Abstract: Polyphosphoinositides (PPIns) are signalling messengers representing less than five per cent of the total phospholipid concentration within the cell. Despite their low concentration, these lipids are critical regulators of various cellular processes, including cell cycle, differentiation, gene transcription, apoptosis and motility. PPIns are generated by the phosphorylation of the inositol head group of phosphatidylinositol (PtdIns). Different pools of PPIns are found at distinct subcellular compartments, which are regulated by an array of kinases, phosphatases and phospholipases. Six of the seven PPIns species have been found in the nucleus, including the nuclear envelope, the nucleoplasm and the nucleolus. The identification and characterisation of PPIns interactor and effector proteins in the nucleus have led to increasing interest in the role of PPIns in nuclear signalling. However, the regulation and functions of PPIns in the nucleus are complex and are still being elucidated. This review summarises our current understanding of the localisation, biogenesis and physiological functions of the different PPIns species in the nucleus.
- phosphoinositides
- PtdIns(4,5)P2
- signalling messengers
- lipid
- kinases
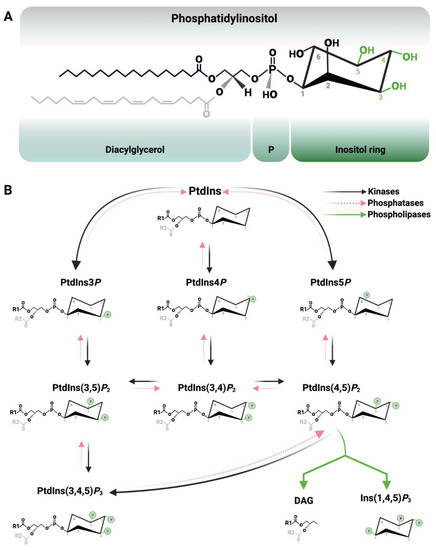
1. Introduction
| Protein Domain | Phosphoinositide(s) Bound | References |
|---|---|---|
| Pleckstrin homology (PH) domain | PtdIns3P, PtdIns4P, PtdIns(4,5)P2, PtdIns(3,4)P2 PtdIns(3,4,5)P3 |
[10,11,12,13,14,33,34,35,36] |
| Phox homology (PX) domain | PtdIns3P, PtdIns(3,4)P2, PtdIns(4,5)P2 | [37,38,39] |
| Plant homeodomain (PHD) | PtdIns3P, PtdIns5P | [40,41] |
| FYVE domain | PtdIns3P | [42] |
| ENTH domain | PtdIns(4,5)P2 | [43] |
| ANTH domain | PtdIns(4,5)P2 | [44] |
| Polybasic domains | PtdIns(4,5)P2 | [45,46,47] |
| Tubby | PtdIns(4,5)P2 | [48,49] |
2. The Nucleus and Nuclear PPIn Transport
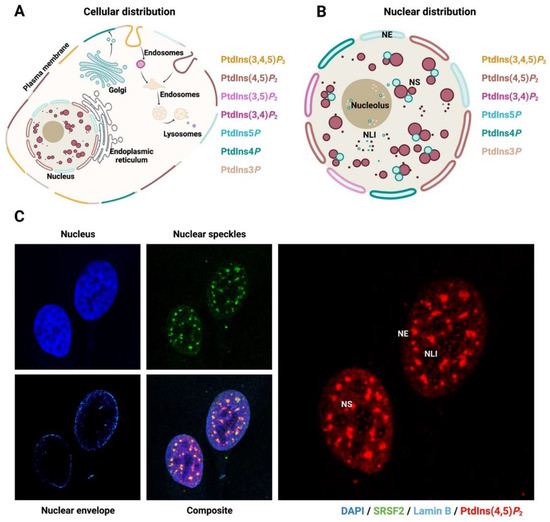
3. Biogenesis of Nuclear Phosphoinositides and Their Metabolising Enzymes
3.1. PtdIns3P
3.2. PtdIns4P
3.3. PtdIns5P
3.4. PtdIns(3,4)P2
3.5. PtdIns(4,5)P2
3.6. PtdIns(3,4,5)P3
4. Physiological Functions of Nuclear Phosphoinositide
4.1. Nuclear PPIns as Regulators of Histone Modifications
4.2. Nuclear PPIns and Their Role in Defining How Histone Modifications Drive Downstream Signalling Outputs
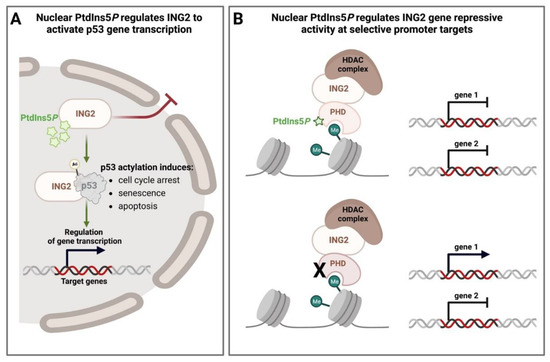
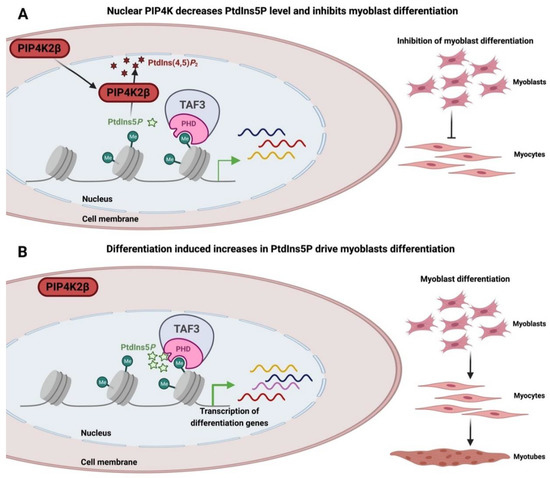
4.3. Nuclear PPIns and Regulation of Transcription Factors
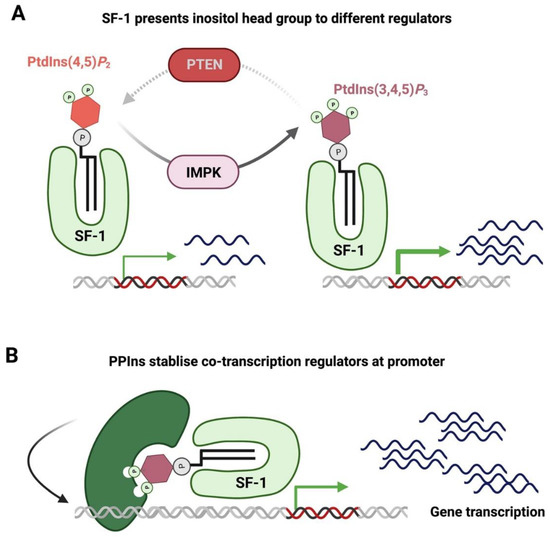
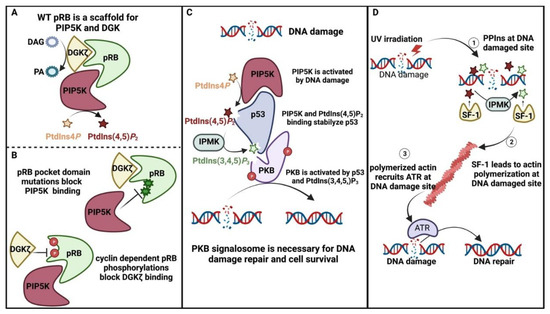
4.4. Nuclear PPIns and Their Role in Protecting the Genome
4.5. Nuclear Speckle and mRNA Machinery
5. Concluding Notes
In the early 1980s, key discoveries in nuclear PPIns centred around biochemical studies aimed at ensuring that the nuclear pool of PPIns was not a consequence of contamination from the much larger pool in the cytoplasm (plasma membrane, ER, Golgi) and, subsequently, on demonstrating that the nuclear pool could be regulated distinctly from other pools of PPIns. Further studies began to define nuclear targets that potentially interact with nuclear pools of PPIns, which effectively define downstream signalling pathways. In fact, we now have a plethora of signalling proteins that are nuclear, bind PPIns and potentially impact on all aspects of nuclear functions, though we lack considerable knowledge of how these pathways are controlled and coordinated. We still do not understand how the nuclear pool of PPIns is established, how it is maintained and how it is controlled