Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 2 by Rita Xu and Version 1 by Inês L. S. Delgado.
The success of the intracellular parasite Toxoplasma gondii in invading host cells relies on the apical complex, a specialized microtubule cytoskeleton structure associated with secretory organelles. The T. gondii genome encodes three isoforms of both α- and β-tubulin, which undergo specific post-translational modifications (PTMs), altering the biochemical and biophysical proprieties of microtubules and modulating their interaction with associated proteins. Tubulin PTMs represent a powerful and evolutionarily conserved mechanism for generating tubulin diversity, forming a biochemical ‘tubulin code’ interpretable by microtubule-interacting factors.
- Toxoplasma gondii
- tubulin glutamylation
- tubulin post-translational modifications
1. Introduction
The phylum Apicomplexa compromises around 5000 obligate intracellular parasites, many of which are highly significant in veterinary and medical contexts [1]. Key members include species in the genus Plasmodium, responsible for malaria in humans, a disease with severe consequences [2]; Eimeria, a pathogen affecting poultry and cattle [3,4][3][4]; Cryptosporidium, an opportunistic pathogen affecting both humans and animals [5]; as well as Besnoitia, Babesia, and Theileria, parasites impacting cattle [6,7,8][6][7][8]. The tissue-cyst-forming coccidian Toxoplasma gondii affects both domestic and wild animals [9]. In humans, it commonly causes congenital neurological and ocular defects [10], and poses a serious threat to immunocompromised individuals [11]. Its ability to spread through water and food has led to its categorization as a category B priority pathogen by the National Institute for Allergy and Infectious Diseases (NIAID) [12].
Understanding how parasites enter host cells and proliferate is crucial for comprehending diseases and may aid in identifying targets for the development of novel therapeutic approaches. The invasion process of Apicomplexa zoites and the molecular mechanisms underlying it appear to be conserved. To penetrate host cells, apicomplexans employ a system of adhesion-based motility known as gliding, which has been observed to depend on actin/myosin interactions [13]. In this process, the apical complex, a microtubule (MT)-based cytoskeletal structure localized in the anterior region of the cell, plays a pivotal role in the interaction with the host cell [14]. Importantly, the molecular composition of this structure remains incompletely understood, but it is likely enriched in proteins involved in MT assembly and dynamics, as well as in proteins participating in MT-associated processes and their interplay with other cellular systems (e.g., actin, vesicles). As a crucial component of the apical complex, comprehending how this specialized class of MTs is assembled, maintained, and functionally interacts with other cellular structures is paramount. Current understanding of the biology of the apical complex suggests that tubulin post-translational modifications (PTMs) and the machineries responsible for their generation and removal play pivotal roles in the assembly and functions of this structure during host cell invasion.
2. Microtubule Cytoskeleton
Throughout evolution, eukaryotic cells developed highly sophisticated and specialized cytoskeleton systems, including intermediate filaments, actin filaments, and MTs. Despite their specific roles, these structures crosstalk and cooperate, such as in supporting membrane structures like nuclear and plasma membranes, thereby imparting shape and mechanical resistance to the cell [15]. Moreover, eukaryotic cytoskeletons are involved in various processes, including cytoplasmic organization, organelle assembly and maintenance, cell division, cell polarity, cell migration, intracellular transport, and cell signaling [16,17][16][17]. In multicellular organisms, the cytoskeleton also plays crucial roles in establishing cell–cell contacts and cell–extracellular matrix interactions, thereby contributing to tissue integrity [18].
MTs are dynamic polymers composed of heterodimers of the structurally and functionally conserved α- and β-tubulins, both of which are GTP-binding proteins and are ubiquitous across all studied eukaryotes [19]. Higher organisms, including humans and mice, possess extensive gene families encoding multiple α- and β-tubulin isotypes [20]. Significantly, certain tubulin isotypes are expressed in a tissue-specific manner, playing key roles in the assembly of specialized functional classes of MTs. For instance, the proper expression of specific tubulin isotypes (e.g., β3-tubulin) is essential for neuronal differentiation and survival in mammals, and mutations in their coding genes are linked to neuronal disorders [21,22,23][21][22][23]. In contrast, lower eukaryotes such as Saccharomyces [24,25][24][25] and Tetrahymena [26,27][26][27] typically harbor one or two α- and two β- canonical tubulin-coding genes.
