Stomph; Wen Jiang; Paul C. Struik; Zinc biofortification of cereals: rice differs from wheat and barley. Trends in Plant Science 2009, 14, 123-4, 10.1016/j.tplants.2009.01.001.
- S-Y Zee; Vascular Tissue and Transfer Cell Distribution in the Rice Spikelet. Australian Journal of Biological Sciences 1972, 25, 411, 10.1071/bi9720411.
- Xiaoba Wu; Jinxin Liu; Dongqi Li; Chun-Ming Liu; Rice caryopsis development I: Dynamic changes in different cell layers. Journal of Integrative Plant Biology 2016, 58, 772-785, 10.1111/jipb.12440.
- Jesse T. Beasley; Julien P. Bonneau; Jose T. Sánchez-Palacios; Laura T. Moreno-Moyano; Damien Callahan; Elad Tako; Raymond P. Glahn; Enzo Lombi; Alexander A.T. Johnson; Metabolic engineering of bread wheat improves grain iron concentration and bioavailability. Plant Biotechnology Journal 2019, 17, 1514-1526, 10.1111/pbi.13074.
- Bianca Kyriacou; Katie Moore; David Paterson; Martin D. De Jonge; Daryl L. Howard; James Stangoulis; Mark Tester; Enzo Lombi; Alexander A.T. Johnson; Localization of iron in rice grain using synchrotron X-ray fluorescence microscopy and high resolution secondary ion mass spectrometry. Journal of Cereal Science 2014, 59, 173-180, 10.1016/j.jcs.2013.12.006.
- Alexander A.T. Johnson; Bianca Kyriacou; Damien Callahan; Lorraine Carruthers; James Stangoulis; Enzo Lombi; Mark Tester; Constitutive Overexpression of the OsNAS Gene Family Reveals Single-Gene Strategies for Effective Iron- and Zinc-Biofortification of Rice Endosperm. PLOS ONE 2011, 6, e24476, 10.1371/journal.pone.0024476.
- Takeshi Senoura; Emi Sakashita; Takanori Kobayashi; Michiko Takahashi; May Sann Aung; Hiroshi Masuda; Hiromi Nakanishi; Naoko K. Nishizawa; The iron-chelate transporter OsYSL9 plays a role in iron distribution in developing rice grains. Plant Molecular Biology 2017, 95, 375-387, 10.1007/s11103-017-0656-y.
- Sébastien Thomine; Grégory Vert; Grégory Vert; Iron transport in plants: better be safe than sorry. Current Opinion in Plant Biology 2013, 16, 322-327, 10.1016/j.pbi.2013.01.003.
- Cecile Nouet; Patrick Motte; Marc Hanikenne; Chloroplastic and mitochondrial metal homeostasis. Trends in Plant Science 2011, 16, 395-404, 10.1016/j.tplants.2011.03.005.
- Gianpiero Vigani; Ï¿½D�M Solti; S�Bastien Thomine; Katrin Philippar; Ádám Solti; Sébastien Thomine; Essential and Detrimental — an Update on Intracellular Iron Trafficking and Homeostasis. Plant And Cell Physiology 2019, 60, 1420-1439, 10.1093/pcp/pcz091.
- Khurram Bashir; Yasuhiro Ishimaru; Hugo Shimo; Seiji Nagasaka; Masaru Fujimoto; Hideki Takanashi; Nobuhiro Tsutsumi; Gynheung An; Hiromi Nakanishi; Naoko K. Nishizawa; The rice mitochondrial iron transporter is essential for plant growth. Nature Communications 2011, 2, 322, 10.1038/ncomms1326.
- Jianhui Han; Xiufang Song; Peng Li; Huijun Yang; Liping Yin; Maize ZmFDR3 localized in chloroplasts is involved in iron transport. Science in China Series C: Life Sciences 2009, 52, 864-871, 10.1007/s11427-009-0108-2.
- Xiu-Yue Zhang; Xi Zhang; Qi Zhang; Xiao-Xi Pan; Luo-Chen Yan; Xiao-Juan Ma; Wei-Zhong Zhao; Xiao-Ting Qi; Li-Ping Yin; Zea maysFe deficiency-related 4 (ZmFDR4) functions as an iron transporter in the plastids of monocots. The Plant Journal 2017, 90, 147-163, 10.1111/tpj.13482.
