Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 1 by Sotiria D. Psoma and Version 2 by Conner Chen.
Biosensor technology is a multidisciplinary field where biology, engineering and nanotechnology promise solutions for healthcare challenges enabling personalised medicine for disease prognosis, diagnosis and drug delivery. The ability to use a point-of-care sensor to measure insulin concurrently with glucose would allow for a much better assessment of endogenous insulin activity, enabling real-time adjustments in insulin dosing to be made while minimising the likelihood of occurrence of extremes of hypoglycaemia or hyperglycaemia.
- insulin
- biosensor
- diabetes mellitus
1. Biosensor Technology
Biosensor technology is a multidisciplinary field where biology, engineering and nanotechnology promise solutions for healthcare challenges enabling personalised medicine for disease prognosis, diagnosis and drug delivery. It combines cutting-edge technology with integrated platforms of microfluidics necessary for analyte delivery and real-time monitoring, and bioelectronics for fast and automated signal processing and analysis to enable decision making [1][48]. Typical microfluidic platforms are formed of polymers such as polydimethylsiloxane (PDMS), silicon or glass, and integrated micropumps. The fast evolution of lab-on-a-chip (LOC) technology where microfluidics, different detection methods and nanomaterials are integrated offers capabilities for multi-analyte, reliable and cost-effective point-of-care diagnostic platforms/biosensors with enhanced sensitivity and lower detection limits compared to conventional methods [2][49]. In addition, recent advances in 3D-printing technologies offer a number of advantages in biosensor production such as higher resolution and low cost, and enable end-user customisation and prototyping [3][50]. Further miniaturisation of bioelectronics and their placement close or under the microfluidic channels where reactions take place can significantly contribute to overcoming issues of lag-time in detection by the biosensor, which is important for closed-automated-loop insulin delivery systems [4][51]. These developments also support the commercialisation pipeline of innovative wearable biosensors.
The ability to use a point-of-care sensor to measure insulin concurrently with glucose would allow for a much better assessment of endogenous insulin activity, enabling real-time adjustments in insulin dosing to be made while minimising the likelihood of occurrence of extremes of hypoglycaemia or hyperglycaemia. A point-of-care biosensor would be tailored to the needs of an individual allowing for diet and exercise patterns to be incorporated and applied into their insulin dosing plan. In recent decades, affinity biosensors based on antibodies, nucleic acid aptamers and molecularly imprinted polymers (MIPs) have emerged as powerful and promising point-of-care diagnostic devices. Particularly, electrochemical and optical affinity biosensors have attracted attention [5][52]. In the literature, most of the insulin biosensors are based on aptamers and molecularly imprinted polymers (MIPs), which imitate or improve on antibody-based biosensors [6][7][8][9][10][11][53,54,55,56,57,58]. They offer portability, fast response, selectivity, specificity, high stability and reproducibility, low detection limits and low production costs.
2. Aptamer-Based Insulin Biosensors
Aptamers are single-stranded nucleic acids that can form secondary and tertiary structures, and like antibodies, they can recognise proteins specifically and with high affinity. However, unlike antibodies, they are small, heat- and pH-resistant, less expensive to produce and have the added advantage in that they do not interfere with endogenous insulin antibodies that diabetic patients who receive insulin often develop. Aptamers are isolated from nucleic acid libraries by a process known as SELEX (systemic evolution of ligands by exponential enrichment) by incubating the library with a target ligand molecule, and they can subsequently be produced synthetically. They have been explored for their potential use in biosensors for insulin in diabetes management with promising results [12][13][14][15][16][17][18][19][20][21][22][59,60,61,62,63,64,65,66,67,68,69].