and manipulated. The first two are rather perplexing, given that the immuno-localisation studies for PtdIns(4,5)P2 suggest that it is highly localised in membrane-less nuclear bodies, such as splicing speckles, islets and the nucleolus. This begs some simple questions. How do PPIns enter the nucleus and, once there, how are membrane-loving lipids maintained in membrane-less compartments or at specific genomic regions? PITP and or SF-1 may provide novel mechanisms to transport PPIns to specific genomic regions, such as promoters, where the exposed inositol head group can be presented to regulate epigenetic signalling. Alternatively, the regulation of subsets of genes by PPIns may occur through selective localisation of genes next to PPIns-rich regions of the nucleus. Proximity-based TSA assays have illustrated how genes that are upregulated often become more closely associated with nuclear splicing speckles where PtdIns(4,5)P2 is localized [208]. Finally, how enzymes that modulate nuclear PPIns are controlled is not well understood, which is critical, as this underpins how environmental cues (growth factors, DNA damage, etc.) impact on the levels of nuclear PPIns pools. In part, this lack of knowledge is driven by the lack of nuclear-specific isoforms that only regulate nuclear pools of PPIns. In most instances, the enzymes shuttle between the two compartments and control their cognate lipid in both compartments. For example, PIP4K2 has a nuclear localisation sequence that allows it to shuttle between compartments [106] and is phosphorylated in response to activation of the p38-MAPK stress pathway [173]. PIP5K1 also shuttles between the cytoplasm and nucleus. PIP5K1 is sumoylated at three different lysine residues, and while sumoylation at lysine 244 controls nuclear entry, sumoylation at lysine 490 controls subnuclear localisation to the nucleolus. Sumoylation at Lys-490 induces association of PIP5K1 with the chromatin silencing machinery, including heterochromatin protein 1 (HP1 ) and the epigenetic histone modification H3K9me3, to inhibit the expression of target ribosomal DNA (rDNA). Phospholipase C 1, which hydrolyses nuclear PtdIns(4,5)P2 to generate DAG and nuclear Ins(1,4,5)P3 shuttles between the cytoplasm and the nucleus. Of the two known isoforms of PLC 1 (1A and 1B), PLC 1B was suggested to be more localised to the nucleus compared to PLC 1A, due to the presence of a nuclear export sequence in an alternatively spliced exon in PLC 1A. PLC 1 is phosphorylated by the P42-MAPK, which leads to an increase in its nuclear activity [209] and subsequent phosphorylation by PKC acts to switch off PLC 1 [210]. Analysing the activity of these enzymes in vivo is even more challenging, given the lack of tools to study phosphoinositide dynamics in real-time in the nucleus. Clearly, we need a much more detailed map of how nuclear PPIns are modulated distinctly to the cytoplasmic pool if specific targeting of the nuclear PPIn pathway is to be used for therapeutic benefit.
References
1. Balla, T.; Szentpetery, Z.; Kim, Y.J.; Dingjan, I.; Linders, P.T.A.; Verboogen, D.R.J.; Revelo, N.H.; ter Beest, M.; Bogaart, G.v.D.; Sbrissa, D.; et al. Phosphoinositide Signaling: New Tools and Insights. Physiology 2009, 24, 231–244. [CrossRef]
2. Balla, T. Phosphoinositides: Tiny LipidsWith Giant Impact on Cell Regulation. Physiol. Rev. 2013, 93, 1019–1137. [CrossRef]
3. Morelec-Coulon, M.J.; Faure, M. Glycerol-inositol-phosphatidic acid. IV. Molecular structure. Bull. Soc. Chim. Biol. 1958, 40, 1307–1313.
