Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 2 by Lindsay Dong and Version 1 by Rosangela Addesso.
The rhizosheath, defined as the complex of root hair, exudates and soil that strongly adheres to plant roots, is a promising root adaptive trait in facing conditions of water and nutrient deficits, as well as acidic soil. Several beneficial ecological functions are attributed to the rhizosheath, such as enhancing water and nutrient uptake; protecting from dehydration, heat and acid stresses; and stimulating microbial activities. It has been described in several Angiosperm species, including crops grown in severe habitats.
- rhizosheath
- sustainable agriculture
- climate change
1. Introduction
The climate crisis is now a subject urgency for humanity, given its negative consequences for agricultural productivity that jeopardize global food provisioning [1,2,3,4,5][1][2][3][4][5]. According to the latest reports by the Intergovernmental Panel on Climate Change (IPCC), in the coming decades, the planet will confront an intensification of extreme climatic events (e.g., drought, heatwaves, heavy rainfall) which will alter the water cycle and its properties, leading to alarming and unprecedented repercussions on hydrological systems and aquifer recharge [6,7][6][7]. Furthermore, increasing temperatures and atmospheric levels of carbon dioxide may entail negative cascade effects influencing the physical, chemical and biological properties of all natural ecosystems of the biosphere, first and foremost the soil, thus limiting the essential ecological functions on which fertility and productivity strongly depend [8,9,10,11][8][9][10][11]. Heavy precipitation can promote soil hypoxia due to water stagnation, as well as nutrient leaching, especially for water-soluble nutrients (e.g., nitrates, sulfates, calcium, magnesium, silicon); on the other hand, drought conditions can lead to soil salinization because of increased water evaporation processes, as well as to a loss of particle cohesion that, in turn, promotes wind erosion [12]. All of this affects microbial communities’ compositions and activities due to the impairment of the soil biogeochemical cycles, compromising plant development through changes in growth and functioning [13,14,15,16][13][14][15][16].
Therefore, urgent and accelerated adaptation actions aimed at climate risk mitigation and at increasing the ecosystem’s resilience, are needed, considering the global demand for food by a growing population and in view of alternative sustainable practices to conventional intensive agriculture [5,17,18][5][17][18]. A rhizosheath is defined as a complex of root hair, exudates and strongly adhering soil. It ranks among the most promising adaptive traits to focus on for the improvement of agricultural sustainability [19]. Despite a recent increase in scientific literature on the rhizosheath, knowledge about the factors involved in its genesis and its ecological functions is still limited; moreover, most of the studies were conducted in controlled laboratory conditions, without validation in the field [20,21,22][20][21][22]. Firstly described in several grass species from arid habitats, and, recently, in plants essential for human and animal consumption (e.g., cereals and legumes), the development of such a particular “cylindrical muff” surrounding the root can be associated with various abiotic and biotic factors. Among them are the soil texture and moisture; the genetic determinants controlling root traits, such as root hairs and their architecture; as well as mucilage secretion and relevant microbial (fungal and bacterial) activities [22,23,24,25,26][22][23][24][25][26]. The rhizosheath plays a crucial role in coping with environmental stresses, protecting the root system against dry and acid soil conditions and improving the uptake of water and nutrients, which is probably related to better contact at the interface between soil and root surfaces [27,28,29,30][27][28][29][30]. Therefore, the adaptation strategy of these particular plants is considered as a trait of great agronomic importance and a potential feature for breeding and management. It is promising for efforts to enhance the sustainability of the crop production and to shift towards the second green revolution [20,31,32][20][31][32].
2. Rhizosheath
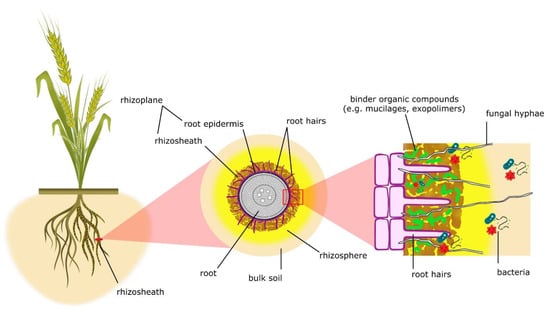
Described for the first time in the 19th century and identified as rhizosheath in the late 1970s, this particular coating of root-enveloping soil has been indicated as an extraordinary drought-adaptive feature of plants [21,33,34][21][33][34]. In detail, the rhizosheath can be described as the root surface and soil clinging to the roots thanks to the physicochemical and biological action of root hairs and the binding organic compounds secreted by microbes and roots, such as mucilages and other exopolymers [29,35,36,37,38][29][35][36][37][38] (Figure 1). According to other definitions, the rhizosheath, when combined with the root surface (root epidermis), forms the rhizoplane. The rhizosphere extends far beyond the rhizosheath’s limits, representing the whole soil volume influenced by plant roots [32,33,39][32][33][39] (Figure 1).
