Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 2 by Lindsay Dong and Version 1 by Jie Cheng.
Microfluidic technology has become an important high-throughput screening technology due to its fast speed, low cost, high automation, and high screening throughput, and it has developed rapidly. Droplet-based microfluidic high-throughput screening has been widely used in various fields, such as strain/enzyme activity screening, pathogen detection, single-cell analysis, drug discovery, and chemical synthesis, and has been widely applied in industries such as those involving materials, food, chemicals, textiles, and biomedicine.
- droplet-based microfluidics
- high-throughput screening
- ultraviolet spectrum
- visible spectrum
- fluorescence spectrum
1. Introduction
Natural microbial strains and enzymes often require evolution or modification. However, the probability of beneficial mutations is very low (<10−5), and conventional screening efficiency is limited by low screening throughput, resulting in high costs and long times for screening a large number of mutants [1]. It is crucial to develop methods for rapid screening of microbial strains and enzyme mutants [1]. With the development of automation equipment and rapid detection methods, various high-throughput screening (HTS) strategies have been established. Therefore, HTS strategies have been widely used in the isolation and screening of high-yielding microbial cell factories [2,3][2][3]. In the screening of industrial microbes, effective target recognition is a key factor, and high-throughput screening based on ultraviolet, visible, and fluorescence spectra in droplet-based microfluidics can greatly enhance target recognition [4,5,6][4][5][6].
Microfluidic technology has emerged as a response to multidisciplinary convergence. It utilizes chemistry, fluid physics, microelectronic materials, nanotechnology, and biotechnology [7]. By manipulating small amounts of fluids in channels at the micrometer or nanometer scale, it brings the screening process onto a chip, achieving automation and intelligence [8,9][8][9]. Moreover, microfluidics combined with fluorescence-activated cell sorting (FACS) systems equipped with flow cytometers (FC) overcome the limitations of traditional FACS. This integration is known as microfluidic FACS [10]. Droplet microfluidics (DM) technology utilizes droplets as microreactor units, performing formation, manipulation, reactions, analysis, and screening operations using microchannels or microstructures. This technology has been widely applied in fields such as drug screening, protein crystallization, evolution, single-cell analysis, enzyme activity detection, and strain screening [11,12][11][12]. Its advantage lies in the ability to encapsulate single cells in droplets, where each droplet serves as an independent reaction system capable of culturing cells and producing metabolites or enzymes. This overcomes the limitations of traditional flow cytometry and fluorescence-activated cell sorting, which are restricted to detecting fluorescent signals directly associated with cells [13,14][13][14]. Fluorescence-activated droplet sorting (FADS) and absorbance-activated droplet sorting (AADS) are typical droplet microfluidic technologies. Microfluidic high-throughput screening technology involves detection, screening, small molecule reactions, and dynamic process analyses. Similar to the FACS principle, the FADS system has the advantage of being able to sort signals within cells, on cells, secreted outside of cells, and in cell-free systems. Additionally, due to its configuration of high-speed cameras, FADS enables the visualization of these sorting processes [13].
Absorption spectroscopy is widely used in colorimetry, protein quantification, enzyme kinetics, and other areas [15]. Compared to FADS, the challenge of AADS detection is that absorbance is directly proportional to the path length, which means that the small volume of droplets and the resulting short optical path length will affect the sensitivity of detection [16]. However, absorption-activated droplet sorting (AADS) is currently approximately 10 times slower than the typical fluorescence-activated droplet sorting (FADS), which means that, compared to FADS, due to throughput limitations, a larger proportion of the sequence space is not accessible. Elliot et al. [17] has improved AADS in order to achieve a sorting speed of kHz. This is primarily achieved using refractive index matching oil and eliminating side scattering to enhance signal quality. Additionally, a sorting algorithm has been implemented to enable sorting at higher frequencies. It has been widely applied in industries such as those involving materials, food, chemical, textile, and biopharmaceuticals (as shown in Figure 1).