4. Hawthornr the, J.N. The inositol phospholipids. J. Lipid Res. 1960, 1, 255–280. [CrossRef] [PubMed]
5. Hawthorne, J.N.; Kemp, P.; Ellis, R.B. Phosphoinositides. 2. The inositol 1-phosphate structure in liver phosphatidylinositol. Biochem. J. 1960, 75, 501. [CrossRef] [PubMed]
6. Irvine, R.F. Nuclear limpid signalling. Nat. Rev. Mol. Cell Biol. 2003, 4, 349–361. [CrossRef]
7. Lemmon, M.A. Phosphoinosietide Recognition Domains. Traffic 2003, 4, 201–213. [CrossRef]
8. Lemmon, M. Plecksartrin homology (PH) domains and phosphoinositides. Biochem. Soc. Symp. 2007, 74, 81–93. [CrossRef]
9. Hammond, G.R.; Baclla, T. Polyphosphoinositide binding domains: Key to inositol lipid biology. Biochim. Biophys. Acta (BBA)—Mol. Cell Biol. Lipids 2015, 1851, 746–758. [CrossRef]
10. Ferguson, K.M.; Lemmon, M.A.; Schlessinger, J.; Sigler, P.B. Structure of the high affinity complex of inositol trisphosphate with a phospholipase C pleckstrin homology domain. Cell 1995, 83, 1037–1046. [CrossRef]
11. Harlan, J.E.; Hajduk, P.J.; Yoon, H.S.; Fase refesik, S.W. Pleckstrin homology domains bind to phosphatidylinositol-4,5-bisphosphate. Nature 1994, 371, 168–170. [CrossRef] [PubMed]
12. Singh, N.; Rey to the Res-Ordoñez, A.; Compagnone, M.A.; Moreno, J.F.; Leslie, B.J.; Ha, T.; Chen, J. Redefining the specificity of phosphoinositide-binding by human PH domain-containing proteins. Nat. Commun. 2021, 12, 4339. [CrossRef] [PubMed]
13. Stauffer, T.P.; Ahn, S.; Meyer, T. Receptor-induced transient reduction in plasma membrane PtdIns(4,5)P2 concentration monitored in living cells. Curr. Biol. 1998, 8, 343–346. [CrossRef]
14. Guillou, H.; Lécureuil, C.; Anderson, K.A.; Suire, S.; Ferguson, G.J.; Ellson, C.D.; Gray, A.; Divecha, N.; Hawkins, P.T.; Stephens, L.R. Use of the GRP1 PH domain as a tool to measure the relative levels of PtdIns(3,4,5)P3 through a protein-lipid overlay approach. J. Lipid Res. 2007, 48, 726–732. [CrossRef] [PubMed]
15. Clark, J.; Anderson, K.E.; Juvin, V.; Smith, T.S.; Karpe, F.; Wakelam, M.J.O.; Stephens, L.R.; Hawkins, P.T. Quantification of PtdInsP3 molecular species in cells and tissues by mass spectrometry. Nat. Methods 2011, 8, 267–272. [CrossRef]
16. Milne, S.B.; Ivanova, P.T.; (DeCamp, D.; Hsueh, R.C.; Brown, H.A. A targeted mass spectrometric analysis of phosphatidylinositol phosphate species. J. Lipid Res. 2005, 46, 1796–1802. [CrossRef]
17. Vadnal, R.E.; Parthasarathy, R. The identification of a novel inositol lipid, phosphatidylinostiol trisphosphate (PIP3), in rat cerebrum using in vivo techniques. Biochem. Biophys. Res. Commun. 1989, 163, 995–1001. [CrossRef]
18. Lands,W.E. Metabolism of glycerolipids. 2. The enzymatic acylation of lysolecithin. J. Biol. Chem. 1960, 235, 2233–2237. [CrossRef]
19. Murphy, R.C.; Folco, G. Lysophospholipid acyltransferases and leukotriene biosynthesis: Intersection of the Lands cycle and the arachidonate PI cycle. J. Lipid Res. 2019, 60, 219–226. [CrossRef]
20. Anderson,K.E.; Kielkowska,A.;Durrant, T.N.; Juvin,V.; Clark, J.; Stephens, L.R.;Hawkins, P.T. Lysophosphatidylinositol-Acyltransferase-1 (LPIAT1) Is Required toMaintain Physiological Levels of PtdIns and PtdInsP2 in theMouse. PLoS ONE 2013, 8, e58425. [CrossRef]
21. Barneda, D.; Cosulich, S.; Stephens, L.; Hawkins, P. How is the acyl chain composition of phosphoinositides created and does it matter? Biochem. Soc. Trans. 2019, 47, 1291–1305. [CrossRef] [PubMed]
22. Patki, V.; Virbasius, J.; Lane, W.S.; Toh, B.-H.; Shpetner, H.S.; Corvera, S. Identification of an early endosomal protein regulated by phosphatidylinositol 3-kinase. Proc. Natl. Acad. Sci. USA 1997, 94, 7326–7330. [CrossRef] [PubMed]
23. Fiume, R.; Keune,W.J.; Faenza, I.; Bultsma, Y.; Ramazzotti, G.; Jones, D.R.; Martelli, A.M.; Somner, L.; Follo, M.Y.; Divecha, N. Nuclear phosphoinositides: Location, regulation and function. Phosphoinosit. II Divers. Biol. Funct. 2012, 59, 335–361.
24. Di Paolo, P.G.; De, C.P. Phosphoinositides in cell regulation and membrane dynamics. Nature 2006, 443, 651–657. [CrossRef][PubMed]
25. Giuriato, S.; Blero, D.; Robaye, B.; Bruyns, C.; Payrastre, B.; Erneux, C. SHIP2 overexpression strongly reduces the proliferation rate of K562 erythroleukemia cell line. Biochem. Biophys. Res. Commun. 2002, 296, 106–110. [CrossRef]
26. Clément, S.; Krause, U.; Desmedt, F.; Tanti, J.-F.; Behrends, J.; Pesesse, X.; Sasaki, T.; Penninger, J.; Doherty, M.; Malaisse, W.; et al. The lipid phosphatase SHIP2 controls insulin sensitivity. Nature 2001, 409, 92–97. [CrossRef]
27. Luo, J.-M.; Yoshida, H.; Komura, S.; Ohishi, N.; Pan, L.; Shigeno, K.; Hanamura, I.; Miura, K.; Iida, S.; Ueda, R.; et al. Possible dominant-negative mutation of the SHIP gene in acute myeloid leukemia. Leukemia 2003, 17, 1–8. [CrossRef] [PubMed]
28. Marion, E.; Kaisaki, P.J.; Pouillon, V.; Gueydan, C.; Levy, J.C.; Bodson, A.; Krzentowski, G.; Daubresse, J.-C.; Mockel, J.; Behrends, J.; et al. The Gene INPPL1, Encoding the Lipid Phosphatase SHIP2, Is a Candidate for Type 2 Diabetes In Rat and Man. Diabetes 2002, 51, 2012–2017. [CrossRef]
29. Halstead, J.R.; Jalink, K.; Divecha, N. An emerging role for PtdIns(4,5https://doi.org/10.3390/biom13071049)P2-mediated signalling in human disease. Trends Pharmacol. Sci. 2005, 26, 654–660. [CrossRef]
30. McCrea, H.J.; De Camilli, P.; Inoue, K.; Ishibe, S.; Silswal, N.; Parelkar, N.K.; Wacker, M.J.; Brotto, M.; Andresen, J. Mutations in Phosphoinositide Metabolizing Enzymes and Human Disease. Physiology 2009, 24, 8–16. [CrossRef]
31. Carrat, G.R.; Haythorne, E.; Tomas, A.; Haataja, L.; Müller, A.; Arvan, P.; Piunti, A.; Cheng, K.; Huang, M.; Pullen, T.J.; et al. The type 2 diabetes gene product STARD10 is a phosphoinositide-binding protein that controls insulin secretory granule biogenesis. Mol. Metab. 2020, 40, 101015. [CrossRef] [PubMed]
32. Bridges, D.; Saltiel, A.R. Phosphoinositides in Insulin Action and Diabetes. In Current Topics in Microbiology and Immunology; Springer: Berlin/Heidelberg, Germany, 2012; p. 362.
