Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 1 by Gloria Borgstahl and Version 2 by Mona Zou.
Human Replication Protein A (RPA) was historically discovered as one of the six components needed to reconstitute simian virus 40 DNA replication from purified components. RPA is now known to be involved in all DNA metabolism pathways that involve single-stranded DNA (ssDNA). Heterotrimeric RPA comprises several domains connected by flexible linkers and is heavily regulated by post-translational modifications (PTMs). The structure of RPA has been challenging to obtain. Various structural methods have been applied, but a complete understanding of RPA’s flexible structure, its function, and how it is regulated by PTMs has yet to be obtained.
- Replication Protein A (RPA)
- phosphorylation
- homologous recombination
- AlphaFold
- protein-ssDNA interactions
- cell cycle
- DNA metabolism
- double-strand break repair
1. The Function of RPA in DNA Double-Strand Break Repair
RPA is critical in coordinating the cell cycle with DNA replication and any DNA repair pathway involving ssDNA. The cell cycle is crucial in completing a series of events that allow cells to grow and divide and includes the following stages: interphase (I), G1 phase, S phase, G2 phase, and mitosis (M). Regulation is key to successfully replicating DNA and normal cell division within these cell cycle phases. Several laboratories have studied RPA in cell cycle regulation. RPA was determined not to be phosphorylated in the G1 phase, but RPA34 in humans and S. cerevisiae is phosphorylated in cells that enter the S phase and then are dephosphorylated in the M phase [1][2][3][45,87,88]. Various researchers have focused on RPA’s role in recruiting proteins involved in DNA repair. RPA is involved in most DNA repair pathways, including base excision repair (BER), nucleotide excision repair (NER), mismatch repair (MMR), and HR [4][5][6][7][8][36,89,90,91,92]. It is also important to note that HR occurs only during S and G2 phases [9][10][40,44].
RPA plays a critical role in HR to repair DSBs. In HR, sections of homologous sequences, typically on a sister chromatid or nearby repeated sequence, will fill in the gap left by the DSB. On each side of this DSB, a 5′ strand is resected, leaving a 3′ overhang on the DNA. This resection is done by the MRN complex (composed of RAD50, mitotic recombination 11 (MRE11), and Nijmegen breakage syndrome 1 (NBS1)), along with other proteins, including CtBP-interacting protein (CtIP), exonuclease 1 (EXO1), DNA replication helicase/nuclease DNA2, and blood syndrome RecQ like helicase (BLM). Once there is a 3′ ssDNA overhang, RPA will bind to this ssDNA and follow the canonical HR pathway that will recruit a complex that includes breast cancer type 1 susceptibility protein (BRCA1), breast cancer type 2 susceptibility protein (BRCA2), and partner and localizer of BRCA2 (PALB1). This BRCA2 complex will recruit radiation sensitivity gene 51 (RAD51) and undergo ssDNA transfer from RPA to BRCA2 complex to RAD51, and RAD51 will perform a homology search on the sister chromatid, eventually repairing the DSB. Secondly, an alternative pathway for HR includes radiation sensitivity gene 52 (RAD52). RAD52 has similar functionality to the BRCA2 complex by undergoing a ssDNA handoff from RPA and allowing RAD51 nucleation of the ssDNA again to perform a homology search on the sister chromatid, eventually repairing the DSB [9][11][12][13][14][40,53,93,94,95].
2. ssDNA and Protein Interactions involving RPA
RPA involves an assorted array of interactions with ssDNA and proteins. The function of RPA binding to ssDNA has been studied in numerous ways, for example, through atomic force microscopy (AFM), circular dichroism (CD), optical tweezers, single-molecule fluorescence resonance energy transfer (smFRET), and many more [15][16][17][61,96,97]. The interaction between ssDNA and RPA is a rapid but extremely stable process that is difficult to evaluate due to the binding strength. A study in 2000 concluded that the zinc-finger motif is necessary to form a stable RPA:ssDNA complex, which depends on redox reactions [18][98]. Others focused on the binding and unfolding of non-canonical ssDNA structures by RPA in DNA replication. The heterotrimeric core of RPA selectively binds to a G-quadruplex forming sequence. G-quadruplexes create a unique problem for RPA to unfold, and various ligands could hinder DNA replication by not allowing RPA to unfold these G-quadruplexes correctly [19][99].