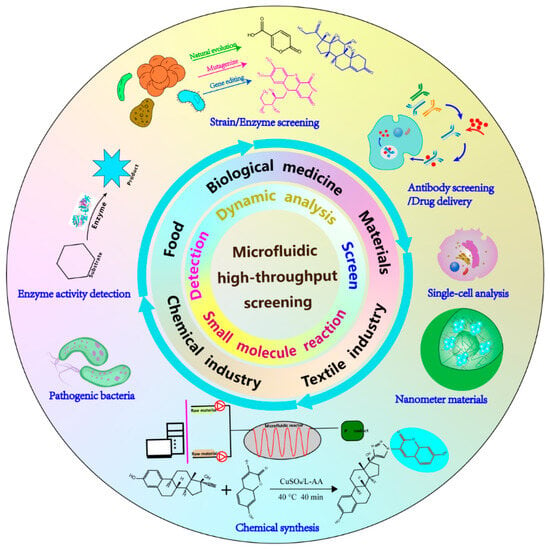
Figure 1.
Application of microfluidic high-throughput screening.
Droplet-based microfluidics plays an important role in directed evolution, enzyme activity detection, and strain screening. The throughput of the screening determines the size of the mutant library, thus determining the possibility of screening high-performance enzymes and strains from large mutant libraries. Through oil encapsulation, the adsorption in the flow channel can be reduced. Droplet encapsulation of individual bacteria allows for the synchronized directional evolution of enzymes during the screening process, making it widely applicable in enzyme-directed evolution.
2. Detection and Screening Based on Ultraviolet Spectroscopy
Generally, the detection of inorganic and organic compounds is based on the Lambert–Beer law, which quantifies the target substance based on the absorption of ultraviolet/visible light due to the structural and color differences of the substance itself. For reactions with specific absorbance, the change in the OD value is caused by the substance itself, enzyme-catalyzed reactions, or absorbent reagents in coupled assays. By measuring the absorbance, products with complex molecular structures such as avilamycin [45][18], cephalosporin C [46][19], lycopene [47][20], and ferulic acid [48][21] can be screened. In the high-throughput screening of strains and enzymes, the direct quantitative identification of metabolic products can be used to screen for beneficial mutants. Mendoza et al. [49][22] directly measured metabolites at a wavelength of 272 nm to identify enzyme activity in the high-throughput screening of lipases with high sn-2 specificity and validated the activity with human pancreatic lipase, LIP2 lipase from Candida antarctica, and lipase from Saccharomyces cerevisiae. For substances which do not have obvious absorbance characteristics, detection can be performed by adding pH indicators, metal ion chelators, enzyme reactions, or chemical reaction coupling methods.
3. Detection and Screening Based on the Visible Light Spectrum
The combination of high-throughput screening strategies based on the visible light spectrum and droplet microfluidic systems is closely related and is mainly used for enzyme activity assays, the screening of enzyme variants, and strain selection [29,30,31][23][24][25]. Similar to the UV spectrum screening strategy, the absorbance of the target substance in the visible light range is used as the screening criterion, and it can be divided into the following steps: direct measurement, addition of pH indicators, addition of metal ion chelating agents, and enzyme-catalyzed reactions. Gielen et al. [54][26] established a coupled droplet microfluidic system for enzyme-directed evolution by embedding optical fibers in the droplet channel. They conducted miniaturized coupled analysis of NAD+-dependent amino acid dehydrogenase and successfully enriched the phenylalanine dehydrogenase activity variant by 2800-fold and identified a mutant with 4.5-times-increased activity in the lysate.4. Detection and Screening Based on Fluorescence Spectroscopy
4.1. Detection and Screening Based on Conventional Strategies of Fluorescence Spectroscopy
Fluorescence analysis has the advantages of high selectivity, sensitivity, and easy integration with chips, making it the most widely used detection method in droplet microfluidic analysis, promoting the development of research fields such as genomics, molecular biology, and synthetic biology. Currently, various industrial microorganisms and enzymes have been screened using fluorescence spectroscopy-based screening strategies. By relying on the specific fluorescence signals of substrates or metabolites, direct screening can be conducted. Wagner et al. [55][27] screened for yeast strains with increased riboflavin production by correlating the secretion level of riboflavin with its yield. The target can emit fluorescence signals by adding fluorescent dyes or metal ion chelators. The majority of screening based on fluorescence spectroscopy relies on enzyme reactions. For direct screening using endogenous enzymes, fluorescence substrates serve as sensitive means to detect enzyme activity, as they do not possess significant fluorescence signals themselves, but the enzymatic reaction products exhibit strong fluorescence signals. Based on the change in fluorescence intensity before and after the reaction, enzyme activity can be quantitatively analyzed (Figure 2A).