33. Kavran, J.M.; Klein, D.E.; Lee, A.; Falasca, M.; Isakoff, S.J.; Skolnik, E.Y.; Lemmon, M.A. Specificity and Promiscuity in Phosphoinositide Binding by Pleckstrin Homology Domains. J. Biol. Chem. 1998, 273, 30497–30508. [CrossRef] [PubMed]
34. Paterson, H.F.; Savopoulos, J.W.; Perisic, O.; Cheung, R.; Ellis, M.V.; Williams, R.L.; Katan, M. Phospholipase C d1 requires a pleckstrin homology domain for interaction with the plasma membrane. Biochem. J. 1995, 312, 661–666. [CrossRef] [PubMed]
35. Yagisawa, H.; Sakuma, K.; Paterson, H.F.; Cheung, R.; Allen, V.; Hirata, H.;Watanabe, Y.; Hirata, M.;Williams, R.L.; Katan, M. Replacements of single basic amino acids in the pleckstrin homology domain of phospholipase C-delta1 alter the ligand binding, phospholipase activity, and interaction with the plasma membrane. J. Biol. Chem. 1998, 273, 417–424. [CrossRef]
36. Kwon, I.-S.; Lee, K.-H.; Choi, J.W.; Ahn, J.-Y. PI(3,4,5)P3 regulates the interaction between Akt and B23 in the nucleus. BMB Rep. 2010, 43, 127–132. [CrossRef]
37. Kanai, F.; Liu, H.; Field, S.; Akbary, H.; Matsuo, T.; Brown, G.E.; Cantley, L.; Yaffe, M.B. The PX domains of p47phox and p40phox bind to lipid products of PI(3)K. Nature 2001, 3, 675–678. [CrossRef] [PubMed]
38. Ellson, C.D.; Andrews, S.; Stephens, L.R.; Hawkins, P.T. The PX domain: A new phosphoinositide-binding module. J. Cell Sci. 2002, 115, 1099–1105. [CrossRef]
39. Chandra, M.; Chin, Y.K.-Y.; Mas, C.; Feathers, J.R.; Paul, B.; Datta, S.; Chen, K.-E.; Jia, X.; Yang, Z.; Norwood, S.J.; et al. Classification of the human phox homology (PX) domains based on their phosphoinositide binding specificities. Nat. Commun. 2019, 10, 1528. [CrossRef]
40. Gozani, O.; Karuman, P.; Jones, D.R.; Ivanov, D.; Cha, J.; Lugovskoy, A.A.; Baird, C.L.; Zhu, H.; Field, S.J.; Lessnick, S.L.; et al. The PHD Finger of the Chromatin-Associated Protein ING2 Functions as a Nuclear Phosphoinositide Receptor. Cell 2003, 114, 99–111.[CrossRef]
41. Kaadige, M.R.; Ayer, D.E. The Polybasic Region That Follows the Plant Homeodomain Zinc Finger 1 of Pf1 Is Necessary and Sufficient for Specific Phosphoinositide Binding. J. Biol. Chem. 2006, 281, 28831–28836. [CrossRef]
42. Stenmark, H.; Aasland, R.; Driscoll, P. The phosphatidylinositol 3-phosphate-binding FYVE finger. FEBS Lett. 2002, 513, 77–84. [CrossRef]
43. Itoh, T.; Koshiba, S.; Kigawa, T.; Kikuchi, A.; Yokoyama, S.; Takenawa, T. Role of the ENTH domain in phosphatidylinositol-4,5-bisphosphate binding and endocytosis. Science 2001, 291, 1047–1051. [CrossRef] [PubMed]
44. Ford, M.G.; Pearse, B.M.; Higgins, M.K.; Vallis, Y.; Owen, D.J.; Gibson, A.; Hopkins, C.R.; Evans, P.R.; McMahon, H.T. Simultaneous binding of PtdIns(4,5)P2 and clathrin by AP180 in the nucleation of clathrin lattices on membranes. Science 2001, 291, 1051–1055. [CrossRef]
45. Lewis, A.E.; Sommer, L.; Arntzen, M.; Strahm, Y.; Morrice, N.A.; Divecha, N.; D’Santos, C.S. Identification of Nuclear Phosphatidylinositol
4,5-Bisphosphate-Interacting Proteins by Neomycin Extraction. Mol. Cell. Proteom. 2011, 10, S1–S15. [CrossRef]
46. Brown, D.A. PIP2Clustering: From model membranes to cells. Chem. Phys. Lipids 2015, 192, 33–40. [CrossRef] [PubMed]
47. Sztacho, M.; Šalovská, B.; Cˇ ervenka, J.; Balaban, C.; Hoboth, P.; Hozák, P. Limited Proteolysis-Coupled Mass Spectrometry Identifies Phosphatidylinositol 4,5-Bisphosphate Effectors in Human Nuclear Proteome. Cells 2021, 10, 68. [CrossRef] [PubMed]
48. Santagata, S.; Boggon, T.J.; Baird, C.L.; Gomez, C.A.; Zhao, J.; Shan, W.S.; Myszka, D.G.; Shapiro, L. G-Protein Signaling Through Tubby Proteins. Science 2001, 292, 2041–2050. [CrossRef]
49. Quinn, K.V.; Behe, P.; Tinker, A. Monitoring changes in membrane phosphatidylinositol 4,5-bisphosphate in living cells using a domain from the transcription factor tubby. J. Physiol. 2008, 586, 2855–2871. [CrossRef]
50. Doucet, C.M.; Hetzer, M.W. Nuclear pore biogenesis into an intact nuclear envelope. Chromosoma 2010, 119, 469–477. [CrossRef]
51. la Cour, T.; Kiemer, L.; Mølgaard, A.; Gupta, R.; Skriver, K.; Brunak, S. Analysis and prediction of leucine-rich nuclear export signals. Protein Eng. Des. Sel. 2004, 17, 527–536. [CrossRef]
52. Cokol, M.; Nair, R.; Rost, B. Finding nuclear localization signals. EMBO Rep. 2000, 1, 411–415. [CrossRef]
53. Holmer, L.; Worman, H. Inner nuclear membrane proteins: Functions and targeting. Cell. Mol. Life Sci. 2001, 58, 1741–1747. [CrossRef] [PubMed]
54. Misteli, T. Beyond the Sequence: Cellular Organization of Genome Function. Cell 2007, 128, 787–800. [CrossRef]