For the assembly of the α/β-tubulin heterodimer, tubulins undergo a complex folding process assisted by molecular chaperones (prefoldin and CCT) [28,29,30,31][28][29][30][31] and tubulin cofactors (TBCA-E) [32]. Besides their roles in heterodimer assembly, tubulin cofactors also contribute to quality control and recycling of heterodimers released from depolymerized MTs. Thus, by regulating the pool of free tubulin dimers competent to polymerize, the tubulin folding pathway controls MT dynamics [33,34,35,36,37][33][34][35][36][37]. Once folded and assembled, tubulin heterodimers polymerize in a polarized head-to-tail manner to form protofilaments, which then assemble into the characteristic hollow structure of MTs, typically composed of 13 protofilaments [19]. During the polymerization process, β-tubulin hydrolyses its GTP to GDP, and upon MT depolymerization, the GDP is exchanged to GTP to enable β-tubulin polymerization again. In contrast, GTP bound to α-tubulin remains not hydrolyzed during polymerization [38,39][38][39].
Due to the polar nature of MTs, one end (the minus end) is comprised of α-tubulin subunits, while the other end (the plus end) consists of β-tubulin. Furthermore, the two ends of MTs exhibit distinct dynamic properties, with the minus end presenting slow growth and the plus end undergoing rapid polymerization [19]. Typically, the minus end is associated with MT organizing centers (MTOCs), such as spindle pole bodies in fungi and centrosomes and the Golgi apparatus in animal cells. MTOCs exhibit structural variability across different eukaryotic groups but are consistently enriched in proteins that facilitate MT nucleation (e.g., gamma-tubulin) and anchoring [40,41,42,43][40][41][42][43].
MTs can present varied dynamic properties and stability, which are influenced by factors such as the preferential incorporation of specific tubulin isotypes (products of different tubulin genes) and by tubulin PTMs such as acetylation, detyrosination, glutamylation, and glycylation [44,45][44][45]. Tubulin PTMs can selectively and reversibly affect distinct MT subpopulations [46]. These modifications are evolutionarily conserved and contribute to what is termed the “tubulin code”.
For instance, α-tubulin can undergo acetylation at K40, which is the sole known PTM that localizes in the luminal surface of MTs [44,45][44][45]. Additionally, its C-terminal tail can undergo a reversible modification through the deletion of the terminal tyrosine (detyrosination) or an irreversible modification by deletion of the last two residues (Δ2). The C-terminal tails of both α- and β-tubulins can also be reversibly modified by glutamylation and glycylation [44,45][44][45]. While the K40 PTM has been linked to MT stability [47], C-terminal PTMs alter MT interactions with associated proteins, thereby influencing sensitivity to MT-targeting drugs.
Tubulin PTMs play a crucial role in the binding of MT-associated proteins (MAPs), such as MT motors and MT-severing proteins, to MTs. Consequently, they are essential for the assembly and maintenance of MT-based organelles such as centrioles, cilia, and flagella, as well as MT cortical structures found in unicellular organisms like Tetrahymena thermophila and T. gondii [46,48][46][48]. Thus, centriolar and axonemal MTs exhibit high levels of acetylation and glutamylation compared to cytoplasmic MTs [49]. In cilia, although the precise impact of tubulin PTMs on intra-flagellar transport is not fully understood, tubulin glutamylation has been shown to affect intra-flagellar transport velocity [50] and the localization of MT motor proteins [51]. Importantly, tubulin PTMs can be regulated in response to environmental cues. In Caenorhabditis elegans, for instance, tubulin glutamylation was shown to be upregulated in sensory cilia in response to changes in temperature, osmolality, and dietary conditions [50].
3. The Specialized Microtubule Structures of Toxoplasma gondii
Apicomplexans are classified as alveolate organisms due to the presence of a flattened vesicle system (alveoli) underlying their plasma membrane, forming a structure known as the pellicle [52,53,54][52][53][54]. The pellicle is further divided into three subdomains (apical, central, and basal), each with distinct properties conferred by specific cytoskeleton components. The plasma membrane associated with alveoli form the inner membrane complex (IMC), which extends from the apical polar ring (APR) to the basal pole, leaving the extreme apical region of the parasite solely enclosed by plasma membrane. While γ-tubulin does not localize to the APR [55], this structure is regarded as an MTOC because the minus ends of subpellicular MTs (SPMTs) are anchored there via cogwheel-like projections, with their plus ends extending distally from this structure [56,57][56][57]. SPMTs extend in a gentle spiral from the APR to a region posterior to the nucleus, contributing to the elongated shape, rigidity, and the maintenance of the highly polarized cell organization [58] (Figure 1). These SPMTs are coated with MAPs [59], among which are Subpellicular Microtubule Proteins 1 and 2 (SPM1 and SPM2), unique to apicomplexan parasites [60]. Up to this point, the proteins identified as components of the APR include TgRNG1, TgRNG2, TgAPR1, and TgKinesin A [56,61,62,63][56][61][62][63]. TgRNG1 becomes detectable at the mature APR only after nuclear division is complete [61]. TgAPR1 is also a marker of the mature APR structure, and parasites lacking TgAPR1 demonstrate a defect in the lytic cycle [56]. The MT motor TgKinesin A, while not essential, plays a role in parasite growth, and parasites lacking this protein exhibit a modest reduction in growth rate. TgKinesin A localizes to emerging daughter buds and is positioned just apical to APR1 at the APR of mature parasites [56].