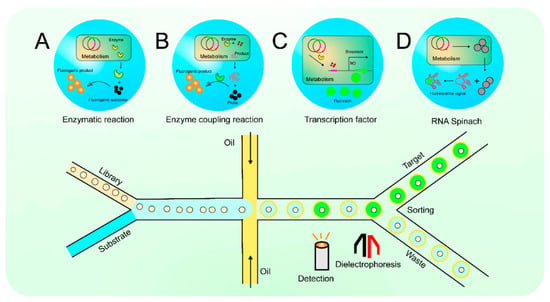
Figure 2. The diagram of droplet microfluidic. Enzymatic reaction (A); Enzyme coupling reaction (B); Transcription factor (C); RNA Spinach (D).
Förster resonance energy transfer (FRET) is a special part of the enzymatic reaction strategy. After the substrate is catalyzed, the fluorescence is activated, and this substrate is usually used as a fluorescent probe. Varadarajan et al. [64][28] designed fluorescence probes and obtained peptide-cleaving enzymes with high selectivity and high catalytic efficiency (Kcat/Km > 104 M−1 s−1).
For methods that involve detecting and screening through the non-enzymatic metabolism of small molecules, it has been observed that many important small molecule metabolites cannot be directly detected through fluorescence, which becomes a bottleneck issue when using droplet microfluidic systems for metabolic pathway and microbial strain screening. The utilization of enzyme-coupled reaction strategies can indirectly convert these small molecules into fluorescent signals, providing an effective approach to address this issue (Figure 2B).
4.2. Detection and Screening Based on Fluorescence Spectroscopy Using Biosensors
Due to the limitations of direct or indirect staining or fluorescence reactions for the detection and screening of target products or key intermediates, the high-throughput screening (HTS) of microorganisms is often challenging. As a complementary and alternative approach, screening strategies using microbial genetic regulatory mechanisms to sense metabolic products have been proposed [70][29]. There are generally two types of biosensors: protein-based biosensors and nucleic acid-based biosensors. Most protein-based biosensors are based on transcription factors (TFs) and engineered fluorescent proteins (FPs). Sun et al. [71][30] divide transcription factor sensors into two categories: endogenous transcription factor biosensors and whole-cell transcription factor biosensors. Various types of TF-based biosensors have been developed, capable of responding to different types of effectors. These biosensors have been widely used in the HTS of metabolic products, including amino acids, organic acids, flavonoids, sugars, and lipids [72,73][31][32] (Figure 2C).
Nucleic acid-based biosensors mainly include RNA riboswitches, RNA spinach [74[33][34],75], and structure-switching DNA biosensors [74,76][33][35]. RNA riboswitches targeting mRNA regulatory regions can detect various metabolites through the RNA aptamer region, thereby regulating the transcription and translation of encoded proteins. Cheng et al. [77][36] used the competitive nature of arginine between arginase and arginine inhibitor to inhibit the expression of eGFP, a molecular sensor, and screened for arginine deamination enzyme mutants with increased activity and substrate affinity. With the rise and development of cell-free synthesis, researchers have found that the traditional genetic characteristics modified and optimized in active cells are no longer applicable to cell-free systems [78,79][37][38]. For example, when transcription factors are used in cell-free systems, they can cause self-transcription effects, response time delays, and an increased translation burden. Ribosome switches have significant advantages, and their construction and screening are particularly important, with a close connection to high-throughput screening using droplet microfluidics [80,81][39][40]. Tabuchi et al. [82][41] were the first to use a droplet microfluidic system to screen for cell-free riboswitches and successfully obtained two types of ribosome switches for histamine, where histamine can either open or close the ribosome switch. RNA Spinach is a newly discovered special structure that produces a stable signal when combined with fluorescent groups. Based on RNA Spinach, biosensors called RNA “drop-in oligonucleotide adaptors” have been developed for the detection of metabolites (Figure 2D).