A more recent paper described how the ssDNA binding complex can change its binding mode to allow for nucleation of RAD51. The spacing between RPA:ssDNA complexes is essential for leaving bare nucleotides in between the complexes to allow for RAD51 nucleation. The exchange of RPA for RAD51 is known to be mediated by the RAD52 middle region [20][100]. Overall, the RPA:ssDNA complex is essential for life and continues to be investigated.
With this insight into the RPA:ssDNA complex, multiple laboratories have investigated how RPA binds to varying lengths of nucleotides (nt), for example, 8-nt, 20-nt, or 30-nt. Kang and coworkers in 2023 investigated a protein involved in Kallmann syndrome: N-methyl-D-aspartate receptor synaptonuclear signaling and neuronal migration factor (NSMF). They determined that NSMF will colocalize and physically interact with RPA during DNA damage, allowing RPA to bind in a 30-nt binding mode. This prepares RPA to become phosphorylated by ATR, an upstream kinase involved in HR [21][101]. They concluded that the 30-nt binding mode of RPA enhances the phosphorylation of RPA32 by ATR, and further phosphorylation will stabilize RPA binding to ssDNA.
RPA has many protein–protein interactions involving DSB repair proteins and others (Table 12). DSB repair binding partners include: BRCA2, RAD52, RAD51, ATR/ATRIP, DNA-PKcs, DSS1, MRE11-RAD50-NBS1, p53, and PP2A [4][6][22][23][24][25][26][27][28][29][30][31][32][33][34][35][36][37][38][39][40][41][42][43][44][45][46][47][48][49][50][51][52][53][54][55][56][57][58][59][60][61][62][63][64][65][66][67][68][69][70][71][72][73][74][75][76][77][78][79][80][81][82][83][84][85][86][87][88][89][90][91][92][93][94][95][96][97][98][99][100][101][102][103][104][105][106][107][108][109][110][111][112][113][114][115][116][117][118][119][120][121][122][123][124][125][126][127][128][129][130][131][132][133][134][135][136][137][138][18,21,36,38,69,72,74,90,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123,124,125,126,127,128,129,130,131,132,133,134,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152,153,154,155,156,157,158,159,160,161,162,163,164,165,166,167,168,169,170,171,172,173,174,175,176,177,178,179,180,181,182,183,184,185,186,187,188,189,190,191,192,193,194,195,196,197,198,199,200,201,202,203,204,205,206,207,208,209,210,211,212]. The interaction of RPA with RAD52 in the alternative HR DSB repair pathway and single-strand annealing is fascinating. Without the RAD52:RPA interaction, the ssDNA handoff from RPA to RAD51 in alternative HR would not be possible, and it is also critical that RPA is phosphorylated to allow this DNA handoff [12][93]. Previous studies identified residues 224-271 on RPA32 and 169-326 on RPA70 that include binding sites for RAD52. It was also found that RAD52 residues 218-303 bind RPA70, along with RPA32 [23][21]. Although the interaction sites have been defined, structural data has yet to be available for the RPA:RAD52 complex.
Table 12. Human proteins that interact with RPA.