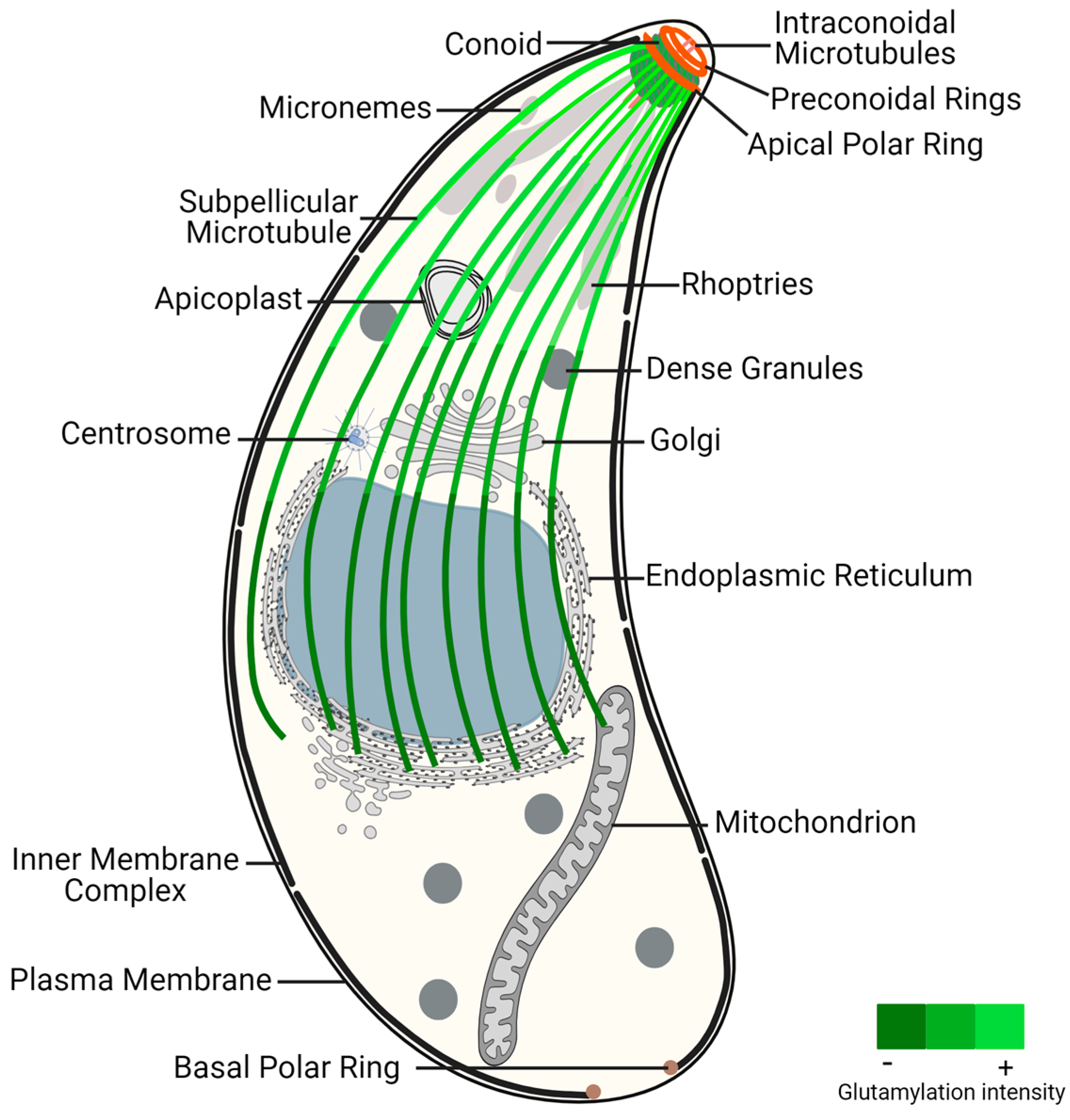
Figure 1. Toxoplasma gondii tachyzoite ultrastructure, highlighting the glutamylation of subpellicular microtubules. This post-translational modification presents increased density near the conoid region, an apical structure critical for parasite invasion and motility, decreasing toward the distal end of subpellicular microtubules. The gradient of glutamylation suggests a functional stratification within the microtubule network, essential for the parasite’s life cycle and pathogenicity.