In the screening system of droplet microfluidics, microdroplets of picoliter (pL) volume as reactors are used to encapsulate enzyme genes, artificially constructed metabolic pathways, gene expression products, etc., which are colored using specific fluorescent probes (Figure 2 top row). In the bottom row of Figure 2, a microfluidic detection/sorting chip for the efficient analysis and sorting of these microdroplets can be seen. Figure 2A shows that screening based on fluorescence spectroscopy is dependent on enzyme reactions. The enzymatic reaction products have a strong fluorescent signal, which is used to quantify the enzyme activity based on the change in fluorescence intensity before and after the reaction [57][42]. Figure 2B shows how small molecules can be indirectly converted to fluorescent signals using an enzyme coupling reaction strategy [68][43]. Figure 2C outlines an assay screening process based on transcription factors (TFs) [73][32]. TFs-based biosensors, which can respond to different types of effectors, are widely used for the HTS of metabolites, including amino acids, organic acids, flavonoids, sugars, and lipids. Figure 2D displays biosensors based on RNA Spinach [53][44]. RNA Spinach is a newly discovered special structure that generates stable signals when combined with fluorescent groups.
5. Detection Screening Based on Other Technologies
As the potential of droplet microfluidic high-throughput screening technology is being explored, researchers have applied it to the screening of more targets. However, they have discovered that some target screenings require the establishment of complex coupling reactions, which may not achieve simplicity and efficiency. Additionally, some target molecules are unable to generate detectable signals. In order to address these issues, label-free detection strategies associated with other detection methods have been established. These methods involve using intrinsic physical or chemical biomarkers for sorting [84][45]. This includes electrochemical detection [85][46], mass spectrometry [86][47], Raman spectroscopy, nuclear magnetic resonance [84][45], Fourier-transform infrared spectroscopy (FTIR), and Fourier-transform near-infrared spectroscopy (FTNIR) [87][48]. Mass spectrometry is not only used for qualitative and quantitative analyses based on the charge-to-mass ratio of the detected substances but can also detect various fragmented substances. This has attracted researchers to combine it with droplet microfluidic technology. However, there are significant barriers between microfluidics and electrospray ionization mass spectrometry. Similar to capillary electrophoresis, the main barrier lies in the interference of surfactants in the electrospray process, which can significantly inhibit ionization efficiency and contaminate the mass spectrometer [88][49]. Surfactants are essential in droplet-based microfluidic systems. They facilitate droplet formation, stabilize droplets, and provide a more important role in creating a biocompatible environment for reactions within the droplets. Therefore, it is necessary to design and synthesize compatible novel surfactants [89][50]. Raman spectroscopy is a detection method that relies on the Raman effect, offering advantages such as rapid and real-time analysis. The discovery of surface-enhanced Raman scattering (SERS) has significantly enhanced the sensitivity of Raman detection. Research has shown that droplet microfluidic technology can effectively overcome the limitations of SERS. Nuclear magnetic resonance (NMR) is a technique for directly analyzing the structure of compounds, and its greatest advantage is its completely non-invasive nature. However, combining NMR with droplet microfluidics is more challenging due to several technical barriers. Unlike fluorescence, NMR detection is based on the principle of magnetic field detection, which requires a high level of uniformity and stability in the droplet samples. The sensitivity of conventional NMR is still insufficient for the microliter- to picoliter-sized samples generated by droplet microfluidic chips [81][40].6. Conclusions