Figure 1. Schematic representation of rhizosheath traits, denoting the several components involved in its formation according to York et al. [33].
The rhizosheath is often operatively defined as the weight of soil that is strongly adhered to the roots after separation from the surrounding soil. Separation techniques include different methods, ranging from shaking to sonication in water. Several works reported in the literature concern a few spontaneous grass species living in extreme arid habitats under water stress, and highlight some of the processes involved in rhizosheath formation and its crucial role in the ecosystem [40,41,42,43,44][40][41][42][43][44]. But recent studies have demonstrated that the rhizosheath is not a specific trait limited to wild species of no agronomic importance; in fact, it is recurrent in several other taxonomic groups among the Angiosperms, both monocotyledons (Poales, Commelinales, etc.) and eudicotyledons (Fabales, Caryophyllales, Brassicales, Solanales, etc.) [22,29,35,45,46,47,48][22][29][35][45][46][47][48]. In view of its beneficial functions on abiotic-stressed plants, rhizosheath have also been investigated in crop species grown in various severe conditions (e.g., drought, low nutrients or acid soil conditions).
3. Factors Involved in Rhizosheath Formation
3.1. Root Hairs
Root hairs are specialized tubular extensions of root epidermal cells, mainly responsible for nutrient and water acquisition, plant anchoring and microbial interactions. They increase the root surface and the access to soil volume [58,59][49][50]. Moreover, they represent the “scaffolding” on which the rhizosheath evolves, providing a physical structure for the stabilization and trapping of soil particles [23,34][23][34]. Therefore, they play a crucial role in rhizosheath formation, since root-hairless species do not present this particular feature, or else they exhibit an underdeveloped rhizosheath, where the chemical adhesive action of exudates is dominant [26,29,60][26][29][51].
Liu et al. [61][52] suggested a highly positive correlation between root hair length and density and the rhizosheath’s dimension and weight in induced dry soil conditions for foxtail millet (Setaria italica L.). They also defined the soil moisture level (10–14% w/w) below which rhizosheath formation is stimulated. Delhaize et al. [27] highlighted a strong and significant relation between root hair length and rhizosheath’s dimensions in wheat (Triticum aestivum L.).
3.2. Root- and Microbial-Derived Mucilages
Soil mucilages are high-molecular-weight organic compounds which are found in soil as a result of the activity of belowground plant organs and microbial production, with several beneficial ecological functions for plants, such as the promotion of nutrients and water uptake, the attraction of favorable microbes and insects and protection against infections [23,62,63,64][23][53][54][55]. Moreover, they promote soil aggregates’ stabilization [65][56] thanks to the adhesion properties related to their nature of viscoelastic gels, which are rich in polysaccharides and glycoproteins, making them the main component in the rhizosheath formation process [23,34,66,67][23][34][57][58]. Together with the extracellular polymeric substances (EPS) secreted by microorganisms, the compounds found in mucilages can affect soil dispersion/flocculation dynamics through compensation for the negative charges on clay; the absorption of organic acid anions; or by influencing the rhizosphere pH, which causes dissolution or precipitation of the binding compounds [68,69][59][60]. Microbial products and root exudates can also influence the mechanical and hydraulic properties of the soil. Mucilages soak the voids between the soil particles, impregnating them; in dry conditions, they increase soil viscosity and strengthen the bond between soil particles and roots, forming a larger and stronger rhizosheath.3.3. Genetics
Plants can respond to environmental abiotic and biotic stresses (e.g., extreme climatic and edaphic conditions, nutrient and water deficiencies, pathogens, etc.) through adaptive molecular strategies controlled by genes and quantitative trait loci (QTLs), allowing them to face hostile conditions. Research in this field is extremely interesting, with a goal of implementing innovative sustainable agricultural practices to allow us to select more tolerant and resilient varieties or to identify relevant and favorable traits for crop breeding. However, at present, only a few studies have highlighted the genetic determinants modulating rhizosheath formation. In response to abiotic stresses, such as dry conditions, acid or P-deficient soil, the upregulation/downregulation of specific genes has been observed with stimulation of the expression of root characteristics and functions, such as root hair development and root exudation [33,44][33][44]. The rhizosheath is a complex, multigenic trait showing a high heritability, with effects differing among species, and the comprehensive understanding of which requires further in-depth analysis [20,33,75][20][33][61].4. Benefits and Ecological Functions of Rhizosheaths