The apical complex is centered around the extensible and retractable conoid, which exhibits active extrusion during host invasion [64]. The conoid, mainly composed of tubulin, adopts a unique polymer form distinct from typical MTs [65]. It comprises two preconoidal rings above the conoid and two intraconoidal MTs [66]. While related alveolates may possess incomplete conoids or pseudoconoids, most apicomplexans were thought to have lost the conoid structure, with coccidian parasites, like T. gondii, retaining a closed conoid [67]. However, recent data indicate that the conoid is a hallmark of invasion mechanisms conserved in all apicomplexans and is also present in other alveolates [68]. Proteomics analysis of the T. gondii conoid/apical complex has identified approximately 200 proteins, representing 70% of T. gondii cytoskeleton proteins. These proteins include several key cytoskeletal components such as actin and actin-binding proteins, varied myosin heavy and light chains, and all three isoforms of β-tubulin [14].
At the apical pole, specialized secretory organelles called rhoptries and micronemes play crucial roles traversing within the conoid to secrete their contents across the plasma membrane at the apical tip of the parasite [69].
As part of the MT cytoskeleton, T. gondii tachyzoites also possess centrioles, which are barrel-shaped structures formed by nine singlet MTs [58]. Unlike non-coccidian apicomplexans such as Plasmodium which lack asexual centrioles, this structure occurs in other coccidians. Coccidian centrioles are relatively short and arranged in a parallel rather than orthogonal configuration [67]. Despite the absence of pericentrin and ninein genes, the term “centrosome” has been used in T. gondii due to its nucleating activity and its ability to function as a signaling platform [70].
The nuclear division in T. gondii relies on the centrocone, a domain of the nuclear envelope, and occurs without the nuclear envelope breaking down [71,72][71][72]. Unlike in other organisms, chromosome segregation in T. gondii occurs without chromosome condensation. Spindle MTs originate in the cytoplasm and traverse the nuclear pores of the centrocone. These MTs are crucial for linking centrosomes with the centrocone and for segregating chromosomes into daughter nuclei [71,72][71][72]. EB1 proteins bind to the positive ends of dynamic MTs, promoting stability and elongation of the MTs. In T. gondii, the EB1 homolog is a nuclear protein that localizes to the centrocone after spindle assembly [73]. Moreover, in addition to the nuclear protein remaining at the centrocone until cytokinesis, there exists a small pool of cytoplasmic TgEB1 that transiently associates with the tips of the daughter buds’ SPMTs after nuclear division is complete [73]. Studies in T. gondii have revealed that SPMTs continue to polymerize during daughter cell assembly. However, during later stages of cell division, SPMTs exhibit high resistance to various conditions that typically lead to MT depolymerization, such as cold, antimitotic agents, detergents, and high pressure [58].
MAPs play a critical role in influencing MT stability and imparting different properties to MT populations. While MT motors (dyneins and kinesins), centriole components (SAS-6, centrin, CEP250), and regulatory proteins (EB1) are highly conserved among eukaryotes, proteins associated with SPMTs and the conoid are predominantly specific to these organisms, reflecting the specialized functions of these structures (Morrissette and Gubbels, 2020 [67]). SPMTs are extensively coated with MAPs such as TgSPM1, TgSPM2, TgTrxL1, TgTrxL2, TgTLAP1, TgTLAP2, TgTLAP3, and TgTLAP4. While some of these proteins are distributed throughout SPMTs, others localize to specific subregions [58].
References
- Levine, N.D. Progress in Taxonomy of the Apicomplexan Protozoa. J. Protozool. 1988, 35, 518–520.
- WHO. World Malaria Report 2020: 20 Years of Global Progress and Challenges; WHO: Geneva, Switzerland, 2020.
- Blake, D.P.; Tomley, F.M. Securing Poultry Production from the Ever-Present Eimeria Challenge. Trends Parasitol. 2014, 30, 12–19.
- Chartier, C.; Paraud, C. Coccidiosis Due to Eimeria in Sheep and Goats, a Review. Small Rumin. Res. 2012, 103, 84–92.
- Tzipori, S.; Ward, H. Cryptosporidiosis: Biology, Pathogenesis and Disease. Microbes Infect. 2002, 4, 1047–1058.
- Álvarez-García, G.; García-Lunar, P.; Gutiérrez-Expósito, D.; Shkap, V.; Ortega-Mora, L.M. Dynamics of Besnoitia Besnoiti Infection in Cattle. Parasitology 2014, 141, 1419–1435.