Using SELEX, Yoshida et al. [16][63] isolated several DNA aptamers against insulin (Table 12) and they selected IGA3 for further testing. IGA3 is a G-rich aptamer which folds into a G-quadruplex and demonstrates the highest affinity for insulin. An enzyme system using IGA3 coupled to a thrombin-inhibiting aptamer was designed so that upon binding of insulin, thrombin is released, and its activity can be measured in a clotting assay. Although enzyme activity correlated with insulin levels, the system lacked sensitivity and was not tested using physiological samples. Several studies have since used IGA3 as an insulin recognition element for further biosensor development [19][20][21][22][23][24][66,67,68,69,70,71]. Wu et al. [21][68] developed a more sensitive electrochemical insulin biosensor where IGA3 was immobilised on a gold electrode surface and binding of insulin resulted in a conformational change detected by changes in electron transfer. Insulin in buffer was detected to 20 nmol/L, with a rapid response time of 60 s. Furthermore, the sensor was specific and selective and therefore has the potential for further development for clinical applications. The basic principle of an aptamer-based electrochemical insulin biosensor is shown in a simplified schematic in Figure 1. Electrochemical insulin biosensors have also been developed based on the DNA sequence of the insulin-linked polymorphic region (ILPR), which is located in the promoter of the human insulin gene and therefore can be considered as a natural insulin aptamer, and low detection limits down to 50 nmol/L were achieved [16][19][63,66].
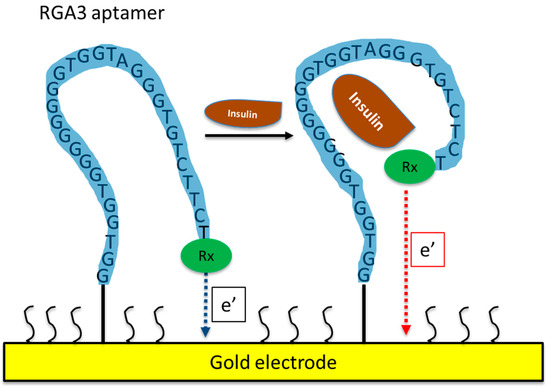
Figure 1. Schematic of an aptamer-based electrochemical insulin biosensor. Binding of insulin to the aptamer causes changes in its conformation and in electron transfer between a redox label and the gold electrode.
Table 12.
Sequences of aptamers sensitive to insulin used in biosensor technology.
| Aptamer Sequences | References | |||
|---|---|---|---|---|
| ILPR: 5′-CAGGGGTGTGGGGACAGGGGTGTGGGG-3′ | [16][19][63,66] | |||
| IGA1: 5′-GGAGGTGGATGGGGAGGGGGAGGTGTGTTT-3′ | [16][25][9,63] | |||
| IGA2: 5′-GGAGGGGGTGGGGAGGGGGCTGGTTGTCC-3′ | [16][ | Electrochemical63] | ||
| Buffer solution and human plasma | 26 fmol/L (buffer) and 81 fmol/L plasma) | 50–2000 pmol/L | [ | 29][75] |
| Electrochemical | Buffer solution | 1.9 pmol/L | 20–70 pmol/L | [30][76] |
| Electrochemical | Buffer solution | 33 fmol/L | 0.050–1.40 pmol/L | [31][77] |
4. Label-Free Insulin Biosensors
Another interesting category of insulin biosensors being developed is label-free and mainly employs electrochemical or optical transducers. Label-free detection is achieved through the intrinsic qualities of the target such as its charge, molecular mass and electrical impedance. Label-free systems offer simplicity at low costs and avoid a labelling stage that can result in sample degradation [32][78].
A label-free porous silicon-based optical biosensor was used to compare antibodies versus aptamers as bioreceptors for insulin, and interferometric reflectance spectroscopy (IRS) was used for detection [33][79]. Both antibodies and aptamers were highly selective for insulin although the aptamer-based approach demonstrated a faster response and lower limit of detection of 1.9 μg/mL. Servarayan et al. [34][80] presented a label-free fluorescence-based biosensor for the detection of insulin in human serum using novel naturally existing chromene mimic receptors. Its working range was from 10 fmol/L to 600 pmol/L, with a limit of detection of 7.07 fmol/L, which is reliable for the clinical detection of insulin. Chen and his colleagues [35][81] presented a label-free aptamer-based optical liquid-crystal (LC) biosensor for insulin. When an aptamer (IGA3) adsorbed to cetyltrimethylammonium bromide (CTAB) is bound to insulin, it undergoes a conformational change at the aqueous–liquid crystal interface, which results in a change in optical appearance from dark to bright that is then detected by polarised optical microscopy. This liquid crystal biosensor demonstrated a rapid response time of 5 min and high specificity and sensitivity for insulin in the range of 0.1–1.0 nmol/L in human urine and serum samples. Xu et al. [36][82] reported a label-free electrochemical sensor capable of ultrasensitive detection of insulin concentrations in blood serum. They immobilised insulin antibodies on gold electrodes and used electrochemical impedance spectroscopy (EIS) to monitor changes associated with the binding of insulin to the electrode surface. The sensor detected insulin across a clinically relevant range with a low detection limit of 4.7 pmol/L and was robust and could be regenerated without loss of sensitivity. Another EIS-based, rapid and label-free insulin biosensor with high sensitivity and accuracy was presented by Malcok et al. [37][83]. They used a similar principle to Xu et al. [10][57] of immobilising an insulin antibody to a gold electrode and used the imaginary impedance of EIS, to determine the optimal binding frequency (OBF) of insulin as 810.5 Hz and changes in imaginary impedance that correlated with insulin concentrations within a physiological range and with a low limit of insulin detection at 2.26 pmol/L.
Gobi et al. [38][84] developed a surface plasmon resonance (SPR) immunosensor based on a novel surface functionalisation method of covalent immobilisation of insulin on a self-assembled polyethylene glycol (PEG) monolayer. They used the principle of a competitive immunoassay using an antibody to insulin and demonstrated high sensitivity, specificity in detecting insulin with a response time of less than 5 min and a low-detection-limit of 1 ng/mL. The immunoreaction was followed in real time and the sensor could be regenerated for repeated use, making it suitable for further clinical development for point-of-care monitoring of insulin.
Hao et al. [39][85] reported an approach to a label-free, real-time insulin detection using a graphene aptameric nanosensor. It was based on a graphene field-effect transistor (GFET) and monitored the affinity binding between insulin and its specific aptameric receptor IGA3. The authors suggested that their biosensor could detect insulin with a limit of 35 pmol/L and hence could be developed for clinical use.
A comparison of the detection/signal principle, sample applications, detection limit and linear concentration ranges of different label-free-based biosensors for insulin detection is presented in Table 45.
Table 45.
Comparison of label-free-based biosensors for detection of insulin.
| Transducer | Sample | Detection Limit | Linear Range | References |
|---|---|---|---|---|
| IRS | Buffer solution and human islets | 4.299 nmol/L | 11.235–112.35 nmol/L | [33][79] |
| Fluorescence | Buffer solution and human serum | 7.07 fmol/L | 10 fmol/L–600 pmol/L | [34][80] |
| Fluorescence | Buffer solution, human urine and serum | 0.1 nmol/L | 0.1–1.0 nmol/L | [35] |
| IGA3: 5′-GGTGGTGGGGGGGGTGGTAGGGTGTCTTCT-3′ | [16][19][20][21][22][23][24][63,66,67,68,69,70,71] | |||
| clGA3: 5′-CCCCACACCCCTGTCCCCACACCCCTG-3′ | [22][69] | |||
| IBA2: 5′-CTCTCTCGGTGGTGGGGGGGGTTAGGGTGTCTTCCTCTCTC-3′ | [18][65] |
Sun et al. [20][67] developed a chemiluminescence biosensor for insulin also based on IGA3 and functionalised gold nanoparticles that are released upon the binding of insulin and catalyse the conversion of luminol. The biosensor was highly selective for insulin and had a low detection limit of 1.6 pmol/L. Wu et al. [22][69] found that insulin was bound to a C-rich-containing aptamer (cIGA3) with higher affinity and faster kinetics than the G-rich IGA3 aptamer, but specificity was poor for both aptamers and biological molecules such as haemoglobin and albumin were also bound. Their data contrast those by other investigators who found IGA3 to be highly specific for insulin with little interference by a range of molecules including albumin [20][21][67,68]. Aggregates of insulin can form and influence binding and, therefore, biological sample conditions need to be taken into consideration, especially when designing biosensors for clinical use [22][69]. Taghdisi et al. [18][65] developed a highly selective insulin biosensor utilising a triple helix switch consisting of another aptamer sequence that binds to insulin (IBA2, Table 12) and a fluorescent oligonucleotide probe. The probe is released upon insulin binding and results in the quenching of fluorescence. The sensor exhibited a low limit of detection of 9.7 nmol/L in serum and urine.
A comparison of detection techniques, sample applications, detection limits and linear concentration ranges of different aptamer-based biosensors for insulin can be found in Table 23.
Table 23.
Comparison of aptamer-based biosensors for detection of insulin.
| Transducer | Sample | Detection Limit | Linear Range | References | |||||
|---|---|---|---|---|---|---|---|---|---|
| Fluorescence | Rat serum and human urine | 9.97 nmol/L | 0–50 nmol/L | [18][65] | |||||
| Fluorescence | Human serum | 2 nmol/L | 2–70 nmol/L | [23][70] | |||||
| Fluorescence resonance energy transfer (FRET) | Human plasma | 0.6 pmol/L | 1 pmol/L–2.0 nmol/L | [24][71] | |||||
| [ | 81 | ] | Electrochemical | Buffer solution | 10 nmol/L | 10–200 nmol/L | [19][66] | ||
| Electrochemical | Buffer solution and human serum | 1.2 pmol/L | 5 pmol/L–50 nmol/L | [36][82] | Electrochemical | Buffer solution | 20 nmol/L | 0.02–5 μmol/L | [21][68] |
| Flow injection chemiluminescence | Buffer solution | 1.6 pmol/L | 7.5 pmol/L–5.0 nmol/L | [20][67] |
3. MIP-Based Insulin Biosensors
Molecular imprinting has received significant attention for nearly half a century. The fabrication of molecularly imprinted polymers (MIPs) involves the polymerisation of functional monomers around a template to form a cast with special recognition sites for the template. Upon removal of the template, the polymer can recognise the target molecule like antibodies recognise specific proteins. Molecular imprinting produces high-affinity polymers that can recognise not only proteins, but also nucleic acids, drugs and other targets and have several advantages over antibodies, as they are more stable, specific, easy and inexpensive to prepare and can be miniaturised for biosensor development.
Different molecular imprinting techniques can be employed in the construction of MIP-based sensors. These include bulk imprinting and surface imprinting [26][72]. In the first approach, the whole of the template molecule is imprinted in the polymer matrix and is then removed following polymerisation. While this approach increases selectivity for a specific molecule, in the case of insulin, smaller polypeptides and degradation products may also cross-react with the template. In addition, proteins may not remain in their correct conformation during the polymerisation process, thus also reducing selectivity. These drawbacks can be overcome by surface imprinting, where recognition sites on the surface of the template are used, are more accessible to the target and offer better binding kinetics.
Using quartz crystal microbalances (QCMs) as biosensor transducers, Schirhagl et al. [27][73] compared insulin detection by natural antibodies with detection using either a directly imprinted surface polyurethane polymer or a double-imprinted insulin antibody replicate. The synthetic coatings showed insulin selectively like the antibody, with a detection limit below 1 μg/mL. However, the antibody replicate showed significantly enhanced sensitivity and can be produced in bulk in a cost-effective manner, making this technology attractive for further development. Kartal et al. [28][74] also developed a QCM-based sensor by imprinting a complex of N-methacryloyl-(l)-histidine methyl ester with insulin onto gold QCM chips. Detection studies were carried out using insulin in aqueous solution and in artificial plasma samples and demonstrated a linear relationship between 0.008 and 1 ng/mL, high selectivity and stability, and a low detection limit (0.008 ng/mL).
In 2020, Piletsky and his colleagues [29][75] synthesised insulin MIP nanoparticles and immobilised them on screen-printed platinum electrodes to form a stable insulin biosensor for clinical applications. The sensor was selective for insulin with a linear response over a range of 50 to 2000 pmol/L and a limit of detection of 81 fmol/L in human plasma. The sensor showed good stability at room temperature and can be mass-produced at relatively low cost, thus fulfilling many requirements for production of a point-of-care insulin sensor. Zidarič et al. [30][76] developed an MIP receptor using an electrochemical technique to polymerise pyrrol in the presence of insulin on a carbon electrode using cyclic voltammetry. They used single-drop analysis to detect insulin in pharmaceutical samples. The biosensor successfully detected insulin in linear concentration ranges from 20 to 70 pmol/L (R2 = 0.9991) with a limit of detection at 1.9 pmol/L. Wardani et al. [31][77] also developed an electrochemical insulin sensor with an even lower reported limit of detection of 33 fmol/L and a linear range of 0.05 to 1.4 pmol/L. They used a gold electrode modified with carboxylated multiwalled carbon nanotubes (f-MWCNTs) and MIP cryogel. The sensor was highly selective for insulin and was stable when stored at room temperature. Abstracted information of the main parameters on the above MIP-based biosensors for insulin detection is provided in Table 34.
Table 34.
Comparison of MIP-based biosensors for detection of insulin.
| Transducer | Sample | Detection Limit | Linear Range | References |
|---|---|---|---|---|
| Dual-electrode QCM | Buffer solution | 2.247 nmol/L | 2.247–224.7 nmol/L | [27][73] |
| QCM chips | Aqueous solution and artificial plasma | 18 fmol/L | 18 fmol/L–2.247 pmol/L | [28][74] |
| Electrochemical | ||||
| Buffer solution | 2.26 pmol/L | 50–1500 pmol/L | [ | 37][83] |
| SPR | Buffer solution and human serum | 2.247 pmol/L | 2.247–674.1 pmol/L | [38][84] |
| Graphene electrical conductance | Buffer solution | 35 pmol/L | 100 pmol/L–1 μmol/L | [39][85] |
5. Other Types of Insulin Biosensors
Further to aptamer- and MIP-based insulin biosensors, there are a few other types reported in the literature, including a sensor based on cyclic voltammetry developed using cobalt hydroxide nanoparticles onto a carbon ceramic electrode [40][86]. The sensor was used for the detection of insulin in human serum samples and showed high stability, reproducibility and high selectivity, with a limit of detection and sensitivity of 0.11 nmol/L and 11.8 nA/nM, respectively. In 2018, Tan et al. [41][87] reported a colorimetric assay detecting glucose and insulin simultaneously using gold shell nanorods that possess peroxidase-like activity. In the presence of a peroxidase substrate and glucose oxidase, glucose levels were quantified calorimetrically, and a linear-concentration-dependent relationship was established. An insulin aptamer immobilised on the surface of the nanorods enabled simultaneous insulin detection by masking the catalytic activity of the peroxidase when insulin was bound. The method was developed to measure the glucose/insulin ratio and could potentially be used to differentiate between type I and type II diabetes. An electrochemiluminescent (ECL) biosensor based on carboxyl poly(9,9-dioctyfluorenyl-2,7-diyl) dots (PFO dots) and 3,4,9,10-perylenetetracar-boxylic acid (PTCA) to form an ECL-resonance energy transfer system by labelling two antibodies to detect insulin was published [42][88]. The biosensor presented good performance with a wide linear range, low detection limit, good stability and selectivity for insulin. Regonda et al. [43][89] demonstrated the advantages of utilising multiple Si nanochannels (NCs) or nanogratings (NGs) instead of the conventional single nanochannel or nanowire design in Si nanowire field effect transistors for biosensor application for insulin detection in human serum. The NG devices improved performance, reproducibility and device stability and insulin could be detected to a low limit of 10 fmol/L.