The role of rhizosheaths in plant survival and productive behavior is still controversial and not entirely proven, given that there are many genetic and physiological factors involved in this process that are not easy to discern [34,50][34][62]. Being described for the first time in species from desert habitats, the rhizosheath was intuitively considered an adaptive trait making plants more tolerant of and resilient to dry conditions. They were thought to protect roots from drought and heat stress, increasing water uptake and decreasing dehydration [50,76][62][63]. Over time, various studies highlighted that rhizosheath size was negatively correlated with soil water content and positively correlated to root hair length [27,36,55,61][27][36][52][64]. The greater rhizosheaths that formed in cactus species in dry conditions showed water potential similar to that of root surfaces and higher than that of bulk soil; this proves a reduced level of water loss from the sheathed root epidermis. On the other hand, in wet conditions, the rhizosheath has been shown to improve soil–root contact, favoring root water uptake [77][65]. The rhizosheath helps plants to tolerate soil acidity. Delhaize et al. [27] showed a positive correlation between rhizosheath size and root hair length in wheat grown in acidic soil. Root hair elongation in acidic soil has been associated with improved water and nutrient absorption and a greater tolerance to Al3+ toxicity. This is due to mechanisms regulated by several genes independently of the well-known TaALMT1 gene encoding for the Al3+ tolerance of root hairs. Moreover, the rhizosheath mucilage increases water retention and contributes to creating a microenvironment with stable water content compared to the surrounding soil due to the hydrophilic/hydrophobic behaviors of mucilage [49,69,78][60][66][67].5. Methods for Rhizosheath Investigation
5.1. Rhizosheath Sampling
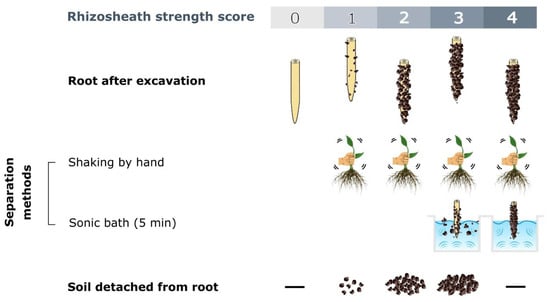
The first cause of misunderstandings may arise from rhizosheath collection methodology, which often affects the results of empirical tests, with a consequent possible underestimation of the rhizosheath’s features. In fact, the lack of a definition of a universal sampling technique may be associated with considerable errors for its basic identification: for instance, erroneous sampling can break the roots, with a consequent loss of the rhizosheath. A major source of ambiguity is the incorrect definition of the soil domain to be sampled around the roots: some works in the literature refer to the rhizosphere, but they are actually conducted on the rhizosheath [29,34][29][34]. Brown et al. [29] developed an efficient method to define the presence/absence and strength scores of rhizosheaths based on different degrees of soil cohesion to roots. These scores range from (0), indicating no attached soil particles; (1), indicating a few that are eliminating after its agitation; (2), indicating attached soil that is detached from the roots by shaking; (3), indicating root-adhering soil that remains after shaking and is partially removed after 5 min in a sonic bath (75 W; 35–45 kHz); and (4), removed the soil remains attached both after shaking and a sonic bath (Figure 2).
Figure 2. Rhizosheath strength score scheme based on the different degrees of soil cohesion to the roots and separation methods, according to Brown et al. [29]. (—) indicates an undetected feature.
5.2. Rhizosheath Quantification
For an initial quantitative characterization of rhizosheath, it should be weighed and oven dried at 105 °C to obtaine the absolute rhizosheath dry weight [24[24][26][37][44][68],26,37,44,52], which can be also calculated as the difference between the root–rhizosheath complex weight and the root weight after being cleaned of the adhering soil in fresh or dry mass conditions [36,51,54,61][36][52][69][70].
The specific rhizosheath weight is given by the ratio between the rhizosheath’s dry mass (root-adhering soil, RAS) and root tissue (RT, RAS/RT, g g−1) after being dried at 65 °C [51,55,82][64][69][71]. However, it can be also calculated as the ratio between the RAS and the total length of the root (RL, RAS/RL, mg cm−1) for each plant. The specific rhizosheath weight allows us to estimate its size [20].