55. Carrero, G.; Hendzel, M.; de Vries, G. Modelling the compartmentalization of splicing factors. J. Theor. Biol. 2006, 239, 298–312. [CrossRef]
56. Cremer, T.; Cremer, C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat. Rev. Genet. 2001, 2, 292–301. [CrossRef] [PubMed]
57. Ho, L.; Crabtree, G.R. Chromatin remodelling during development. Nature 2010, 463, 474–484. [CrossRef]
58. Clapier, C.R.; Iwasa, J.; Cairns, B.R.; Peterson, C.L. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat. Rev. Mol. Cell Biol. 2017, 18, 407–422. [CrossRef] [PubMed]
59. Spector, D.; Lamond, A. Nuclear Speckles. Cold Spring Harb. Perspect. Biol. 2011, 3, a000646. [CrossRef] [PubMed]
60. Cremer, T.; Kreth, G.; Koester, H.; Fink, R.H.A.; Heintzmann, R.; Cremer, M.; Solovei, I.; Zink, D.; Cremer, C. Chromosome Territories, Interchromatin Domain Compartment, and Nuclear Matrix: An Integrated View of the Functional Nuclear Architecture. Crit. Rev. Eukaryot. Gene Expr. 2000, 10, 38. [CrossRef]
61. Osborne, S.; Thomas, C.; Gschmeissner, S.; Schiavo, G. Nuclear PtdIns(4,5)P2 assembles in a mitotically regulated particle involved in pre-mRNA splicing. J. Cell Sci. 2001, 114, 2501–2511. [CrossRef]
62. Kalasova, I.; Fáberová, V.; Kalendová, A.; Yildirim, S.; Uliˇcná, L.; Venit, T.; Hozák, P. Tools for visualization of phosphoinositide in the cell nucleus. Histochem. Cell Biol. 2016, 145, 485–496. [CrossRef]
63. Divecha, N.; Banfi´c, H.; Irvine, R.F. Inositides and the nucleus and inositides in the nucleus. Cell 1993, 74, 405–407. [CrossRef]
64. Divecha, N.; Banfic, H.; Irvine, R.F. The polyphosphoinositide cycle exists in the nuclei of Swiss 3T3 cells under the control of a receptor (for IGF-1) in the plasma membrane, and stimulation of the cycle increases nuclear diacylglycerol and apparently induces translocation of protein kinase. EMBO J. 1991, 10, 3207–3214. [CrossRef] [PubMed]
65. Banfi´c, H.; Žižak, M.; Divecha, N.; Irvine, R.F. Nuclear diacylglycerol is increased during cell proliferation in vivo. Biochem. J. 1993, 290, 633–636. [CrossRef] [PubMed]
66. Vann, L.R.; Wooding, P.F.; Irvine, R.F.; Divecha, N. Metabolism and possible compartmentalization of inositol lipids in isolated rat-liver nuclei. Biochem. J. 1997, 327, 569–576. [CrossRef]
67. Cocco, L.; Gilmour, R.S.; Ognibene, A.; Letcher, A.J.; Manzoli, F.A.; Irvine, R.F. Synthesis of polyphosphoinositides in nuclei of Friend cells. Evidence for polyphosphoinositide metabolism inside the nucleus which changes with cell differentiation. Biochem. J. 1987, 248, 765–770. [CrossRef]
68. Boronenkov, I.V.; Loijens, J.C.; Umeda, M.; Anderson, R.A. Phosphoinositide signaling pathways in nuclei are associated with nuclear speckles containing pre-mRNA processing factors. Mol. Biol. Cell 1998, 9, 3547–3560. [CrossRef] [PubMed]
69. De Vries, K.J.; Heinrichs, A.A.J.; Cunningham, E.; Brunink, F.; Westerman, J.; Somerharju, P.J.; Cockcroft, S.; Wirtz, K.W.A.; Snoek, G.T. An isoform of the phosphatidylinositol-transfer protein transfers sphingomyelin and is associated with the Golgi system. Biochem. J. 1995, 310, 643–649. [CrossRef] [PubMed]
70. Rubbini, S.; Cocco, L.; Manzoli, L.; Lutterman, J.; Billi, A.; Matteucci, A.; Wirtz, K. Phosphoinositide Signalling in Nuclei of Friend Cells: DMSO-Induced Differentiation Reduces the Association of Phosphatidylinositol-Transfer Protein with the Nucleus. Biochem. Biophys. Res. Commun. 1997, 230, 302–305. [CrossRef]
71. de Vries, K.; Westerman, J.; Bastiaens, P.; Jovin, T.; Wirtz, K.; Snoek, G. Fluorescently Labeled Phosphatidylinositol Transfer Protein Isoforms ( and ), Microinjected into Fetal Bovine Heart Endothelial Cells, Are Targeted to Distinct Intracellular Sites. Exp. Cell Res. 1996, 227, 33–39. [CrossRef]
72. Tribble, E.K.; Ivanova, P.T.; Grabon, A.; Alb, J.G.; Faenza, I.; Cocco, L.; Brown, H.A.; Bankaitis, V.A. Quantitative profiling of the endonuclear glycerophospholipidome of murine embryonic fibroblasts. J. Lipid Res. 2016, 57, 1492–1506. [CrossRef]
73. Carrillo, N.D.; Chen,M.; Cryns, V.L.; Anderson, R.A. Lipid transfer proteins initiate nuclear phosphoinositide signaling. bioRxiv 2023. [CrossRef]
74. Prasanth, K.V.; Sacco-Bubulya, P.A.; Prasanth, S.G.; Spector, D.L. Sequential Entry of Components of Gene Expression Machinery into Daughter Nuclei. Mol. Biol. Cell 2003, 14, 1043–1057. [CrossRef] [PubMed]
75. Sobol, M.; Yildirim, S.; Philimonenko, V.V.; Marášek, P.; Castaño, E.; Hozák, P. UBF complexes with phosphatidylinositol 4,5-bisphosphate in nucleolar organizer regions regardless of ongoing RNA polymerase I activity. Nucleus 2013, 4, 478–486. [CrossRef] [PubMed]
