The full set of relationship between 61 codons and 3 stop signals that specify the 20 naturals amino acids is called the genetic code. The fundamental role of symmetry in the genetic code is to decrease disorder (entropy) between codons and to preserve the integrity of system during evolution [1,2].
- supersymmetry genetic code
- standard genetic code
- genetic code symmetry
- Introduction
1. Introduction
In 1961 Nirenberg and collaborators deciphered the genetic code, determining experimentally which codons correspond to each of 20 natural amino acids, but without considering codon’s regularity in the form of genetic code table. That challenge was addressed by Nobel laureate F. H. Crick, and in 1968 a solution was published [3][1][2][3] under the name Universal Genetic Code table and readily included in biology and genetics textbooks. In search for symmetry in that genetic code table, the guideline was Watson-Crick A↔T and C↔G base pairing, that was discovered in 1953 for the structure of DNA molecule. This goal was achieved as a half-way result only, and the whole concept of creation of the genetic code table was comparison completed with the random Crick’s “frozen accident hypothesis” as the code inability to accept the new codon variations.
In the meantime, more than thirty different codes have been discovered for genomes of some bacteria and archaea as well as for some organellar mitochondrial and eukaryotic nuclear genomes. Because of that the name of the Universal Genetic Code was changed to the Standard Genetic Code.
The Standard Genetic Code has U-C-A-G horizontal and vertical ordering of bases with only alphabetical symmetry, but no physicochemical symmetry. Therefore, the third base in codons does not differentiate among sixteen codon boxes and were ignored by many studies to this day. The fact that the Standard Genetic Code is degenerate, i.e., that more than one codon can code the same amino acid, the search for symmetries cannot be completely successful by amino acids arrangement. The result was also studied by algebraic approaches to the degeneracy of genetic code, and hypothesis of evolution of the genetic code through progressive symmetry breaking [4,5].
We realized that the key to the genetic code symmetries must be between codon purines (A, G) and pyrimidines (U, C) as the starting point. This approach led us to discovery of the Supersymmetry Genetic Code (SSyGC) table (Figure 1), characterized by codon physicochemical symmetries with the purine – pyrimidine symmetry net as a “golden rule”, which is common for all RNA and DNA species and unchangeable during evolution.
- The Supersymmetry Genetic Code table
2. The Supersymmetry Genetic Code table
On the contrary, with our SSyGC table we have proved that the third base is crucial point for discovery of physicochemical symmetries on the principle of Watson-Crick base pairing (A↔U and C↔G like codon/anticodon) and for discovery of double mirror symmetry. We have also proved that our classification of trinucleotides/codons [2, 6] has the same symmetries, as well as the DNA quadruplets [2].
Our symmetry-based theory of genetic code broadens horizon for understanding evolution. Because of complete physicochemical symmetries of the SSyGC table it is not necessary to involve “the frozen accident”. All more than thirty alternative genetic codes with slight departure from the standard code, can be incorporated in the SSyGC table [2]. Thus, the SSyGC table fulfills all physicochemical criteria on the origin of the genetic code [7,8]. Of special importance is the meaning of complete physicochemical symmetries in the SSyGC table, common for all living RNA and DNA species and unchanged during the whole evolution. In this theoretical approach, there is no evolution of the genetic code, but instead it has a power of natural law in analogy to Nӧther՚s theorem [1][2][3][4][5][6][7][8][9]em [9] for the natural law of energy conservation as a consequence of time symmetry (Ref.) and Einstein՚s paradigm to put symmetries as a dominant concept in the fundamental law of physics to the phenomenon of life.
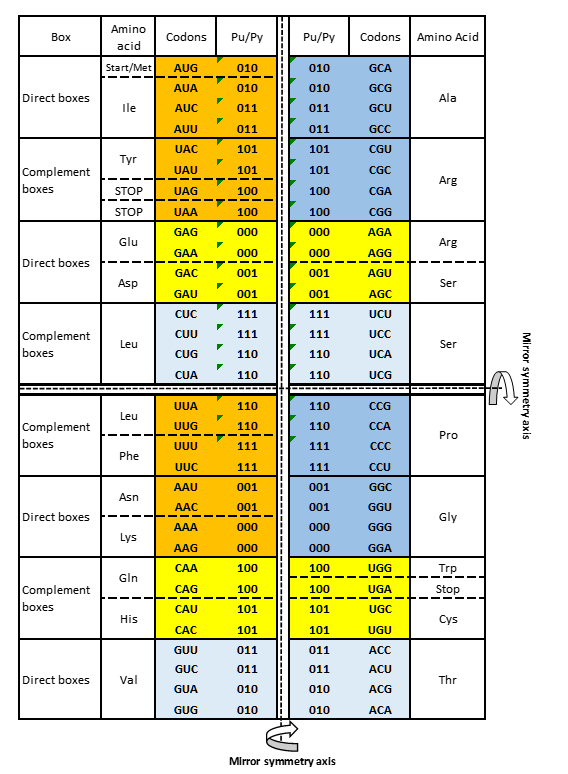
Purine-pyrimidine symmetry net
Figure 1. The Supersymmetry genetic code (SSyGC) table – common for all RNA and DNA species. It has the same distribution of purine/pyrimidine profile in both columns, and simultaneously the same profile distribution pairs of codon rows within each box. There are five symmetries present: 1) purine – pyrimidine symmetry between bases and codons, 2) direct – complement symmetry of codons between boxes, 3) A+U rich and C+G rich symmetry of codons between two columns, 4) symmetrical position of split boxes with codons for more than one amino acid, and complete boxes with codons for only one amino acid. 5) Superior dominant double mirror symmetry as a core symmetry of the SSyGC table is present between all purines and pyrimidines of the whole genetic code. With horizontal and vertical central mirror symmetry axis it created the purine – pyrimidine symmetry net as “the golden rule” for all RNA and DNA species which is unchangeable during evolution.