In recent years, droplet microfluidics has gradually moved from the field of chemistry to biology. With the advancement of droplet microfluidic technology and the creation of various detection strategies, an important step has been taken towards the discovery of strains/enzymes with new or improved functions. Especially in the development of synthetic biology, the detection of natural bioactive compounds, the directed evolution of strains/enzymes, and the screening and identification of enzyme semi-rational or rational designs are intricately intertwined with droplet microfluidics. With its advantages of low cost, speed, high automation, and detection sensitivity, it has replaced traditional well-plate screening and regular flow cytometry, becoming the preferred high-throughput screening technology in the field of synthetic biology. Among the high-throughput screening strategies based on UV, visible, and fluorescence spectra, the fluorescence detection strategy has the widest application and is the most mature, serving as a crucial means for the industrialization and commercialization of strain/enzyme screening. UV–visible light detection is known for its simplicity but has limitations and limited applications. Biosensors are an important branch of fluorescence detection. Their essence lies in enhancing fluorescence intensity or overcoming the difficulties of generating fluorescence signals, thus compensating for the shortcomings of conventional strategies. Enzymes are the most important biocatalysts in nature and have been applied in various fields after thousands of years of natural evolution. However, natural enzymes often cannot meet the requirements of industry, making artificial selection and screening increasingly important. Directed evolution has been a key strategy for generating enzymes with desired characteristics such as high selectivity, but experimental barriers and the cost of analyzing large mutant libraries have limited these efforts. However, the biggest problem faced by directed evolution is that traditional microplate screening methods cannot meet the demands of high-capacity library screening. Meanwhile, the industrial and pharmaceutical sectors continue to have a fast-growing demand for novel and improved microbial catalysts. Therefore, the directed evolution of enzymes and the discovery of new enzymes have directly benefited from the development of droplet microfluidics, which provide individual reaction environments and larger screening capacities for enzyme reactions. Enzymes that may require directed evolution in the future include esterases, cellulases, glucose dehydrogenases, and plastic-degrading enzymes.References
- Moragues, T.; Arguijo, D.; Beneyton, T.; Modavi, C.; Simutis, K.; Abate, A.R.; Baret, J.C.; DeMello, A.J.; Densmore, D.; Griffiths, A.D. Droplet-based microfluidics. Nat. Rev. Methods Primers 2023, 3, 32.
- Chiu, F.; Stavrakis, S. High-throughput droplet-based microfluidics for directed evolution of enzymes. Electrophoresis 2019, 40, 2860–2872.
- Jiang, J.; Yang, G.; Ma, F. Fluorescence coupling strategies in fluorescence-activated droplet sorting (FADS) for ultrahigh-throughput screening of enzymes, metabolites, and antibodies. Biotechnol. Adv. 2023, 66, 108173.
- Muretta, J.M.; Rajasekaran, D.; Blat, Y.; Little, S.; Myers, M.; Nair, C.; Burdekin, B.; Yuen, S.L.; Jimenez, N.; Guhathakurta, P.; et al. HTS driven by fluorescence lifetime detection of FRET identifies activators and inhibitors of cardiac myosin. SLAS Discov. Adv. Life Sci. R D 2023, 28, 223–232.
- Sesen, M.; Alan, T.; Neild, A. Droplet control technologies for microfluidic high throughput screening (μHTS). Lab Chip 2017, 17, 2372–2394.
- Rao, C.; Huisman, D.H.; Vieira, H.M.; Frodyma, D.E.; Neilsen, B.K.; Chakraborty, B.; Hight, S.K.; White, M.A.; Fisher, K.W.; Lewis, R.E. A Gene Expression High-Throughput Screen (GE-HTS) for Coordinated Detection of Functionally Similar Effectors in Cancer. Cancers 2020, 12, 3143.
- Yang, D.; Yu, Z.; Zheng, M.; Yang, W.; Liu, Z.; Zhou, J.; Huang, L. Artificial intelligence-accelerated high-throughput screening of antibiotic combinations on a microfluidic combinatorial droplet system. Lab Chip 2023, 23, 3961–3977.