76. Smith, C.D.;Wells, W.W. Phosphorylation of rat liver nuclear envelopes. I. Characterization of in vitro protein phosphorylation. J. Biol. Chem. 1983, 258, 9360–9367. [CrossRef]
77. Shah, Z.H.; Jones, D.R.; Sommer, L.; Foulger, R.; Bultsma, Y.; D’Santos, C.; Divecha, N. Nuclear phosphoinositides and their impact on nuclear functions. FEBS J. 2013, 280, 6295–6310. [CrossRef]
78. Gaullier, J.-M.; Simonsen, A.; D’Arrigo, A.; Bremnes, B.; Stenmark, H.; Aasland, R. FYVE fingers bind PtdIns(3)P. Nature 1998, 394, 432–433. [CrossRef]
79. Stenmark, H.; Aasland, R. FYVE-finger proteins--effectors of an inositol lipid. J. Cell Sci. 1999, 112, 4175–4183. [CrossRef]
80. Gillooly, D.J.; Morrow, I.C.; Lindsay, M.; Gould, R.; Bryant, N.J.; Gaullier, J.; Parton, R.G.; Stenmark, H. Localization of phosphatidylinositol 3-phosphate in yeast and mammalian cells. EMBO J. 2000, 19, 4577–4588. [CrossRef]
81. Norris, F.A.; Majerus, P.W. Hydrolysis of phosphatidylinositol 3,4-bisphosphate by inositol polyphosphate 4-phosphatase isolated by affinity elution chromatography. J. Biol. Chem. 1994, 269, 8716–8720. [CrossRef]
82. Norris, F.A.; Auethavekiat, V.; Majerus, P.W. The Isolation and Characterization of cDNA Encoding Human and Rat Brain Inositol Polyphosphate 4-Phosphatase. J. Biol. Chem. 1995, 270, 16128–16133. [CrossRef] [PubMed]
83. Devereaux, K.; Dall’armi, C.; Alcazar-Roman, A.; Ogasawara, Y.; Zhou, X.; Wang, F.; Yamamoto, A.; De Camilli, P.; Di Paolo, G. Regulation of Mammalian Autophagy by Class II and III PI 3-Kinases through PI3P Synthesis. PLoS ONE 2013, 8, e76405. [CrossRef] [PubMed]
84. Franco, I.; Gulluni, F.; Campa, C.C.; Costa, C.; Margaria, J.P.; Ciraolo, E.; Martini, M.; Monteyne, D.; De Luca, E.; Germena, G.; et al. PI3K Class II Controls Spatially Restricted Endosomal PtdIns3P and Rab11 Activation to Promote Primary Cilium Function. Dev. Cell 2014, 28, 647–658. [CrossRef] [PubMed]
85. Gulluni, F.; De Santis, M.C.; Margaria, J.P.; Martini, M.; Hirsch, E. Class II PI3K Functions in Cell Biology and Disease. Trends Cell Biol. 2019, 29, 339–359. [CrossRef]
86. Didichenko, S.A.; Thelen, M. Phosphatidylinositol 3-Kinase C2 Contains a Nuclear Localization Sequence and Associates with Nuclear Speckles. J. Biol. Chem. 2001, 276, 48135–48142. [CrossRef] [PubMed]
87. Višnjic´, D.; Crljen, V.; C´ uric´, J.; Batinic´, D.; Volinia, S.; Banfic´, H. The activation of nuclear phosphoinositide 3-kinase C2 in all-trans-retinoic acid-differentiated HL-60 cells. FEBS Lett. 2002, 529, 268–274. [CrossRef]
88. Banfic, H.; Visnjic, D.; Mise, N.; Balakrishnan, S.; Deplano, S.; Korchev, Y.E.; Domin, J. Epidermal growth factor stimulates translocation of the class II phosphoinositide 3-kinase PI3K-C2 to the nucleus. Biochem. J. 2009, 422, 53–60. [CrossRef] [PubMed]
89. Huang, P.T.; Einav, S.; Asquith, C.R.M. PIKfyve: A lipid kinase target for COVID-19, cancer and neurodegenerative disorders. Nat. Rev. Drug Discov. 2021, 20, 730. [CrossRef]
90. Clague, M.J.; Lorenzo, Ó. The Myotubularin Family of Lipid Phosphatases. Traffic 2005, 6, 1063–1069. [CrossRef]
91. Robinson, F.L.; Dixon, J.E. Myotubularin phosphatases: Policing 3-phosphoinositides. Trends Cell Biol. 2006, 16, 403–412. [CrossRef]
92. Divecha, N.; Letcher, A.J.; Banfic, H.H.; Rhee, S.G.; Irvine, R.F. Changes in the components of a nuclear inositide cycle during differentiation in murine erythroleukaemia cells. Biochem. J. 1995, 312, 63–67. [CrossRef] [PubMed]
93. Fáberová, V.; Kalasová, I.; Krausová, A.; Hozák, P. Super-Resolution Localisation of Nuclear PI(4)P and Identification of Its Interacting Proteome. Cells 2020, 9, 1191. [CrossRef] [PubMed]
94. Mao, Y.; Manford, A.; Xia, T.; Saxena, A.; Stefan, C.; Hu, F.; Emr, S. Crystal structure of the Yeast Sac1: Implications for its phosphoinositide phosphatase function. EMBO J. 2010, 29, 1489–1498. [CrossRef]
95. Zewe, J.P.;Wills, R.C.; Sangappa, S.; Goulden, B.D.; Hammond, G.R. SAC1 degrades its lipid substrate PtdIns4P in the endoplasmic reticulum to maintain a steep chemical gradient with donor membranes. eLife 2018, 7, e35588. [CrossRef] [PubMed]
96. Clarke, J.H.; Letcher, A.J.; D’Santos, C.S.; Halstead, J.R.; Irvine, R.F.; Divecha, N. Inositol lipids are regulated during cell cycle progression in the nuclei of murine erythroleukaemia cells. Biochem. J. 2001, 357, 905–910. [CrossRef]
97. Clarke, J.H.; Irvine, R.F. Evolutionarily conserved structural changes in phosphatidylinositol 5-phosphate 4-kinase (PI5P4K) isoforms are responsible for differences in enzyme activity and localization. Biochem. J. 2013, 454, 49–57. [CrossRef]