| Interacting Protein | Interaction Site on RPA | DNA Metabolism Pathway | Citation |
|---|---|---|---|
| AID | RPA32 | Immunoglobulin diversification | [33][107] |
| * Ajuba | RPA70 | DNA damage response (DDR) | [34][35][108,109] |
| ** ATR/ATRIP | RPA70-F | Checkpoint signaling, DNA repair | [36][37][38][110,111,112] |
| BID | RPA70-F | Replication stress response | [39][113] |
| * BLM | RPA70-A/B | DNA unwinding, resection | [40][41][51][114,115,125] |
| ** BRCA2 | ? | Homologous Recombination (HR) | [31][105] |
| DDX11 | ? | Chromosome segregation | [119][193] |
| ** DNA-PKcs | ? | DNA repair | [22][42][18,116] |
| ** DSS1 | RPA70 | HR | [47][121] |
| * ETAA1 | RPA70-F/RPA32 | ATR activation, repair at stalled replication forks |
[43][44][45][46][117,118,119,120] |
| * EXO5 | RPA70-F | Intrastrand crosslink repair | [107][181] |
| * FANCJ | RPA70 | DNA repair, genome stability | [49][123] |
| * FBH1 | RPA32 | DNA unwinding, resection | [119][193] |
| HELB | RPA70-F | Replication stress response | [105][106][179,180] |
| HERC2 | RPA70 | Replication | [50][51][124,125] |
| HIRA | RPA70-C | Chromatin remodeling | [52][126] |
| Histones H3 and H4 | RPA70-F | Chromatin remodeling | [53][127] |
| * HLTF | RPA70 | Genome stability | [121][122][195,196] |
| HSF1 | RPA70 | Gene expression | [48][122] |
| * Menin | RPA32 | Genome stability | [55][56][129,130] |
| ** MRE11-RAD50-NBS1 | RPA70-F | HR | [57][58][131,132] |
| Nucleolin | RPA14 | Replication (stress) | [59][60][61][133,134,135] |
| NSMF | RPA32 | DDR | [21][101] |
| ** p53 | RPA70-F | HR | [62][63][64][65][66][136,137,138,139,140] |
| * p53BP1 | RPA70/RPA32 | DDR | [32][106] |
| * PALB2 | RPA32 | Recovery of stalled replication forks | [67][141] |
| Papillomavirus E1 | RPA70-A | Replication | [108][109][182,183] |
| Parvovirus NS1 | RPA70/RPA32 | DNA unwinding, resection | [113][187] |
| PCNA | RPA70 | Replication | [117][191] |
| Polδ | RPA70 | Replication | [114][188] |
| Pol-Prim | RPA70-F/A/B | Replication restart, DNA damage tolerance |
[28][129][102,203] |
| ** PP2A | RPA32 | DDR | [130][204] |
| * PRP19/BCAS2 | RPA70-F/C | DNA repair | [70][144] |
| * PTEN | RPA70 | Genome stability | [71][145] |
| * RAD9/RAD1/HUS1 (9-1-1) | RPA70/RPA32 | DDR | [72][146] |
| RAD17 | RPA70-F | DDR, replication stress response | [73][104][131][147,178,205] |
| ** RAD51 | RPA70-A | Recombination | [29][76][77][132][103,150,151,206] |
| ** RAD52 | RPA70-A/B & RPA32-wHLH | DNA repair | [23][25]81][82][133][21[78][79][,6980,152],153,154[,155,156,207] |
| RECQL1 | RPA70 | DNA unwinding | [124][198[125],199] |
| RECQ5β | ? | DNA unwinding | [126][200[127],201] |
| RFC | RPA70 | DNA unwinding | [114][188[115],189] |
| * RFWD3 | RPA32-wHLH | DNA repair | [83][84][121][157,158,195] |
| * RNaseH | ? | Transcription, DNA repair | [85][159] |
| * RNF4 | RPA70 | DNA DSB repair by HR | [128][202] |
| SENP6 | RPA70 | Unperturbed DNA replication | [24][38] |
| SMARCAL1/HARP | RPA32-wHLH | Replication fork restart | [86][87][88][134][160,161,162,208] |
| SV40 Large T antigen | RPA70-A/B & RPA32-wHLH | Replication | [108][109][110][111][112][182,183,184,185,186] |
| * Tipin | RPA32-wHLH | DDR | [90][91][164,165] |
| * UDG | RPA32-wHLH | Base excision repair | [25][92][116][69,166,190] |
| * UNG2 | RPA32-wHLH | Base excision repair | [25][69[92],166] |
| * WRN | RPA70-A/B | DNA unwinding, resection | [41][115[96],170] |
| * XPA | RPA70-A & RPA32-wHLH | Nucleotide excision repair (NER) | [25][92][97][135][136][137][69,166,171,209,210,211] |
| * XPF-ERCC1 | RPA70 | NER | [100][101][102][174,175,176] |
| * XPG | ? | NER | [100][102][138][174,176,212] |
* Involved in DNA repair; ** Involved in DNA DSB repair.