- Jacob, S.S.; Sengupta, P.P.; Paramanandham, K.; Suresh, K.P.; Chamuah, J.K.; Rudramurthy, G.R.; Roy, P. Bovine Babesiosis: An Insight into the Global Perspective on the Disease Distribution by Systematic Review and Meta-Analysis. Vet. Parasitol. 2020, 283, 109136.
- Mans, B.J.; Pienaar, R.; Latif, A.A. A Review of Theileria Diagnostics and Epidemiology. Int. J. Parasitol. Parasites Wildl. 2015, 4, 104–118.
- Delgado, I.L.S.; Zúquete, S.; Santos, D.; Basto, A.P.; Leitão, A.; Nolasco, S. The Apicomplexan Parasite Toxoplasma gondii. Encyclopedia 2022, 2, 189–211.
- Khan, K.; Khan, W. Congenital Toxoplasmosis: An Overview of the Neurological and Ocular Manifestations. Parasitol. Int. 2018, 67, 715–721.
- Wang, Z.-D.; Liu, H.-H.; Ma, Z.-X.; Ma, H.-Y.; Li, Z.-Y.; Yang, Z.-B.; Zhu, X.-Q.; Xu, B.; Wei, F.; Liu, Q. Toxoplasma gondii Infection in Immunocompromised Patients: A Systematic Review and Meta-Analysis. Front. Microbiol. 2017, 8, 389.
- Slifko, T.R.; Smith, H.V.; Rose, J.B. Emerging Parasite Zoonoses Associated with Water and Food. Int. J. Parasitol. 2000, 30, 1379–1393.
- Frénal, K.; Dubremetz, J.-F.; Lebrun, M.; Soldati-Favre, D. Gliding Motility Powers Invasion and Egress in Apicomplexa. Nat. Rev. Microbiol. 2017, 15, 645–660.
- Hu, K.; Johnson, J.; Florens, L.; Fraunholz, M.; Suravajjala, S.; DiLullo, C.; Yates, J.; Roos, D.S.; Murray, J.M. Cytoskeletal Components of an Invasion Machine—The Apical Complex of Toxoplasma gondii. PLoS Pathog. 2006, 2, e13.
- Fletcher, D.A.; Mullins, R.D. Cell Mechanics and the Cytoskeleton. Nature 2010, 463, 485–492.
- Hohmann, T.; Dehghani, F. The Cytoskeleton-A Complex Interacting Meshwork. Cells 2019, 8, 362.
- Moujaber, O.; Stochaj, U. The Cytoskeleton as Regulator of Cell Signaling Pathways. Trends Biochem. Sci. 2020, 45, 96–107.
- Sluysmans, S.; Vasileva, E.; Spadaro, D.; Shah, J.; Rouaud, F.; Citi, S. The Role of Apical Cell-Cell Junctions and Associated Cytoskeleton in Mechanotransduction. Biol. Cell 2017, 109, 139–161.
- Goodson, H.V.; Jonasson, E.M. Microtubules and Microtubule-Associated Proteins. Cold Spring Harb. Perspect. Biol. 2018, 10, a022608.
- Findeisen, P.; Mühlhausen, S.; Dempewolf, S.; Hertzog, J.; Zietlow, A.; Carlomagno, T.; Kollmar, M. Six Subgroups and Extensive Recent Duplications Characterize the Evolution of the Eukaryotic Tubulin Protein Family. Genome Biol. Evol. 2014, 6, 2274–2288.
- Jaglin, X.H.; Poirier, K.; Saillour, Y.; Buhler, E.; Tian, G.; Bahi-Buisson, N.; Fallet-Bianco, C.; Phan-Dinh-Tuy, F.; Kong, X.P.; Bomont, P.; et al. Mutations in the Beta-Tubulin Gene TUBB2B Result in Asymmetrical Polymicrogyria. Nat. Genet. 2009, 41, 746–752.
- Tischfield, M.A.; Baris, H.N.; Wu, C.; Rudolph, G.; Van Maldergem, L.; He, W.; Chan, W.-M.; Andrews, C.; Demer, J.L.; Robertson, R.L.; et al. Human TUBB3 Mutations Perturb Microtubule Dynamics, Kinesin Interactions, and Axon Guidance. Cell 2010, 140, 74–87.
- Tischfield, M.A.; Engle, E.C. Distinct Alpha- and Beta-Tubulin Isotypes Are Required for the Positioning, Differentiation and Survival of Neurons: New Support for the “multi-Tubulin” Hypothesis. Biosci. Rep. 2010, 30, 319–330.
- Neff, N.F.; Thomas, J.H.; Grisafi, P.; Botstein, D. Isolation of the Beta-Tubulin Gene from Yeast and Demonstration of Its Essential Function In Vivo. Cell 1983, 33, 211–219.