- Isabelle Fobis-Loisy; Karine Loridon; Stéphane Lobréaux; Michel Lebrun; Jean-François Briat; Structure and Differential Expression of two Maize Ferritin Genes in Response to Iron and Abscisic Acid. JBIC Journal of Biological Inorganic Chemistry 1995, 231, 609-619, 10.1111/j.1432-1033.1995.tb20739.x.
- Stein, R.J.; Ricachenevsky, F.K.; Fett, J.P. Differential regulation of the two rice ferritin genes (OsFER1 and OsFER2). Plant Sci. 2009, 177, 563–569.
- Søren Borg; Henrik Brinch-Pedersen; Birgitte Tauris; Lene Heegaard Madsen; Behrooz Darbani; Shahin Noeparvar; Preben Bach Holm; Wheat ferritins: Improving the iron content of the wheat grain. Journal of Cereal Science 2012, 56, 204-213, 10.1016/j.jcs.2012.03.005.
- Yu Zhang; Yong-Han Xu; Hong-Yin Yi; Ji-Ming Gong; Vacuolar membrane transporters OsVIT1 and OsVIT2 modulate iron translocation between flag leaves and seeds in rice. The Plant Journal 2012, 72, 400-410, 10.1111/j.1365-313x.2012.05088.x.
- James Connorton; Eleanor R. Jones; Ildefonso Rodríguez-Ramiro; Susan Fairweather-Tait; Cristobal Uauy; Janneke Balk; Wheat Vacuolar Iron Transporter TaVIT2 Transports Fe and Mn and Is Effective for Biofortification. Plant Physiology 2017, 174, 2434-2444, 10.1104/pp.17.00672.
- Jing Che; Kengo Yokosho; Naoki Yamaji; Jian Feng Ma; A Vacuolar Phytosiderophore Transporter Alters Iron and Zinc Accumulation in Polished Rice Grains. Plant Physiology 2019, 181, 276-288, 10.1104/pp.19.00598.
- Li, L.; Ye, L.; Kong, Q.; Shou, H. A vacuolar membrane ferric-chelate reductase, OsFRO1, alleviates Fe toxicity in rice (Oryza sativa L.). Front. Plant Sci. 2019, 10, 1–11.
- Takanori Kobayashi; Motofumi Suzuki; Haruhiko Inoue; Reiko Nakanishi Itai; Michiko Takahashi; Hiromi Nakanishi; Satoshi Mori; Naoko K. Nishizawa; Expression of iron-acquisition-related genes in iron-deficient rice is co-ordinately induced by partially conserved iron-deficiency-responsive elements. Journal of Experimental Botany 2005, 56, 1305-1316, 10.1093/jxb/eri131.
- Yan Li; Nian Wang; Fengtao Zhao; Xuejiao Song; Zhaohua Yin; Rong Huang; Chunqing Zhang; Changes in the transcriptomic profiles of maize roots in response to iron-deficiency stress. Plant Molecular Biology 2014, 85, 349-363, 10.1007/s11103-014-0189-6.
- Meng Wang; Yuta Kawakami; Navreet K. Bhullar; Molecular Analysis of Iron Deficiency Response in Hexaploid Wheat. Frontiers in Sustainable Food Systems 2019, 3, 67, 10.3389/fsufs.2019.00067.
- Seiji Nagasaka; Michiko Takahashi; Reiko Nakanishi-Itai; Khurram Bashir; Hiromi Nakanishi; Satoshi Mori; Naoko K. Nishizawa; Time course analysis of gene expression over 24 hours in Fe-deficient barley roots. Plant Molecular Biology 2008, 69, 621-631, 10.1007/s11103-008-9443-0.
- Takanori Kobayashi; Naoko K. Nishizawa; Iron sensors and signals in response to iron deficiency. Plant Science 2014, 224, 36-43, 10.1016/j.plantsci.2014.04.002.
- Kobayashi, T.; Ogo, Y.; Itai, R.N.; Nakanishi, H.; Takahashi, M.; Mori, S.; Nishizawa, N.K; The transcription factor IDEF1 regulates the response to and tolerance of iron deficiency in plants. Proc. Natl. Acad. Sci. USA 2007, 104, 19150–19155.