5.3. Genetic Studies
Key genetic determinant studies associated with the expression of rhizosheath characteristics, focused on QTL mapping and gene identification, were conducted on several crop species, among which were cereals (e.g., wheat, pearl barley, millet, rice); others, such as lupine and tomato; and wild relatives [29,44][29][44]. Delhaize et al. [83][72] used multiparent whole-genome analysis (MPWGAIM) in wheat (Triticum aestivum L.) populations grown on non-acid soils, and identified six QTLs located on chromosomes 2B, 4D, 5A, 5B, 6A and 7A. Some of them were probably linked to the basic helix–loop–helix (bHLHs) transcription factor family, influencing the root hair elongation in Arabidopsis and rice and determining the rhizosheath extent. Others were located close to Rht genes influencing the plant structure, such as the height or the root length. On the other hand, five QTLs located on different chromosomes (i.e., 1D, 3A, 3B, 6A2, 7B) contributed to rhizosheath size in wheat (Triticum aestivum L.) grown in acid soil, with a notable improvement in phosphorus acquisition [28]. George et al. [36] found genomic regions in barley (Hordeum vulgare L.) that were significantly associated with rhizosheath weight on the chromosome 2H. They contained a glutamate receptor and several putative candidate genes which modulate the root system development in rice and Arabidopsis in abiotic stress conditions, such as cold and drought, or during the early and delicate plant growth stages. Drought treatment on foxtail millet (Setaria italica) increased the expression of five root-hair-elongation-associated genes (Seita.3G196500, Seita.2G057800, Seita.9G333500, Seita. 8G104600, Seita.7G190800). This was revealed by qRT-PCR analysis demostrating the development of a larger root hairs as appendage to which soil particles bond [42].5.4. Microbial Investigations
The advances in genomic sequencing methods developed during the last decades has allowed us to overcome the lack of information related to non-readily culturable microbes from several environments [85][73]. Through these revolutions, technological applications investigating the composition and the physiology of the rhizobiome can shed light on its key role in the soil ecosystem, unraveling the vital mutualistic interactions among soil, roots and microbes [86][74]. Recent studies have suggested that rhizosheath-associated microbial communities are pivotal in its building processes, as well as in the plant-growth-promoting services provided by root–bacteria relationships. A rhizosheath was defined as an edaphic “mini-oasis” in arid habitats, where several microbial taxa presenting high functional redundancy are in competition to conquer ecological niches that support the beneficial functions of plant biofertilization, biopromotion and bioprotection [79][75]. Root-associated microorganisms can implement several strategies aimed at improving essential nutrient availability; producing biostimulants (exopolysaccharides, phytohormones, volatile compounds, etc.) for plant growth; and at mitigating abiotic/biotic stresses affecting plants [81][76]. Several studies based on bacterial and fungal strain inoculation techniques have demonstrated their particular role in root system development, as well as in rhizosheath evolution. Chen et al. [67][58], in the rhizosheath of Kengyilia hirsute, individuated the enrichment of Massilia and Arthrobacter species that are likely related to plant molecular mechanisms for specific taxa selection and accumulation involved in rhizosheath formation. Trichoderma harzianum T-22 increased the rhizosheath amount in several ancient and modern wheat varieties, affecting their root systems’ architectures differently [51][69]. The endophytic fungus Piriformospora indica is able to modulate auxin production under moderate soil drying, enhancing the growth of rice root hairs for soil exploration by seeking water and providing a more suitable physical structure for the formation of the rhizosheath. This leads to an enrichment of Bacillus cereus in both the rhizosphere and the rhizosheath, suggesting a strong bacteria–fungi interaction involved in the exudate compounds use [52][68].References
- Giorgi, F. Climate Change Hot-Spots. Geophys. Res. Lett. 2006, 33, L08707.
- Hare, W.L.; Cramer, W.; Schaeffer, M.; Battaglini, A.; Jaeger, C.C. Climate Hotspots: Key Vulnerable Regions, Climate Change and Limits to Warming. Reg. Environ. Chang. 2011, 11, 1–13.
- Turco, M.; Palazzi, E.; von Hardenberg, J.; Provenzale, A. Observed Climate Change Hotspots: OBSERVED CLIMATE CHANGE HOTSPOTS. Geophys. Res. Lett. 2015, 42, 3521–3528.
- Islam, S.N.; Winkel, J. Climate Change and Social Inequality; DESA Working Paper 152; Department of Economic & Social Affairs, United Nations: New York, NY, USA, 2017.
- FAO. Strategy on Climate Change 2022–2031; Food and Agriculture Organization of the United Nations: Rome, Italy, 2022.
- IPCC. Climate Change 2021: The physical science basis. In Contribution of Working Group I to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change; Masson-Delmotte, V., Zhai, P., Pirani, A., Connors, S.L., Péan, C., Berger, S., Caud, N., Chen, Y., Goldfarb, L., Gomis, M.I., et al., Eds.; Cambridge University Press: Cambridge, UK, 2021.
- IPCC. Summary for policymakers. In Climate Change 2022: Impacts, Adaptation, and Vulnerability. Contribution of Working Group II to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change; Pörtner, H.-O., Roberts, D.C., Tignor, M., Poloczanska, E.S., Mintenbeck, K., Alegría, A., Craig, M., Langsdorf, S., Löschke, S., Möller, V., et al., Eds.; Cambridge University Press: Cambridge, UK, 2022; Available online: https://report.ipcc.ch/ar6wg2/pdf/IPCC_AR6_WGII_SummaryForPolicymakers.pdf (accessed on 1 April 2022).