- Schatz, P.J.; Pillus, L.; Grisafi, P.; Solomon, F.; Botstein, D. Two Functional Alpha-Tubulin Genes of the Yeast Saccharomyces Cerevisiae Encode Divergent Proteins. Mol. Cell. Biol. 1986, 6, 3711–3721.
- Barahona, I.; Soares, H.; Cyrne, L.; Penque, D.; Denoulet, P.; Rodrigues-Pousada, C. Sequence of One α- and Two β-Tubulin Genes of Tetrahymena pyriformis. Structural and Functional Relationships with Other Eukaryotic Tubulin Genes. J. Mol. Biol. 1988, 202, 365–382.
- Gaertig, J.; Thatcher, T.H.; McGrath, K.E.; Callahan, R.C.; Gorovsky, M.A. Perspectives on Tubulin Isotype Function and Evolution Based on the Observation That Tetrahymena thermophila Microtubules Contain a Single α- and β-Tubulin. Cell Motil. Cytoskeleton 1993, 25, 243–253.
- Vainberg, I.E.; Lewis, S.A.; Rommelaere, H.; Ampe, C.; Vandekerckhove, J.; Klein, H.L.; Cowan, N.J. Prefoldin, a Chaperone That Delivers Unfolded Proteins to Cytosolic Chaperonin. Cell 1998, 93, 863–873.
- Hansen, W.J.; Cowan, N.J.; Welch, W.J. Prefoldin-Nascent Chain Complexes in the Folding of Cytoskeletal Proteins. J. Cell Biol. 1999, 145, 265–277.
- Gestaut, D.; Zhao, Y.; Park, J.; Ma, B.; Leitner, A.; Collier, M.; Pintilie, G.; Roh, S.-H.; Chiu, W.; Frydman, J. Structural Visualization of the Tubulin Folding Pathway Directed by Human Chaperonin TRiC/CCT. Cell 2022, 185, 4770–4787.e20.
- Liu, C.; Jin, M.; Wang, S.; Han, W.; Zhao, Q.; Wang, Y.; Xu, C.; Diao, L.; Yin, Y.; Peng, C.; et al. Pathway and Mechanism of Tubulin Folding Mediated by TRiC/CCT along Its ATPase Cycle Revealed Using Cryo-EM. Commun. Biol. 2023, 6, 531.
- Lopez-Fanarraga, M.; Avila, J.; Guasch, A.; Coll, M.; Zabala, J.C. Review: Postchaperonin Tubulin Folding Cofactors and Their Role in Microtubule Dynamics. J. Struct. Biol. 2001, 135, 219–229.
- Serna, M.; Zabala, J.C. Tubulin Folding and Degradation. In Encyclopedia of Life Sciences; Wiley: Hoboken, NJ, USA, 2016; pp. 1–9.
- Gonçalves, J.; Tavares, A.; Carvalhal, S.; Soares, H. Revisiting the Tubulin Folding Pathway: New Roles in Centrosomes and Cilia. Biomol. Concepts 2010, 1, 423–434.
- Nolasco, S.; Bellido, J.; Serna, M.; Carmona, B.; Soares, H.; Zabala, J.C. Colchicine Blocks Tubulin Heterodimer Recycling by Tubulin Cofactors TBCA, TBCB, and TBCE. Front. Cell Dev. Biol. 2021, 9, 656273.
- Francis, J.W.; Goswami, D.; Novick, S.J.; Pascal, B.D.; Weikum, E.R.; Ortlund, E.A.; Griffin, P.R.; Kahn, R.A. Nucleotide Binding to ARL2 in the TBCD∙ARL2∙β-Tubulin Complex Drives Conformational Changes in β-Tubulin. J. Mol. Biol. 2017, 429, 3696–3716.
- Francis, J.W.; Newman, L.E.; Cunningham, L.A.; Kahn, R.A. A Trimer Consisting of the Tubulin-Specific Chaperone D (TBCD), Regulatory GTPase ARL2, and β-Tubulin Is Required for Maintaining the Microtubule Network. J. Biol. Chem. 2017, 292, 4336–4349.
- Spiegelman, B.M.; Penningroth, S.M.; Kirschner, M.W. Turnover of Tubulin and the N Site GTP in Chinese Hamster Ovary Cells. Cell 1977, 12, 587–600.
- Kristensson, M.A. The Game of Tubulins. Cells 2021, 10, 745.
- Chabin-Brion, K.; Marceiller, J.; Perez, F.; Settegrana, C.; Drechou, A.; Durand, G.; Poüs, C. The Golgi Complex Is a Microtubule-Organizing Organelle. Mol. Biol. Cell 2001, 12, 2047–2060.