98. Divecha, N.; Brooksbank, C.E.L.; Irvine, R.F. Purification and characterization of phosphatidylinositol 4-phosphate 5-kinases. Biochem. J. 1992, 288, 637–642. [CrossRef]
99. Brooksbank, C.E.L.; Hutchings, A.; Butcher, G.W.; Irvine, R.F.; Divecha, N. Monoclonal antibodies to phosphatidylinositol 4-phosphate 5-kinase: Distribution and intracellular localization of the C isoform. Biochem. J. 1993, 291, 77–82. [CrossRef]
100. Bazenet, C.E.; Ruano, A.R.; Brockman, J.L.; Anderson, R.A. The human erythrocyte contains two forms of phosphatidylinositol-4-phosphate 5-kinase which are differentially active toward membranes. J. Biol. Chem. 1990, 265, 18012–18022. [CrossRef]
101. Rameh, L.E.; Tolias, K.F.; Duckworth, B.C.; Cantley, L.C. A new pathway for synthesis of phosphatidylinositol-4,5-bisphosphate. Nature 1997, 390, 192–196. [CrossRef]
102. Divecha, N.; Truong, O.; Hsuan, J.J.; Hinchliffe, K.A.; Irvine, R.F. The cloning and sequence of the C isoform of PtdIns4P 5-kinase. Biochem. J. 1995, 309, 715–719. [CrossRef]
103. Castellino, A.M.; Parker, G.J.; Boronenkov, I.V.; Anderson, R.A.; Chao, M.V. A Novel Interaction between the Juxtamembrane Region of the p55 Tumor Necrosis Factor Receptor and Phosphatidylinositol-4-phosphate 5-Kinase. J. Biol. Chem. 1997, 272, 5861–5870. [CrossRef] [PubMed]
104. Boronenkov, I.V.; Anderson, R.A. The Sequence of Phosphatidylinositol-4-phosphate 5-Kinase Defines a Novel Family of Lipid Kinases. J. Biol. Chem. 1995, 270, 2881–2884. [CrossRef] [PubMed]
105. Jones, D.R.; Divecha, N. Linking lipids to chromatin. Curr. Opin. Genet. Dev. 2004, 14, 196–202. [CrossRef]
106. Ciruela, A.; Hinchliffe, K.A.; Divecha, N.; Irvine, R.F. Nuclear targeting of the beta isoform of type II phosphatidylinositol phosphate kinase (phosphatidylinositol 5-phosphate 4-kinase) by its alpha-helix 7. Biochem. J. 2000, 346, 587–591. [CrossRef]
107. Hasegawa, J.; Strunk, B.S.; Weisman, L.S. PI5P and PI(3,5)P2: Minor, but Essential Phosphoinositides. Cell Struct. Funct. 2017, 42, 49–60. [CrossRef]
108. Poli, A.; Zaurito, A.E.; Abdul-Hamid, S.; Fiume, R.; Faenza, I.; Divecha, N. Phosphatidylinositol 5 Phosphate (PI5P): From Behind the Scenes to the Front (Nuclear) Stage. Int. J. Mol. Sci. 2019, 20, 2080. [CrossRef] [PubMed]
109. Bulley, S.J.; Droubi, A.; Clarke, J.H.; Anderson, K.E.; Stephens, L.R.; Hawkins, P.T.; Irvine, R.F. In B cells, phosphatidylinositol 5-phosphate 4-kinase– synthesizes PI(4,5)P2 to impact mTORC2 and Akt signaling. Proc. Natl. Acad. Sci. USA 2016, 113, 10571–10576. [CrossRef]
110. Lundquist, M.R.; Goncalves, M.D.; Loughran, R.M.; Possik, E.; Vijayaraghavan, T.; Yang, A.; Pauli, C.; Ravi, A.; Verma, A.; Yang, Z.; et al. Phosphatidylinositol-5-Phosphate 4-Kinases Regulate Cellular Lipid Metabolism By Facilitating Autophagy. Mol. Cell 2018, 70, 531–544.e9. [CrossRef]
111. Vicinanza, M.; Korolchuk, V.; Ashkenazi, A.; Puri, C.; Menzies, F.M.; Clarke, J.; Rubinsztein, D.C. PI(5)P Regulates Autophagosome Biogenesis. Mol. Cell 2015, 57, 219–234. [CrossRef]
112. Poli, A.; Pennacchio, F.A.; Ghisleni, A.; di Gennaro, M.; Lecacheur, M.; Nastały, P.; Crestani, M.; Pramotton, F.M.; Iannelli, F.; Beznusenko, G.; et al. PIP4K2B is mechanoresponsive and controls heterochromatin-driven nuclear softening through UHRF1. Nat. Commun. 2023, 14, 1432. [CrossRef] [PubMed]
113. Poli, A.; Abdul-Hamid, S.; Zaurito, A.E.; Campagnoli, F.; Bevilacqua, V.; Sheth, B.; Fiume, R.; Pagani, M.; Abrignani, S.; Divecha, N. PIP4Ks impact on PI3K, FOXP3, and UHRF1 signaling and modulate human regulatory T cell proliferation and immunosuppressive activity. Proc. Natl. Acad. Sci. USA 2021, 118, e2010053118. [CrossRef] [PubMed]
114. Blind, R.D.; Sablin, E.P.; Kuchenbecker, K.M.; Chiu, H.-J.; Deacon, A.M.; Das, D.; Fletterick, R.J.; Ingraham, H.A. The signaling phospholipid PIP 3 creates a new interaction surface on the nuclear receptor SF-1. Proc. Natl. Acad. Sci. USA 2014, 111, 15054–15059. [CrossRef]
115. Ndamukong, I.; Jones, D.R.; Lapko, H.; Divecha, N.; Avramova, Z. Phosphatidylinositol 5-Phosphate Links Dehydration Stress to the Activity of ARABIDOPSIS TRITHORAX-LIKE Factor ATX1. PLoS ONE 2010, 5, e13396. [CrossRef]
116. Sbrissa, D.; Ikonomov, O.C.; Shisheva, A. PIKfyve, a Mammalian Ortholog of Yeast Fab1p Lipid Kinase, Synthesizes 5- Phosphoinositides. J. Biol. Chem. 1999, 274, 21589–21597. [CrossRef]
117. Schaletzky, J.; Dove, S.K.; Short, B.; Lorenzo, O.; Clague, M.J.; Barr, F.A. Phosphatidylinositol-5-Phosphate Activation and Conserved Substrate Specificity of the Myotubularin Phosphatidylinositol 3-Phosphatases. Curr. Biol. 2003, 13, 504–509. [CrossRef]