- Takanori Kobayashi; Reiko Nakanishi Itai; Yuko Ogo; Yusuke Kakei; Hiromi Nakanishi; Michiko Takahashi; Naoko K. Nishizawa; The rice transcription factor IDEF1 is essential for the early response to iron deficiency, and induces vegetative expression of late embryogenesis abundant genes. The Plant Journal 2009, 60, 948-961, 10.1111/j.1365-313x.2009.04015.x.
- Kobayashi, T.; Itai, R.N.; Aung, M.S.; Senoura, T.; Nakanishi, H.; Nishizawa, N.K; The rice transcription factor IDEF1 directly binds to iron and other divalent metals for sensing cellular iron status. Plant J. 2012, 69, 81–91.
- Takanori Kobayashi; Yuko Nakayama; Reiko Nakanishi Itai; Hiromi Nakanishi; Toshihiro Yoshihara; Satoshi Mori; Naoko K. Nishizawa; Identification of novel cis-acting elements, IDE1 and IDE2, of the barley IDS2 gene promoter conferring iron-deficiency-inducible, root-specific expression in heterogeneous tobacco plants. The Plant Journal 2003, 36, 780-793, 10.1046/j.1365-313x.2003.01920.x.
- Lixia Zhang; Reiko Nakanishi Itai; Takashi Yamakawa; Hiromi Nakanishi; Naoko K. Nishizawa; Takanori Kobayashi; The Bowman–Birk Trypsin Inhibitor IBP1 Interacts with and Prevents Degradation of IDEF1 in Rice. Plant Molecular Biology Reporter 2014, 32, 841-851, 10.1007/s11105-013-0695-8.
- Takanori Kobayashi; Yuko Ogo; May Sann Aung; Tomoko Nozoye; Reiko Nakanishi Itai; Hiromi Nakanishi; Takashi Yamakawa; Naoko K. Nishizawa; The spatial expression and regulation of transcription factors IDEF1 and IDEF2. Annals of Botany 2010, 105, 1109-1117, 10.1093/aob/mcq002.
- Yuko Ogo; Reiko Nakanishi Itai; Hiromi Nakanishi; Haruhiko Inoue; Takanori Kobayashi; Motofumi Suzuki; Michiko Takahashi; Satoshi Mori; Naoko K. Nishizawa; Isolation and characterization of IRO2, a novel iron-regulated bHLH transcription factor in graminaceous plants. Journal of Experimental Botany 2006, 57, 2867-2878, 10.1093/jxb/erl054.
- Yuko Ogo; Reiko Nakanishi Itai; Hiromi Nakanishi; Takanori Kobayashi; Michiko Takahashi; Satoshi Mori; Naoko K. Nishizawa; The rice bHLH protein OsIRO2 is an essential regulator of the genes involved in Fe uptake under Fe-deficient conditions. The Plant Journal 2007, 51, 366-377, 10.1111/j.1365-313x.2007.03149.x.
- Shoudong Wang; Lin Li; Yinghui Ying; Jin Wang; Ji Feng Shao; Naoki Yamaji; James Whelan; Jian Feng Ma; Huixia Shou; A transcription factor OsbHLH156 regulates Strategy II iron acquisition through localising IRO2 to the nucleus in rice. New Phytologist 2019, 225, 1247-1260, 10.1111/nph.16232.
- Takanori Kobayashi; Seiji Nagasaka; Takeshi Senoura; Reiko Nakanishi Itai; Hiromi Nakanishi; Naoko K. Nishizawa; Iron-binding haemerythrin RING ubiquitin ligases regulate plant iron responses and accumulation. Nature Communications 2013, 4, 2792, 10.1038/ncomms3792.
- Huimin Zhang; Yang Li; Xiani Yao; Gang Liang; Diqiu Yu; POSITIVE REGULATOR OF IRON HOMEOSTASIS1, OsPRI1, Facilitates Iron Homeostasis. Plant Physiology 2017, 175, 543-554, 10.1104/pp.17.00794.
- Huimin Zhang; Yang Li; Mengna Pu; Peng Xu; Gang Liang; Diqiu Yu; Oryza sativa POSITIVE REGULATOR OF IRON DEFICIENCY RESPONSE 2 (OsPRI2) and OsPRI3 are involved in the maintenance of Fe homeostasis. Plant, Cell & Environment 2020, 43, 261-274, 10.1111/pce.13655.