- Reynolds, W.D.; Bowman, B.T.; Drury, C.F.; Tan, C.S.; Lu, X. Indicators of Good Soil Physical Quality: Density and Storage Parameters. Geoderma 2002, 110, 131–146.
- Heyder, U.; Schaphoff, S.; Gerten, D.; Lucht, W. Risk of Severe Climate Change Impact on the Terrestrial Biosphere. Environ. Res. Lett. 2011, 6, 034036.
- Farkas, C.; Gelybó, G.; Bakacsi, Z.; Horel, Á.; Hagyó, A.; Dobor, L.; Kása, I.; Tóth, E. Impact of Expected Climate Change on Soil Water Regime under Different Vegetation Conditions. Biologia 2014, 69, 1510–1519.
- Mills, R.T.E.; Gavazov, K.S.; Spiegelberger, T.; Johnson, D.; Buttler, A. Diminished Soil Functions Occur under Simulated Climate Change in a Sup-Alpine Pasture, but Heterotrophic Temperature Sensitivity Indicates Microbial Resilience. Sci. Total Environ. 2014, 473–474, 465–472.
- Mondal, S. Impact of Climate Change on Soil Fertility. In Climate Change and the Microbiome; Choudhary, D.K., Mishra, A., Varma, A., Eds.; Soil Biology; Springer International Publishing: Cham, Switzerland, 2021; Volume 63, pp. 551–569. ISBN 978-3-030-76862-1.
- Schimel, J.; Balser, T.C.; Wallenstein, M. Microbial stress-response physiology and its implications for ecosystem function. Ecology 2007, 88, 1386–1394.
- Gelybó, G.; Tóth, E.; Farkas, C.; Horel, Á.; Kása, I.; Bakacsi, Z. Potential Impacts of Climate Change on Soil Properties. Agrokem. És Talaj. 2018, 67, 121–141.
- Patil, A.; Lamnganbi, M. Impact of Climate Change on Soil Health: A Review. Int. J. Chem. Stud. 2018, 6, 2399–2404.
- Smith, J.L.; Halvorson, J.J.; Bolton, H. Soil Properties and Microbial Activity across a 500m Elevation Gradient in a Semi-Arid Environment. Soil Biol. Biochem. 2002, 34, 1749–1757.
- Altieri, M.A.; Nicholls, C.I.; Henao, A.; Lana, M.A. Agroecology and the Design of Climate Change-Resilient Farming Systems. Agron. Sustain. Dev. 2015, 35, 869–890.
- Hallama, M.; Pekrun, C.; Mayer-Gruner, P.; Uksa, M.; Abdullaeva, Y.; Pilz, S.; Schloter, M.; Lambers, H.; Kandeler, E. The Role of Microbes in the Increase of Organic Phosphorus Availability in the Rhizosheath of Cover Crops. Plant Soil 2022, 476, 353–373.
- Etesami, H. Potential Advantage of Rhizosheath Microbiome, in Contrast to Rhizosphere Microbiome, to Improve Drought Tolerance in Crops. Rhizosphere 2021, 20, 100439.
- Ndour, P.M.S.; Heulin, T.; Achouak, W.; Laplaze, L.; Cournac, L. The Rhizosheath: From Desert Plants Adaptation to Crop Breeding. Plant Soil 2020, 456, 1–13.
- Wang, J.; Ding, Y.; Cao, Y.; Xu, W.; Zhang, Y. Rhizosheath Microbes Induce Root Immune Response under Soil Drying. Plant Signal. Behav. 2021, 16, 1920752.
- Honvault, N.; Houben, D.; Firmin, S.; Meglouli, H.; Laruelle, F.; Fontaine, J.; Lounès-Hadj Sahraoui, A.; Coutu, A.; Lambers, H.; Faucon, M. Interactions between Below-ground Traits and Rhizosheath Fungal and Bacterial Communities for Phosphorus Acquisition. Funct. Ecol. 2021, 35, 1603–1619.
- Watt, M.; McCully, M.E.; Canny, M.J. Formation and Stabilization of Rhizosheaths of Zea mays. (Effect of Soil Water Content). Plant Physiol. 1994, 106, 179–186.
- Haling, R.E.; Brown, L.K.; Bengough, A.G.; Valentine, T.A.; White, P.J.; Young, I.M.; George, T.S. Root Hair Length and Rhizosheath Mass Depend on Soil Porosity, Strength and Water Content in Barley Genotypes. Planta 2014, 239, 643–651.
- Zhang, Y.; Du, H.; Xu, F.; Ding, Y.; Gui, Y.; Zhang, J.; Xu, W. Root-Bacteria Associations Boost Rhizosheath Formation in Moderately Dry Soil through Ethylene Responses. Plant Physiol. 2020, 183, 780–792.