118. Zolov, S.N.; Bridges, D.; Zhang, Y.; Lee,W.W.; Riehle, E.; Verma, R.; Lenk, G.M.; Converso-Baran, K.; Weide, T.; Albin, R.L.; et al. In vivo, Pikfyve generates PI(3,5)P2, which serves as both a signaling lipid and the major precursor for PI5P. Proc. Natl. Acad. Sci. USA 2012, 109, 17472–17477. [CrossRef]
119. Ungewickell, A.; Hugge, C.; Kisseleva, M.; Chang, S.C.; Zou, J.; Feng, Y.; Galyov, E.E.; Wilson, M.; Majerus, P.W. The identification and characterization of two phosphatidylinositol-4,5-bisphosphate 4-phosphatases. Proc. Natl. Acad. Sci. USA 2005, 102, 18854–18859. [CrossRef]
120. Zou, J.; Marjanovic, J.; Kisseleva, M.V.;Wilson, M.; Majerus, P.W. Type I phosphatidylinositol-4,5-bisphosphate 4-phosphatase regulates stress-induced apoptosis. Proc. Natl. Acad. Sci. USA 2007, 104, 16834–16839. [CrossRef] [PubMed]
121. Posor, Y.; Eichhorn-Gruenig, M.; Puchkov, D.; Schöneberg, J.; Ullrich, A.; Lampe, A.; Müller, R.; Zarbakhsh, S.; Gulluni, F.; Hirsch, H.; et al. Spatiotemporal control of endocytosis by phosphatidylinositol-3,4-bisphosphate. Nature 2013, 499, 233–237. [CrossRef]
122. Alliouachene, S.; Bilanges, B.; Chicanne, G.; Anderson, K.E.; Pearce, W.; Ali, K.; Valet, C.; Posor, Y.; Low, P.C.; Chaussade, C.; et al. Inactivation of the Class II PI3K-C2 Potentiates Insulin Signaling and Sensitivity. Cell Rep. 2015, 13, 1881–1894. [CrossRef] [PubMed]
123. Yokogawa, T.; Nagata, S.; Nishio, Y.; Tsutsumi, T.; Ihara, S.; Shirai, R.; Morita, K.; Umeda, M.; Shirai, Y.; Saitoh, N.; et al. Evidence that 30-phosphorylated polyphosphoinositides are generated at the nuclear surface: Use of immunostaining technique with monoclonal antibodies specific for PI 3,4-P2. FEBS Lett. 2000, 473, 222–226. [CrossRef] [PubMed]
124. Marat, A.L.; Wallroth, A.; Lo, W.T.; Müller, R.; Norata, G.D.; Falasca, M.; Schultz, C.; Haucke, V. mTORC1 activity repression by late endosomal phosphatidylinositol 3,4-bisphosphate. Science 2017, 356, 968–972. [CrossRef] [PubMed]
125. Gozzelino, L.; De Santis, M.C.; Gulluni, F.; Hirsch, E.; Martini, M. PI(3,4)P2 Signaling in Cancer and Metabolism. Front. Oncol. 2020, 10, 360. [CrossRef]
126. Kerr, W.G. Inhibitor and activator: Dual functions for SHIP in immunity and cancer. Ann. N. Y. Acad. Sci. 2010, 1217, 1–17. [CrossRef]
127. Krystal, G.; Damen, J.E.; Helgason, C.D.; Huber, M.; Hughes, M.R.; Kalesnikoff, J.; Lam, V.; Rosten, P.;Ware, M.D.; Yew, S.; et al. SHIPs ahoy. Int. J. Biochem. Cell Biol. 1999, 31, 1007–1010. [CrossRef]
128. Ooms, L.M.; Binge, L.C.; Davies, E.M.; Rahman, P.; Conway, J.R.; Gurung, R.; Ferguson, D.T.; Papa, A.; Fedele, C.G.; Vieusseux, J.L.; et al. The Inositol Polyphosphate 5-Phosphatase PIPP Regulates AKT1-Dependent Breast Cancer Growth and Metastasis. Cancer Cell 2015, 28, 155–169. [CrossRef]
129. Norris, F.A.; Atkins, R.C.; Majerus, P.W. The cDNA Cloning and Characterization of Inositol Polyphosphate 4-Phosphatase Type II. J. Biol. Chem. 1997, 272, 23859–23864. [CrossRef]
130. Watt, S.A.; Kular, G.; Fleming, I.N.; Downes, C.P.; Lucoqc, J.M. Subcellular localization of phosphatidylinositol 4,5-bisphosphate using the pleckstrin homology domain of phospholipase C 1. Biochem. J. 2002, 363, 657–666. [CrossRef]
131. Miyazawa, A.; Umeda, M.; Horikoshi, T.; Yanagisawa, K.; Yoshioka, T.; Inoue, K. Production and characterization of monoclonal antibodies that bind to phosphatidylinositol 4,5-bisphosphate. Mol. Immunol. 1988, 25, 1025–1031.
132. Fukami, K.; Matsuoka, K.; Nakanishi, O.; Yamakawa, A.; Kawai, S.; Takenawa, T. Antibody to phosphatidylinositol 4,5-bisphosphate inhibits oncogene-induced mitogenesis. Proc. Natl. Acad. Sci. USA 1988, 85, 9057–9061. [CrossRef] [PubMed]
133. Thomas, C.L.; Steel, J.; Prestwich, G.D.; Schiavo, G. Generation of phosphatidylinositol-specific antibodies and their characterization. Biochem. Soc. Trans. 1999, 27, 648–652. [CrossRef]
134. Yildirim, S.; Castano, E.; Sobol, M.; Philimonenko, V.V.; Dzijak, R.; Venit, T.; Hozák, P. Involvement of PIP2 in RNA Polymerase I transcription. J. Cell Sci. 2013, 126, 2730–2739. [CrossRef]
135. Anderson, R.A.; Boronenkov, I.V.; Doughman, S.D.; Kunz, J.; Loijens, J.C. Phosphatidylinositol Phosphate Kinases, a Multifaceted Family of Signaling Enzymes. J. Biol. Chem. 1999, 274, 9907–9910. [CrossRef] [PubMed]
136. Mellman, D.L.; Gonzales, M.L.; Song, C.; Barlow, C.A.;Wang, P.; Kendziorski, C.; Anderson, R.A. A PtdIns4,5P2-regulated nuclear poly(A) polymerase controls expression of select mRNAs. Nature 2008, 451, 1013–1017. [CrossRef]
137. Ishihara, H.; Shibasaki, Y.; Kizuki, N.; Wada, T.; Yazaki, Y.; