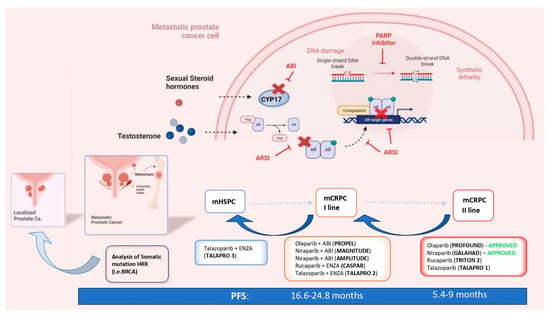
2. The Rationale and the Genetic Profiles That Sustain PARP Inhibitors in PC
To address the underlying issue of genomic erosion, a complex network of DDR mechanisms has evolved
[9][10][9,10]. Genetic abnormalities in DDR response systems are associated with life-threatening diseases such as immune deficiency, premature aging and cancer susceptibility. The following are the principal DNA repair pathways: single-strand break repair (SSR), mismatch repair (MMR), base excision repair (BER), homologous recombination repair (HRR), non-homologous end joining (NHEJ) and nucleotide excision repair (NER). With a single-strand break in DNA, the second intact strand represents a template for BER and NER repair. A double-strand break (DSB) activates the HRR, MMR and NHEJ mechanisms
[11][12][11,12]PARP is a multifunctional protein that plays a critical role in SSR and BER mechanisms, primarily by recruiting DNA repair proteins to the sites of damage. So far, eighteen members of the PARP family have been found, among which PARP-1 is the most important
[13]. PARP’s main enzymatic function is to add ADP-ribose to substrate proteins by cleaving NAD
+ and releasing nicotinamide, thus activating a complex cascade, which ends in the recruitment of different proteins, such as DNA polymerase theta (POLQ), DNA ligase I and XRCC1, to the site of damage. Despite being well-known for its role in SSR and BER, recent evidence suggests that PARP can also affect DSB repair, mainly by controlling the expression of the key HR genes BRCA1 and RAD51
[14][15][16][14,15,16].
Defects in the DDR pathways can result in genomic instability and gene mutations, and, eventually, lead to the development of cancer. On the other hand, germline or acquired mutations in DDR also provide cancer-specific vulnerabilities that can be targeted by synthetic lethality-based therapies
[17]. PARPis are the first successful example of a targeted therapy that uses synthetic lethality to kill cancerous cells with DNA-repairing deficiencies (e.g., BRCA1/2 mutation). These drugs act by competing with NAD
+ for the catalytically active sites of PARP molecules, interfering with SSB repair by BER. The unrepaired SSB can be converted to DSB, and the primary mechanism in the repair of such lesions during the cell cycle is HRR. While HRR-proficient cells can repair the DSB originating from the SSB, thus ensuring genetic stability and cell survival, HRR-deficient cells are unable to do so, resulting in apoptosis and, ultimately, cell death
[16][17][18][16,17,18]. Olaparib was the first drug to be developed in this group, initially approved in 2014 for patients with ovarian cancer with germline BRCA PVs and subsequently for those with breast, pancreatic and prostate cancer
[19].
Germline HRR PVs increase the risk of developing PC by eight times at the age of 65 years and are linked to more aggressive PC, a higher risk of lymph-node invasion and distant metastasis at the time of diagnosis
[20]. Moreover, germline BRCA 1/2 PVs are associated with a higher probability of failure in patients with localized disease undergoing active surveillance and a higher risk of recurrence in patients who have undergone curative treatment
[21][22][21,22]. Several studies have analyzed the prevalence of germline and somatic mutations in HRR genes, both in metastatic and localized PC. A 2019 systematic review by Lang et al.
[23] found a median prevalence of germline HRR PVs, and specifically of BRCA2 PVs, in patients with familiar PC rates of 29.3% (range, 7.3–91.67%) and 3.7% (range, 1.3–7.9%), respectively. In the same study, for unselected patients, the frequencies of somatic and germline mutations were, respectively, 3.9% and 1.5% for ATM, 1.1% and 0.6% for BRCA1, 4.9% and 1.1% for BRCA2, 1.3% and 0.5% for PALB2 and 1.5% and 0.5% for RAD51C. The overall prevalence of somatic DDR gene PVs ranged between 4.9 and 22%, while the rate of germline DDR PVs was 17.2–19%.
Regarding metastatic PC, the incidence of germline PVs in HRR genes was found to be between 11% and 33% in a pan-cancer analysis of whole genomes
[23]. Similarly, 8% of germline and 23% of somatic HRR PVs were found in 150 metastatic PCs by the International Stand Up to Cancer/Prostate Cancer Foundation team (SU2C-PCF)
[24]. In this cohort, BRCA2 was the most prevalent mutation (13%) followed by ATM (7.3%), MSH2 (2%) and BRCA1. More specifically, in mCRPC, the incidence of somatic HRR PVs was 24% (BRCA 13%, ATM 7.3%, MSH2 2%, BRCA1 0.3%) in a study by Eeles et al.
[25] and 28% in the Profound study, which analyzed the results of 2792 biopsies of mCRPC patients.
Considering localized PC, the rates of PVs in HRR seem to be lower than those seen in mPC. Despite this, a 2019 study by Kim et al.
[26] found an overall incidence of HRR pathway alteration in localized PC of 29.9%, higher than that suggested by a previous study by Marshall et al.
[27] (11% in Gleason Grade Group 5 and 15.8% in cT3 patients).
The Profound study represents the biggest resource of tissue samples, with a total of 4858 centrally tested
[28]. Next-Generation Sequencing (NGS) was performed mainly on primary tumor samples (83%). It is relevant to underline that of these primary samples, 96% were archival and 4% newly obtained. Samples from metastases were obtained in 17% of cases, and again, most were archival (60% archival and 33% newly obtained). This sample was found to be suitable for testing at similar percentages between the newly obtained and archival samples (63.9% vs. 56.9%) and between the metastatic and primary samples (63.9% vs. 56.2%). On the contrary, the NGS results significantly declined with increasing sample age.
The somatic analysis of HRR mutations in PC should be used instead of germline analysis, when possible. A biopsy of metastatic lesions is the ideal method to identify the molecular changes that occur during disease progression. Despite this, biopsies of metastatic sites are challenging and not always feasible, and results from a single site may underestimate tumor heterogeneity. In the future, the analysis of free circulating DNA (cDNA) might overcome the issues associated with tissue biopsy, such as invasiveness, the need for anesthesia and pain. The first study to analyze cDNA in this field was the GHALAND study
[29], where an analysis of treatment efficacy was performed on the basis of the amount of circulating tumor cells present from the eighth week of treatment. The BRCA cohort obtained better results when compared to the non-BRCA cohort, with a 24% CTC (circulating tumor cell) response. However, incongruences with the different commercial tests currently available for the analysis of cDNA are as high as 40%, with the risk of patients receiving inadequate or no treatment.