Purines A and G marked 0, pyrimidines C and U marked 1. 0 pu, purine; 1 py, pyrimidine; dark yellow, two pairs of split boxes with direct – complement symmetry between codons; dark blue, two pairs of no-split boxes with direct – complement symmetry between codons; light yellow, two pairs of split boxes with purine ↔ purine, pyrimidine ↔ pyrimidine transformation between codons; light blue, two pairs of non-split boxes with purine ↔ purine, pyrimidine ↔ pyrimidine transformation between codons. From work Marija Rosandić and Vladimir Paar: Standard Genetic code vs. Supersymmetry Genetic Code – Alphabetical table vs. physicochemical table. Biosystems 2022, 104695 published by ELSEVIER and reproduced by permission of publisher.
Characteristics of the SSyGC table are:
- The SSyGC table starts with AUG start signal.
- All 61codons and three stop signals are arranged in 2x8 boxes, which alternate in each row on the principle of A+T rich and C+G rich codons but with the same purine and pyrimidine ordering.
- Split and complete boxes in the whole supersymmetry code are symmetrically position.
- The ordering of purines and pyrimidines in both columns is identical.
- The ordering of purines and pyrimidines in each two rows is also identical.
- Vertically, between direct and complement boxes, purines, and pyrimidines as well as codons are regularly ordered on the principle of Watson – Crick pairing.
- Horizontally in the same row, purine in the first column transforms in the purine of the second column, and analogously pyrimidine in pyrimidine creating alternately A+T rich and C+G rich codons.
- Left column contains dominant A+T rich codons (24 A+T rich, 8 C+G rich), but second base of all codons is week A and U.
- Right column contains dominant C+G rich codons (24 C+G rich, 8 A+U rich), but second base of all codons is strong C and G.
- Only in the SSyGC table the codons of all amino acids are not scattered, including three sextets for Serine, Arginine and Leucine which codons are positioned in continuity.
- The SSyGC table has a central vertical and horizontal purine-pyrimidine double mirror symmetry according to the mirror symmetry axis.
- Mirror symmetry with horizontal symmetry axis is also present between the second and third bases of codons.
- In this way, the physicochemical unique symmetry net of the whole SSyGC table is structured creating full symmetries between bases, codons, and amino acids.
- The symmetry net is unique and common for all RNA and DNA living species on Earth.
- The symmetry net is common also for more than 30 nuclear and mitochondrial genetic codes which differ from the Standard genetic code table.
- Due to the symmetry net, codons of the SSyGC table directly transform in the DNA molecule with Watson – Crick pairing (32 codons direct and 32 codons their complement) (Rosandić and Paar 2023).
- The SSyGC table with its unique symmetry net has remained unchanged during evolution and has the power of the natural law for the origin of life.
- The protection of the genetic code and DNA molecule symmetries during all of evolution reveal their role in decreasing entropy (disorder) and the preservation of species integrity.
Now, more than 60 years after Nirenberg’s empirical discovery, physicochemical symmetries of the SSyGC table for all RNA and DNA species on Earth are discovered, shedding together with DNA symmetries a new light on evolutionary development of species.
References
- Rosandić, M., Paar, V. Standard Genetic code vs. Supersymmetry Genetic Code–Alphabetical table vs. physicochemical table. BioSystems . 2022, 14, 110748.
- 2. Rosandić, M., Paar, V. The Evolution of Life Is a Road Paved with the DNA Quadruplet symmetry and the Supersymmetry Genetic Code.. Int. J. Mol. Sci .. 2023, 24, 12029.
- F.H.C. Crick; The origin of the genetic code. J. Mol. Biol.. 1968, 38, 367-379.
- G L Findley; A M Findley; S P McGlynn; Symmetry characteristics of the genetic code.. Proc. Natl. Acad. Sci.. 1982, 79, 7061-7065.
- José Eduardo M. Hornos; Yvone M. M. Hornos; Michael Forger; SYMMETRY AND SYMMETRY BREAKING: AN ALGEBRAIC APPROACH TO THE GENETIC CODE. Int. J. Mod. Phys. B. 1999, 13, 2795-2885.
- Rosandić, M.; Paar, V. Classification of Trinucleotides/Codons. Encyclopedia. Available online: https://encyclopedia.pub/entry/47601 (accessed on 02 February 2024).
- Eugene V. Koonin; Artem S. Novozhilov; Origin and evolution of the genetic code: The universal enigma. IUBMB Life. 2009, 61, 99-111.
- Eugene V. Koonin; Artem S. Novozhilov; Origin and Evolution of the Universal Genetic Code. Annu. Rev. Genet.. 2017, 51, 45-62.
- Nöther, E. Invariante Variantionsprobleme.. Nachr. d. König. Gesellsch. d. Wiss. zu Göttingen, Math-2-phys. Klasse, Gesellsch. d. Wiss.. 1918, 1918, 235–257.