- Sackmann, E.K.; Fulton, A.L.; Beebe, D.J. The present and future role of microfluidics in biomedical research. Nature 2014, 507, 181–189.
- Miansari, M.; Friend, J.R. Acoustic Nanofluidics via Room—Temperature Lithium Niobate Bonding: A Platform for Actuation and Manipulation of Nanoconfined Fluids and Particles. Adv. Funct. Mater. 2016, 26, 7861–7872.
- Zhang, B.; Wu, W.; Zhao, Q.; Yan, S. Geometric optimization of double layered microchannel with grooves array for enabling nanoparticle manipulation. Phys. Fluids 2023, 35, 062009.
- Zeng, W.; Guo, L.; Xu, S.; Chen, J.; Zhou, J. High-Throughput Screening Technology in Industrial Biotechnology. Trends Biotechnol. 2020, 38, 888–906.
- Chou, W.-L.; Lee, P.-Y.; Yang, C.-L.; Huang, W.-Y.; Lin, Y.-S. Recent Advances in Applications of Droplet Microfluidics. Micromachines 2015, 6, 1249–1271.
- Amirifar, L.; Besanjideh, M.; Nasiri, R.; Shamloo, A.; Nasrollahi, F.; de Barros, N.R.; Davoodi, E.; Erdem, A.; Mahmoodi, M.; Hosseini, V.; et al. Droplet-based microfluidics in biomedical applications. Biofabrication 2022, 14, 022001.
- Weitong, Q.; Guangyu, Y. Research and application progress of microdroplets high throughput screening methods. Synth. Biol. 2023, 4, 966–979.
- Sun, G.; Qu, L.; Azi, F.; Liu, Y.; Li, J.; Lv, X.; Du, G.; Chen, J.; Chen, C.H.; Liu, L. Recent progress in high-throughput droplet screening and sorting for bioanalysis. Biosens. Bioelectron. 2023, 225, 115107.
- Hansen, S.K.; Jamali, B.; Hubbuch, J. Selective high throughput protein quantification based on UV absorption spectra. Biotechnol. Bioeng. 2013, 110, 448–460.
- Duncombe, T.A.; Ponti, A.; Seebeck, F.P.; Dittrich, P.S. UV-Vis Spectra-Activated Droplet Sorting for Label-Free Chemical Identification and Collection of Droplets. Anal. Chem. 2021, 93, 13008–13013.
- Tu, R.; Martinez, R.; Prodanovic, R.; Klein, M.; Schwaneberg, U. A Flow Cytometry–Based Screening System for Directed Evolution of Proteases. SLAS Discov. 2011, 16, 285–294.
- Agresti, J.J.; Antipov, E.; Abate, A.R.; Ahn, K.; Rowat, A.C.; Baret, J.C.; Marquez, M.; Klibanov, A.M.; Griffiths, A.D.; Weitz, D. A Ultrahigh-throughput screening in drop-based microfluidics for directed evolution. Proc. Natl. Acad. Sci. USA 2010, 107, 4004–4009.
- Turner, P.; Mamo, G.; Karlsson, E.N. Potential and utilization of thermophiles and thermostable enzymes in biorefining. Microb. Cell Factories 2007, 6, 9.
- Nakkharat, P.; Haltrich, D. Purification and characterisation of an intracellular enzyme with beta-glucosidase and beta-galactosidase activity from the thermophilic fungus Talaromyces thermophilus CBS 236.58. J. Biotechnol. 2006, 123, 304–313.
- Huebner, A.; Olguin, L.F.; Bratton, D.; Whyte, G.; Huck, W.T.; de Mello, A.J.; Edel, J.B.; Abell, C.; Hollfelder, F. Development of quantitative cell-based enzyme assays in microdroplets. Anal. Chem. 2008, 80, 3890–3896.