- Burak, E.; Quinton, J.N.; Dodd, I.C. Root Hairs Are the Most Important Root Trait for Rhizosheath Formation of Barley (Hordeum Vulgare), Maize (Zea Mays) and Lotus Japonicus (Gifu). Ann. Bot. 2021, 128, 45–57.
- Delhaize, E.; James, R.A.; Ryan, P.R. Aluminium Tolerance of Root Hairs Underlies Genotypic Differences in Rhizosheath Size of Wheat (Triticum aestivum) Grown on Acid Soil. New Phytol. 2012, 195, 609–619.
- James, R.A.; Weligama, C.; Verbyla, K.; Ryan, P.R.; Rebetzke, G.J.; Rattey, A.; Richardson, A.E.; Delhaize, E. Rhizosheaths on Wheat Grown in Acid Soils: Phosphorus Acquisition Efficiency and Genetic Control. J. Exp. Bot. 2016, 67, 3709–3718.
- Brown, L.K.; George, T.S.; Neugebauer, K.; White, P.J. The Rhizosheath—A Potential Trait for Future Agricultural Sustainability Occurs in Orders throughout the Angiosperms. Plant Soil 2017, 418, 115–128.
- Aslam, M.M.; Karanja, J.K.; Yuan, W.; Zhang, Q.; Zhang, J.; Xu, W. Phosphorus Uptake Is Associated with the Rhizosheath Formation of Mature Cluster Roots in White Lupin under Soil Drying and Phosphorus Deficiency. Plant Physiol. Biochem. 2021, 166, 531–539.
- Lynch, J.P. Roots of the Second Green Revolution. Aust. J. Bot. 2007, 55, 493.
- Zhu, A.-M.; Wu, Q.; Liu, H.-L.; Sun, H.-L.; Han, G.-D. Isolation of Rhizosheath and Analysis of Microbial Community Structure around Roots of Stipa Grandis. Sci. Rep. 2022, 12, 2707.
- York, L.M.; Carminati, A.; Mooney, S.J.; Ritz, K.; Bennett, M.J. The Holistic Rhizosphere: Integrating Zones, Processes, and Semantics in the Soil Influenced by Roots. J. Exp. Bot. 2016, 67, 3629–3643.
- Pang, J.; Ryan, M.H.; Siddique, K.H.M.; Simpson, R.J. Unwrapping the Rhizosheath. Plant Soil 2017, 418, 129–139.
- Bergmann, D.; Zehfus, M.; Zierer, L.; Smith, B.; Gabel, M. Grass Rhizosheaths: Associated Bacterial Communities and Potential for Nitrogen Fixation. W. N. Am. Nat. 2009, 69, 105–114.
- George, T.S.; Brown, L.K.; Ramsay, L.; White, P.J.; Newton, A.C.; Bengough, A.G.; Russell, J.; Thomas, W.T.B. Understanding the Genetic Control and Physiological Traits Associated with Rhizosheath Production by Barley (Hordeum Vulgare). New Phytol. 2014, 203, 195–205.
- Karanja, J.K.; Aslam, M.M.; Qian, Z.; Yankey, R.; Dodd, I.C.; Weifeng, X. Abscisic Acid Mediates Drought-Enhanced Rhizosheath Formation in Tomato. Front. Plant Sci. 2021, 12, 658787.
- Marasco, R.; Fusi, M.; Mosqueira, M.; Booth, J.M.; Rossi, F.; Cardinale, M.; Michoud, G.; Rolli, E.; Mugnai, G.; Vergani, L.; et al. Rhizosheath–Root System Changes Exopolysaccharide Content but Stabilizes Bacterial Community across Contrasting Seasons in a Desert Environment. Environ. Microb. 2022, 17, 14.
- Puente, M.E.; Bashan, Y.; Li, C.Y.; Lebsky, V.K. Microbial Populations and Activities in the Rhizoplane of Rock-Weathering Desert Plants. I. Root Colonization and Weathering of Igneous Rocks. Plant Biol. 2004, 6, 629–642.
- Haling, R.E.; Richardson, A.E.; Culvenor, R.A.; Lambers, H.; Simpson, R.J. Root Morphology, Root-Hair Development and Rhizosheath Formation on Perennial Grass Seedlings Is Influenced by Soil Acidity. Plant Soil 2010, 335, 457–468.
- Haling, R.E.; Simpson, R.J.; Culvenor, R.A.; Lambers, H.; Richardson, A.E. Effect of Soil Acidity, Soil Strength and Macropores on Root Growth and Morphology of Perennial Grass Species Differing in Acid-soil Resistance. Plant Cell Environ. 2011, 34, 444–456.
- Liu, T.-Y.; Chen, M.-X.; Zhang, Y.; Zhu, F.-Y.; Liu, Y.-G.; Tian, Y.; Fernie, A.R.; Ye, N.; Zhang, J. Comparative Metabolite Profiling of Two Switchgrass Ecotypes Reveals Differences in Drought Stress Responses and Rhizosheath Weight. Planta 2019, 250, 1355–1369.