- Yukawa, M.; Ikebe, C.; Toda, T. The Msd1-Wdr8-Pkl1 Complex Anchors Microtubule Minus Ends to Fission Yeast Spindle Pole Bodies. J. Cell Biol. 2015, 209, 549–562.
- Sanchez, A.D.; Feldman, J.L. Microtubule-Organizing Centers: From the Centrosome to Non-Centrosomal Sites. Curr. Opin. Cell Biol. 2017, 44, 93–101.
- Jaspersen, S.L. Anatomy of the Fungal Microtubule Organizing Center, the Spindle Pole Body. Curr. Opin. Struct. Biol. 2021, 66, 22–31.
- Wloga, D.; Joachimiak, E.; Fabczak, H. Tubulin Post-Translational Modifications and Microtubule Dynamics. Int. J. Mol. Sci. 2017, 18, 2207.
- Carmona, B.; Marinho, H.S.; Matos, C.L.; Nolasco, S.; Soares, H. Tubulin Post-Translational Modifications: The Elusive Roles of Acetylation. Biology 2023, 12, 561.
- Wloga, D.; Gaertig, J. Post-Translational Modifications of Microtubules. J. Cell Sci. 2010, 123, 3447–3455.
- Janke, C.; Montagnac, G. Causes and Consequences of Microtubule Acetylation. Curr. Biol. 2017, 27, R1287–R1292.
- Xiao, H.; El Bissati, K.; Verdier-Pinard, P.; Burd, B.; Zhang, H.; Kim, K.; Fiser, A.; Angeletti, R.H.; Weiss, L.M. Post-Translational Modifications to Toxoplasma gondii α- and β-Tubulins Include Novel C-Terminal Methylation. J. Proteome Res. 2010, 9, 359–372.
- Orbach, R.; Howard, J. The dynamic and structural properties of axonemal tubulins support the high length stability of cilia. Nat Commun. 2019, 10, 1838.
- Kimura, Y.; Tsutsumi, K.; Konno, A.; Ikegami, K.; Hameed, S.; Kaneko, T.; Kaplan, O.I.; Teramoto, T.; Fujiwara, M.; Ishihara, T.; et al. Environmental Responsiveness of Tubulin Glutamylation in Sensory Cilia Is Regulated by the P38 MAPK Pathway. Sci. Rep. 2018, 8, 8392.
- Power, K.M.; Akella, J.S.; Gu, A.; Walsh, J.D.; Bellotti, S.; Morash, M.; Zhang, W.; Ramadan, Y.H.; Ross, N.; Golden, A.; et al. Mutation of NEKL-4/NEK10 and TTLL Genes Suppress Neuronal Ciliary Degeneration Caused by Loss of CCPP-1 Deglutamylase Function. PLoS Genet. 2020, 16, e1009052.
- Leander, B.S.; Keeling, P.J. Morphostasis in Alveolate Evolution. Trends Ecol. Evol. 2003, 18, 395–402.
- Leander, B.S.; Kuvardina, O.N.; Aleshin, V.V.; Mylnikov, A.P.; Keeling, P.J. Molecular Phylogeny and Surface Morphology of Colpodella edax (Alveolata): Insights into the Phagotrophic Ancestry of Apicomplexans. J. Eukaryot. Microbiol. 2003, 50, 334–340.
- Goodenough, U.; Roth, R.; Kariyawasam, T.; He, A.; Lee, J.-H. Epiplasts: Membrane skeletons and epiplastin proteins in euglenids, glaucophytes, cryptophytes, ciliates, dinoflagellates, and apicomplexans. mBio 2018, 9, e02020-18.
- Suvorova, E.S.; Francia, M.; Striepen, B.; White, M.W. A Novel Bipartite Centrosome Coordinates the Apicomplexan Cell Cycle. PLoS Biol. 2015, 13, e1002093.
- Leung, J.M.; He, Y.; Zhang, F.; Hwang, Y.-C.; Nagayasu, E.; Liu, J.; Murray, J.M.; Hu, K. Stability and Function of a Putative Microtubule-Organizing Center in the Human Parasite Toxoplasma gondii. Mol. Biol. Cell 2017, 28, 1361–1378.
- Russell, D.G.; Burns, R.G. The Polar Ring of Coccidian Sporozoites: A Unique Microtubule-Organizing Centre. J. Cell Sci. 1984, 65, 193–207.
- Morrissette, N.S.; Sibley, L.D. Cytoskeleton of Apicomplexan Parasites. Microbiol. Mol. Biol. Rev. 2002, 66, 21–38, table of contents.
- Morrissette, N.S.; Murray, J.M.; Roos, D.S. Subpellicular Microtubules Associate with an Intramembranous Particle Lattice in the Protozoan Parasite Toxoplasma gondii. J. Cell Sci. 1997, 110 Pt 1, 35–42.