- Takanori Kobayashi; Asami Ozu; Subaru Kobayashi; Gynheung An; Jong-Seong Jeon; Naoko K. Nishizawa; OsbHLH058 and OsbHLH059 transcription factors positively regulate iron deficiency responses in rice.. Plant Molecular Biology 2019, 101, 471-486, 10.1007/s11103-019-00917-8.
- Luqing Zheng; Yinghui Ying; Lu Wang; Fang Wang; James Whelan; Huixia Shou; Identification of a novel iron regulated basic helix-loop-helix protein involved in Fe homeostasis in Oryza sativa. BMC Plant Biology 2010, 10, 1–9, 10.1186/1471-2229-10-166.
- Y. Ogo; T. Kobayashi; R. Nakanishi Itai; Hiromi Nakanishi; Y. Kakei; M. Takahashi; S. Toki; S. Mori; N. K. Nishizawa; A Novel NAC Transcription Factor, IDEF2, That Recognizes the Iron Deficiency-responsive Element 2 Regulates the Genes Involved in Iron Homeostasis in Plants. Journal of Biological Chemistry 2008, 283, 13407-13417, 10.1074/jbc.m708732200.
- Lu Wang; Yinghui Ying; Reena Narsai; Lingxiao Ye; Luqing Zheng; Jingluan Tian; James Whelan; Huixia Shou; Identification of OsbHLH133 as a regulator of iron distribution between roots and shoots inOryza sativa. Plant, Cell & Environment 2013, 36, 224-236, 10.1111/j.1365-3040.2012.02569.x.
- Louis Grillet; Ping Lan; Wenfeng Li; Girish Mokkapati; Wolfgang Schmidt; IRON MAN is a ubiquitous family of peptides that control iron transport in plants. Nature Plants 2018, 4, 953-963, 10.1038/s41477-018-0266-y.
- Cristobal Uauy; Assaf Distelfeld; Tzion Fahima; Ann Blechl; Jorge Dubcovsky; A NAC Gene Regulating Senescence Improves Grain Protein, Zinc, and Iron Content in Wheat. Science 2006, 314, 1298-1301, 10.1126/science.1133649.
- Assaf Distelfeld; Stephen P. Pearce; Raz Avni; Beatrice Scherer; Cristobal Uauy; Fernando Piston; Ann Slade; Rongrong Zhao; Jorge Dubcovsky; Divergent functions of orthologous NAC transcription factors in wheat and rice. Plant Molecular Biology 2012, 78, 515-24, 10.1007/s11103-012-9881-6.
- Takanori Kobayashi; Reiko Nakanishi Itai; Takeshi Senoura; Takaya Oikawa; Yasuhiro Ishimaru; Minoru Ueda; Hiromi Nakanishi; Naoko K. Nishizawa; Jasmonate signaling is activated in the very early stages of iron deficiency responses in rice roots. Plant Molecular Biology 2016, 91, 533-47, 10.1007/s11103-016-0486-3.
- Shen, C.; Yue, R.; Sun, T.; Zhang, L.; Yang, Y.; Wang, H. OsARF16, a transcription factor regulating auxin redistribution, is required for iron deficiency response in rice (Oryza sativa L.). Plant Sci. 2015, 231, 148–158.
- Curie, C.; Briat, J.-F; Iron transport and signaling in plants. Annu. Rev. Plant Biol. 2003, 54, 183–206.
- Meng Wang; Wilhelm Gruissem; Navreet K. Bhullar; Nicotianamine synthase overexpression positively modulates iron homeostasis-related genes in high iron rice. Frontiers in Plant Science 2013, 4, 1–15, 10.3389/fpls.2013.00156.
- Longjun Cheng; Fang Wang; Huixia Shou; Fangliang Huang; Luqing Zheng; Fei He; Jinhui Li; Fang-Jie Zhao; Daisei Ueno; Jian Feng Ma; Ping Wu; Mutation in Nicotianamine Aminotransferase Stimulated the Fe(II) Acquisition System and Led to Iron Accumulation in Rice1[C][W][OA]. Plant Physiology 2007, 145, 1647-1657, 10.1104/pp.107.107912.