BRCA2 is an essential protein involved in the canonical HR pathway. BRCA2 also completes the DNA handoff, like RAD52, by interacting with RPA. BRCA2 mutations in women can cause familial, early-onset breast cancer, so understanding these mutations and their potential impact on the interaction with RPA will become vital in understanding DNA repair, specifically in cancer cells. A study in 2003 used a cancer-predisposing BRCA2 mutation (Y42C) to investigate its interaction with RPA. They found that the Y42C mutation inhibited the interaction between RPA:BRCA2, showing that the Y42C mutation has biological importance within the human body [139][213]. The BRCA2 interaction with RAD51 has been studied extensively over the years, as BRCA2 promotes RAD51 nucleation on the ssDNA. Still, there has yet to be data on the interaction between BRCA2 and RPA to specifically understand how and where this interaction occurs [140][214].
3. PTMs That Regulate RPA in DNA Metabolism
PTMs are chemical modifications involved in the functional regulation of cellular proteins. RPA relies on PTMs for its proper function in the cell cycle and DNA repair, and the main PTM will be focused here is phosphorylation. Phosphorylation of RPA has been studied for decades, with most studies focused on the RPA32 N-terminal region. Understanding the importance of phosphorylation on other RPA subunits in RPA’s regulation is still unresolved.
Phosphorylation occurs throughout the cell cycle, as previously described. A paper by Yates and coworkers determined two cell cycle checkpoint kinases in yeast, Mec1 and Ddc2 (ATR and ATRIP, respectively, in humans), that are essential for replication stress response and DDR. These two kinases were shown to recruit ssDNA binding to RPA through Ddc2 by phosphorylation. A yeast Ddc2-RPA structure through X-ray crystallography indicated that this interaction is necessary to mediate RPA phosphorylation during DDR [141][215].
An early study in 1990 by Din and colleagues determined the phosphorylation of RPA32 and RPA70 in human and yeast cells by phosphoamino acid analysis using P32 labeling [142][39]. Then, numerous labs focused on the phosphorylation of RPA because of its critical regulation in the DNA repair pathway. In 2003, Binz and others discovered that the phosphorylation of RPA32 modulates RPA-dsDNA interactions and subsequent destabilization [143][22]. They concluded that an intersubunit interaction between phosphorylated RPA32NT and RPA70 N-terminal domain (RPA70N) was possible. Recently, NMR and docking studies were conducted to investigate the interaction between the RPA70 N-terminal domain (RPA70N) and a phosphomimetic N-terminal peptide of RPA32 with candidate serine/threonine sites mutated to aspartic acid. The study showed the possibility that RPA32 N-terminal phosphorylation could allow for transient interaction with RPA70N, which could alter or enhance other interaction sites for proteins or DNA [144][216]. These conclusions indicated a possible role of RPA32 N-terminal phosphorylation binding with RPA70N to form a less disordered structure if these interactions occur in the context of the heterotrimer.
Not only is RPA involved in RPA-DNA interactions, but RPA phosphorylation also influences subcellular localization, especially when there is mitotic phosphorylation at S23 and S29 in RPA32 [145][217]. When the cell has a DDR, RPA becomes hyperphosphorylated [146][147][50,56]. For example, in response to DSBs, RPA is a substrate of ATM kinase that phosphorylates RPA32 at Thr21 and most likely others [148][43]. In 2014, a study was performed on RPA hyperphosphorylation involving the HR pathway in cell cycle phases where HR DSB repair occurs. In these studies, a human squamous cell carcinoma cell line (UM-SCC-38) was synchronized in the S and G2 phases [149][150][151][218,219,220]. Phosphorylation sites in S and G2 phases with and without DSBs were analyzed with western blots using all available antibodies for phosphorylation sites on RPA32. In the G2 phase, chromatin-bound immunoprecipitation phosphorylation occurred at S4/8, S12, T21, S23, and S33 of RPA32 after DSBs.