- Tran, J.Q.; Li, C.; Chyan, A.; Chung, L.; Morrissette, N.S. SPM1 Stabilizes Subpellicular Microtubules in Toxoplasma gondii. Eukaryot. Cell 2012, 11, 206–216.
- Tran, J.Q.; de Leon, J.C.; Li, C.; Huynh, M.-H.; Beatty, W.; Morrissette, N.S. RNG1 Is a Late Marker of the Apical Polar Ring in Toxoplasma gondii. Cytoskeleton 2010, 67, 586–598.
- Gould, S.B.; Kraft, L.G.K.; van Dooren, G.G.; Goodman, C.D.; Ford, K.L.; Cassin, A.M.; Bacic, A.; McFadden, G.I.; Waller, R.F. Ciliate Pellicular Proteome Identifies Novel Protein Families with Characteristic Repeat Motifs That Are Common to Alveolates. Mol. Biol. Evol. 2011, 28, 1319–1331.
- Katris, N.J.; van Dooren, G.G.; McMillan, P.J.; Hanssen, E.; Tilley, L.; Waller, R.F. The Apical Complex Provides a Regulated Gateway for Secretion of Invasion Factors in Toxoplasma. PLoS Pathog. 2014, 10, e1004074.
- Mondragon, R.; Frixione, E. Ca(2+)-Dependence of Conoid Extrusion in Toxoplasma gondii Tachyzoites. J. Eukaryot. Microbiol. 1996, 43, 120–127.
- Hu, K.; Roos, D.S.; Murray, J.M. A Novel Polymer of Tubulin Forms the Conoid of Toxoplasma gondii. J. Cell Biol. 2002, 156, 1039–1050.
- Nichols, B.A.; Chiappino, M.L. Cytoskeleton of Toxoplasma gondii. J. Protozool. 1987, 34, 217–226.
- Morrissette, N.; Gubbels, M.-J. The Toxoplasma Cytoskeleton: Structures, Proteins, and Processes. In Toxoplasma gondii, 3rd ed.; Louis, M., Weiss, K.K., Eds.; Academic Press: Cambridge, MA, USA, 2020; pp. 743–788.
- Koreny, L.; Zeeshan, M.; Barylyuk, K.; Tromer, E.C.; van Hooff, J.J.E.; Brady, D.; Ke, H.; Chelaghma, S.; Ferguson, D.J.P.; Eme, L.; et al. Molecular characterization of the conoid complex in Toxoplasma reveals its conservation in all apicomplexans, including Plasmodium species. PLoS Biol. 2021, 19, e3001081.
- Mageswaran, S.K.; Guérin, A.; Theveny, L.M.; Chen, W.D.; Martinez, M.; Lebrun, M.; Striepen, B.; Chang, Y.-W. In Situ Ultrastructures of Two Evolutionarily Distant Apicomplexan Rhoptry Secretion Systems. Nat. Commun. 2021, 12, 4983.
- Francia, M.E.; Dubremetz, J.-F.; Morrissette, N.S. Basal Body Structure and Composition in the Apicomplexans Toxoplasma and Plasmodium. Cilia 2015, 5, 3.
- Naumov, A.; Kratzer, S.; Ting, L.-M.; Kim, K.; Suvorova, E.S.; White, M.W. The Toxoplasma centrocone houses cell cycle regulatory factors. mBio 2017, 8, e00579-17.
- Farrell, M.; Gubbels, M.-J. The Toxoplasma gondii Kinetochore Is Required for Centrosome Association with the Centrocone (Spindle Pole). Cell. Microbiol. 2014, 16, 78–94.
- Chen, C.-T.; Kelly, M.; de Leon, J.; Nwagbara, B.; Ebbert, P.; Ferguson, D.J.P.; Lowery, L.A.; Morrissette, N.; Gubbels, M.-J. Compartmentalized Toxoplasma EB1 Bundles Spindle Microtubules to Secure Accurate Chromosome Segregation. Mol. Biol. Cell 2015, 26, 4562–4576.
- Hu, K. Organizational Changes of the Daughter Basal Complex during the Parasite Replication of Toxoplasma gondii. PLoS Pathog. 2008, 4, e10.
- Gubbels, M.-J.; White, M.; Szatanek, T. The Cell Cycle and Toxoplasma gondii Cell Division: Tightly Knit or Loosely Stitched? Int. J. Parasitol. 2008, 38, 1343–1358.
- Blader, I.J.; Coleman, B.I.; Chen, C.-T.; Gubbels, M.-J. Lytic Cycle of Toxoplasma gondii: 15 Years Later. Annu. Rev. Microbiol. 2015, 69, 463–485.
More