- Marasco, R.; Mosqueira, M.J.; Fusi, M.; Ramond, J.-B.; Merlino, G.; Booth, J.M.; Maggs-Kölling, G.; Cowan, D.A.; Daffonchio, D. Rhizosheath Microbial Community Assembly of Sympatric Desert Speargrasses Is Independent of the Plant Host. Microbiome 2018, 6, 215.
- Aslam, M.M.; Karanja, J.K.; Dodd, I.C.; Waseem, M.; Weifeng, X. Rhizosheath: An Adaptive Root Trait to Improve Plant Tolerance to Phosphorus and Water Deficits? Plant Cell Environ. 2022, 45, 2861–2874.
- Bailey, C.; Scholes, M. Rhizosheath Occurrence in South African Grasses. S. Afr. J. Bot. 1997, 63, 484–490.
- Smith, R.J.; Hopper, S.D.; Shane, M.W. Sand-Binding Roots in Haemodoraceae: Global Survey and Morphology in a Phylogenetic Context. Plant Soil 2011, 348, 453–470.
- Cheraghi, M.; Mousavi, S.M.; Zarebanadkouki, M. Functions of Rhizosheath on Facilitating the Uptake of Water and Nutrients under Drought Stress: A Review. Plant Soil 2023, 491, 1–25.
- Mo, X.; Wang, M.; Zeng, H.; Wang, J. Rhizosheath: Distinct Features and Environmental Functions. Geoderma 2023, 435, 116500.
- Bibikova, T.; Gilroy, S. Root Hair Development. J. Plant Growth Regul. 2002, 21, 383–415.
- Grierson, C.; Nielsen, E.; Ketelaarc, T.; Schiefelbein, J. Root Hairs. Arab. Book 2014, 12, e0172.
- McCully, M.E. ROOTS IN SOIL: Unearthing the Complexities of Roots and Their Rhizospheres. Annu. Rev. Plant. Physiol. Plant. Mol. Biol. 1999, 50, 695–718.
- Liu, T.; Ye, N.; Song, T.; Cao, Y.; Gao, B.; Zhang, D.; Zhu, F.; Chen, M.; Zhang, Y.; Xu, W.; et al. Rhizosheath Formation and Involvement in Foxtail Millet (Setaria italica) Root Growth under Drought Stress. J. Integr. Plant Biol. 2019, 61, 449–462.
- Morel, J.L.; Habib, L.; Plantureux, S.; Guckert, A. Influence of Maize Root Mucilage on Soil Aggregate Stability. Plant Soil 1991, 136, 111–119.
- Czarnes, S.; Hallett, P.D.; Bengough, A.G.; Young, I.M. Root- and Microbial-Derived Mucilages Affect Soil Structure and Water Transport: Mucilages, Soil Structure and Sorptivity. Eur. J. Soil Sci. 2000, 51, 435–443.
- Galloway, A.F.; Akhtar, J.; Marcus, S.E.; Fletcher, N.; Field, K.; Knox, P. Cereal Root Exudates Contain Highly Structurally Complex Polysaccharides with Soil-binding Properties. Plant J. 2020, 103, 1666–1678.
- Di Marsico, A.; Scrano, L.; Labella, R.; Lanzotti, V.; Rossi, R.; Cox, L.; Perniola, M.; Amato, M. Mucilage from fruits/seeds of chia (Salvia hispanica L.) improves soil aggregate stability. Plant Soil 2018, 425, 57–69.
- Ahmed, M.A.; Kroener, E.; Holz, M.; Zarebanadkouki, M.; Carminati, A. Mucilage Exudation Facilitates Root Water Uptake in Dry Soils. Functional Plant Biol. 2014, 41, 1129.
- Chen, Y.; Chen, C.; Zhou, Q.; Hu, J.; Lei, Y.; Liu, W. Specific Rhizobacteria Responsible in the Rhizosheath System of Kengyilia Hirsuta. Front. Plant Sci. 2022, 12, 785971.
- Naveed, M.; Brown, L.K.; Raffan, A.C.; George, T.S.; Bengough, A.G.; Roose, T.; Sinclair, I.; Koebernick, N.; Cooper, L.; Hackett, C.A.; et al. Plant Exudates May Stabilize or Weaken Soil Depending on Species, Origin and Time: Effect of Plant Exudates on Rhizosphere Formation. Eur. J. Soil Sci. 2017, 68, 806–816.
- Kroener, E.; Holz, M.; Zarebanadkouki, M.; Ahmed, M.; Carminati, A. Effects of Mucilage on Rhizosphere Hydraulic Functions Depend on Soil Particle Size. Vadose Zone J. 2018, 17, 1–11.