- Zhang, W.; Fu, J.; Wang, Y.; Zhang, X.; Li, J. Enhanced visible-light photocatalytic activity of ZnS/BiOBr/graphene oxide ternary composite. J. Phys. Chem. Solids 2019, 127, 19–27.
- Wang, Q.; Feng, L.R.; Wei, L.; Li, H.G.; Wang, L.; Zhou, Y.; Yu, X.B. Mutation Breeding of Lycopene-Producing Strain Blakeslea Trispora by a Novel Atmospheric and Room Temperature Plasma (ARTP). Appl. Biochem. Biotechnol. 2014, 174, 452–460.
- Zhou, S.; Liu, P.; Chen, J.; Du, G.; Li, H.; Zhou, J. Characterization of mutants of a tyrosine ammonia-lyase from Rhodotorula glutinis. Appl. Microbiol. Biotechnol. 2016, 100, 10443–10452.
- Abalde-Cela, S.; Gould, A.; Liu, X.; Kazamia, E.; Smith, A.G.; Abell, C. High-throughput detection of ethanol-producing cyanobacteria in a microdroplet platform. J. R. Soc. Interface 2015, 12, 20150216.
- Ostafe, R.; Prodanović, R.; Lloyd Ung, W.; Weitz, D.A.; Fischer, R.J.B. A high-throughput cellulase screening system based on droplet microfluidics. Biomicrofluidics 2014, 8, 041102.
- Cheng, F.; Kardashliev, T.; Pitzler, C.; Shehzad, A.; Lue, H.; Bernhagen, J.; Zhu, L.; Schwaneberg, U. A Competitive Flow Cytometry Screening System for Directed Evolution of Therapeutic Enzyme. ACS Synth. Biol. 2015, 4, 768–775.
- Vallejo, D.; Nikoomanzar, A.; Paegel, B.M.; Chaput, J.C. Fluorescence-Activated Droplet Sorting for Single-Cell Directed Evolution. ACS Synth. Biol. 2019, 8, 1430–1440.
- Fu, X.; Zhang, Y.; Xu, Q.; Sun, X.; Meng, F. Recent Advances on Sorting Methods of High-Throughput Droplet-Based Microfluidics in Enzyme Directed Evolution. Front. Chem. 2021, 6, 666867.
- Gu, S.; Lu, Y.; Ding, Y.; Li, L.; Zhang, F.; Wu, Q. Droplet-based microfluidics for dose–response assay of enzyme inhibitors by electrochemical method. Anal. Chim. Acta 2013, 796, 68–74.
- Gasilova, N.; Yu, Q.; Qiao, L.; Girault, H.H. On-chip spyhole mass spectrometry for droplet-based microfluidics. Angew. Chem. (Int. Ed. Engl.) 2014, 53, 4408–4412.
- Pullagura, B.K.; Amarapalli, S.; Gundabala, V. Coupling electrohydrodynamics with photopolymerization for microfluidics-based generation of polyethylene glycol diacrylate (PEGDA) microparticles and hydrogels. Colloids Surf. A Physicochem. Eng. Asp. 2021, 608, 125586.
- Goto, H.; Kanai, Y.; Yotsui, A.; Shimokihara, S.; Shitara, S.; Oyobiki, R.; Fujiwara, K.; Watanabe, T.; Einaga, Y.; Matsumoto, Y.; et al. Microfluidic screening system based on boron-doped diamond electrodes and dielectrophoretic sorting for directed evolution of NAD(P)-dependent oxidoreductases. Lab Chip 2020, 20, 852–861.
- Norris, J.L.; Porter, N.A.; Caprioli, R.M. Mass spectrometry of intracellular and membrane proteins using cleavable detergents. Anal. Chem. 2003, 75, 6642–6647.
- Heinemann, J.; Deng, K.; Shih, S.C.C.; Gao, J.; Adams, P.D.; Singh, A.K.; Northen, T.R. On-chip integration of droplet microfluidics and nanostructure-initiator mass spectrometry for enzyme screening. Lab Chip 2017, 17, 323–331.