A global view of RPA heterotrimer phosphorylation was explained using a high-quality RPA70 antibody and isoelectric focusing (IEF) to separate RPA isoforms based on the number of phosphorylations present. Capillary IEF immunoassay data of the phosphorylated RPA heterotrimeric isoforms with and without DSB treatment, synchronized in the G2 phase of the cell cycle, were collected (Figure 12) [9][40]. In this study, data showed that RPA is always a phosphoprotein in the G2 phase by having up to seven phosphates (Figure 12A, blue line). Hyperphosphorylated isoforms generated after DSBs contained up to 13 or 14 phosphates. IEF identified some of these sites with phosphospecific antibodies (Figure 12B–E). The hyperphosphorylated forms with 13–14 phosphates included pS4/8, pT21, and pS33. Several of the sites had yet to be characterized before. DSBs showed increased phosphorylation by PIKK and CDK kinases in S and G2 phases, providing new information on candidate PIKK and CDK sites for future research.
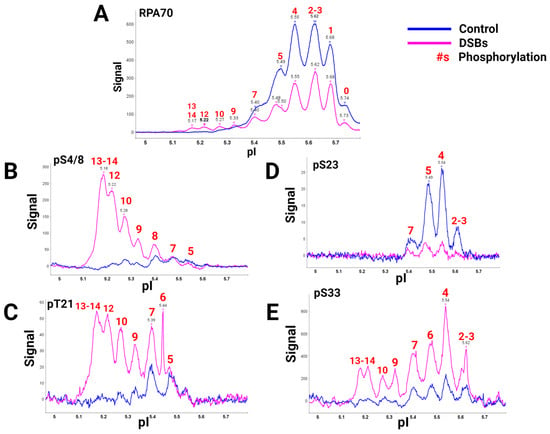
Figure 12. RPA isoforms from cells synced in the G2 phase either with DNA damage (blue) or without DNA damage (pink), separated with isoelectric focusing using a pH gradient from 5-6 and probed with phospho-specific antibodies. RPA heterotrimer probed with (A) anti-RPA70-CT, (B) phosphoRPA32(S4/8), (C) phosphoRPA32(T21), (D) phosphoRPA32(S23), and (E) phosphoRPA32(S33) antibodies. The red number above the corresponding peaks indicates the number of phosphorylations on each isoform. Figure adapted from [9][40].
This research provided pertinent information about RPA heterotrimeric phosphorylation in response to DSBs in the S and G2 phases of the human cell cycle. Several of these phosphorylation sites are PIKK-specific sites in the DNA binding and protein interaction domains of RPA. A summary figure describes possible phosphorylation sites before and after DSBs (Figure 23). This study provided data about phosphatase and kinase action in remodeling RPA isoforms in response to DNA damage [9][40].
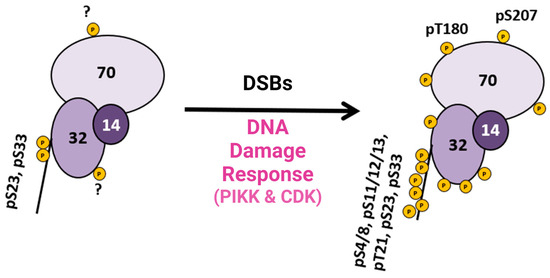
Figure 23. Summary diagram of RPA phosphorylation before and after DSBs. RPA is always phosphorylated (some sites are unknown and indicated with a ?), but after DDR is activated, RPA becomes hyperphosphorylated by the PIKK family of kinases and CDK cell cycle kinases, which phosphorylate RPA. Phosphorylation sites after DNA damage are shown in Figure 1D. Adapted from [9][40].