- Galloway, A.F.; Knox, P.; Krause, K. Sticky Mucilages and Exudates of Plants: Putative Microenvironmental Design Elements with Biotechnological Value. New Phytol. 2020, 225, 1461–1469.
- Basirat, M.; Mousavi, S.M.; Abbaszadeh, S.; Ebrahimi, M.; Zarebanadkouki, M. The Rhizosheath: A Potential Root Trait Helping Plants to Tolerate Drought Stress. Plant Soil 2019, 445, 565–575.
- Young, I.M. Variation in Moisture Contents between Bulk Soil and the Rhizosheath of Wheat (Triticum aestivum L. Cv. Wembley). New Phytol. 1995, 130, 135–139.
- de la Fuente Cantó, C.; Diouf, M.N.; Ndour, P.M.S.; Debieu, M.; Grondin, A.; Passot, S.; Champion, A.; Barrachina, C.; Pratlong, M.; Gantet, P.; et al. Genetic Control of Rhizosheath Formation in Pearl Millet. Sci. Rep. 2022, 12, 9205.
- North, G.B.; Nobel, P.S. Drought-induced changes in soil contact and hydraulic conductivity for roots of Opuntia ficus-indica with and without rhizosheaths. Plant Soil 1997, 191, 249–258.
- Tahir, M.; Mirza, M.S.; Hameed, S.; Dimitrov, M.R.; Smidt, H. Cultivation-Based and Molecular Assessment of Bacterial Diversity in the Rhizosheath of Wheat under Different Crop Rotations. PLoS ONE 2015, 10, e0130030.
- Ahmed, M.A.; Kroener, E.; Benard, P.; Zarebanadkouki, M.; Kaestner, A.; Carminati, A. Drying of Mucilage Causes Water Repellency in the Rhizosphere of Maize: Measurements and Modelling. Plant Soil 2016, 407, 161–171.
- Xu, F.; Liao, H.; Zhang, Y.; Yao, M.; Liu, J.; Sun, L.; Zhang, X.; Yang, J.; Wang, K.; Wang, X.; et al. Coordination of Root Auxin with the Fungus Piriformospora Indica and Bacterium Bacillus Cereus Enhances Rice Rhizosheath Formation under Soil Drying. ISME J. 2022, 16, 801–811.
- Bochicchio, R.; Labella, R.; Vitti, A.; Nuzzaci, M.; Logozzo, G.; Amato, M. Root Morphology, Allometric Relations and Rhizosheath of Ancient and Modern Tetraploid Wheats (Triticum Durum Desf.) in Response to Inoculation with Trichoderma Harzianum T-22. Plants 2022, 11, 159.
- Adu, M.O.; Asare, P.A.; Yawson, D.O.; Ackah, F.K.; Amoah, K.K.; Nyarko, M.A.; Andoh, D.A. Quantifying Variations in Rhizosheath and Root System Phenotypes of Landraces and Improved Varieties of Juvenile Maize. Rhizosphere 2017, 3, 29–39.
- Ndour, P.M.S.; Gueye, M.; Barakat, M.; Ortet, P.; Bertrand-Huleux, M.; Pablo, A.-L.; Dezette, D.; Chapuis-Lardy, L.; Assigbetsé, K.; Kane, N.A.; et al. Pearl Millet Genetic Traits Shape Rhizobacterial Diversity and Modulate Rhizosphere Aggregation. Front. Plant Sci. 2017, 8, 1288.
- Delhaize, E.; Rathjen, T.M.; Cavanagh, C.R. The Genetics of Rhizosheath Size in a Multiparent Mapping Population of Wheat. J. Exp. Bot. 2015, 66, 4527–4536.
- Streit, W.R.; Schmitz, R.A. Metagenomics—The Key to the Uncultured Microbes. Curr. Opin. Microbiol. 2004, 7, 492–498.
- Baldrian, P. The Known and the Unknown in Soil Microbial Ecology. FEMS Microbiol. Ecol. 2019, 95, fiz005.
- Marasco, R.; Fusi, M.; Ramond, J.-B.; Van Goethem, M.W.; Seferji, K.; Maggs-Kölling, G.; Cowan, D.A.; Daffonchio, D. The Plant Rhizosheath–Root Niche Is an Edaphic “Mini-Oasis” in Hyperarid Deserts with Enhanced Microbial Competition. ISME Commun. 2022, 2, 47.
- Pathan, S.I.; Ceccherini, M.T.; Sunseri, F.; Lupini, A. Rhizosphere as Hotspot for Plant-Soil-Microbe Interaction. In Carbon and Nitrogen Cycling in Soil; Datta, R., Meena, R.S., Pathan, S.I., Ceccherini, M.T., Eds.; Springer: Singapore, 2020; pp. 17–43. ISBN 9789811372636.
More