- Holland-Moritz, D.A.; Wismer, M.K.; Mann, B.F.; Farasat, I.; Devine, P.; Guetschow, E.D.; Mangion, I.; Welch, C.J.; Moore, J.C.; Sun, S.; et al. Mass Activated Droplet Sorting (MADS) Enables High-Throughput Screening of Enzymatic Reactions at Nanoliter Scale. Angew. Chem. (Int. Ed. Engl.) 2020, 59, 4470–4477.
- Willner, M.R.; McMillan, K.S.; Graham, D.; Vikesland, P.J.; Zagnoni, M. Surface-Enhanced Raman Scattering Based Microfluidics for Single-Cell Analysis. Anal. Chem. 2018, 90, 12004–12010.
- Wang, X.; Xin, Y.; Ren, L.; Sun, Z.; Zhu, P.; Ji, Y.; Li, C.; Xu, J.; Ma, B. Positive dielectrophoresis-based Raman-activated droplet sorting for culture-free and label-free screening of enzyme function in vivo. Sci. Adv. 2020, 6, eabb3521.
- Liu, W.W.; Zhu, Y. “Development and application of analytical detection techniques for droplet-based microfluidics”—A review. Anal. Chim. Acta 2020, 1113, 66–84.
- Swyer, I.; Soong, R.; Dryden, M.D.M.; Fey, M.; Maas, W.E.; Simpson, A.; Wheeler, A.R. Interfacing digital microfluidics with high-field nuclear magnetic resonance spectroscopy. Lab Chip 2016, 16, 4424–4435.
- Sun, Q.M.; Lu, Y.L.; Shen, X.L.; Sun, X.X.; Wang, J.; Yuan, Q.P. Fluorescence detection-based high-throughput screening systems and devices facilitate cell factories construction. Synth. Biol. 2023, 4, 947–965.
- Shin, J.; Noireaux, V. An E. coli cell-free expression toolbox: Application to synthetic gene circuits and artificial cells. ACS Synth. Biol. 2012, 1, 29–41.
- Hammar, P.; Angermayr, S.A.; Sjostrom, S.L.; van der Meer, J.; Hellingwerf, K.J.; Hudson, E.P.; Joensson, H.N. Single-cell screening of photosynthetic growth and lactate production by cyanobacteria. Biotechnol. Biofuels 2015, 8, 193.
- Choi, K.; Mudrik, J.M.; Wheeler, A.R. A guiding light: Spectroscopy on digital microfluidic devices using in-plane optical fibre waveguides. Anal. Bioanal. Chem. 2015, 407, 7467–7475.
- Qiao, Y.; Zhao, X.; Zhu, J.; Tu, R.; Dong, L.; Wang, L.; Dong, Z.; Wang, Q.; Du, W. Fluorescence-activated droplet sorting of lipolytic microorganisms using a compact optical system. Lab Chip 2017, 18, 190–196.
- Xu, J.G.; Huang, M.S.; Wang, H.F.; Fang, Q. Forming a Large-Scale Droplet Array in a Microcage Array Chip for High-Throughput Screening. Anal. Chem. 2019, 91, 10757–10763.
- Qiao, Y.; Hu, R.; Chen, D.; Wang, L.; Wang, Z.; Yu, H.; Fu, Y.; Li, C.; Dong, Z.; Weng, Y.; et al. Fluorescence-activated droplet sorting of PET degrading microorganisms. J. Hazard Mater. 2022, 424 Pt B, 127417.
- Zhang, G.; Chen, Y.; Li, Q.; Zhou, J.; Li, J.; Du, G. Growth-coupled evolution and high-throughput screening assisted rapid enhancement for amylase-producing Bacillus licheniformis. Bioresour. Technol. 2021, 337, 125467.
- Prodanovic, R.; Ung, W.L.; Durdic, K.I.; Fischer, R.; Weitz, D.A.; Ostafe, R. A High-Throughput Screening System Based on Droplet Microfluidics for Glucose Oxidase Gene Libraries. Molecules 2020, 25, 2418.
More
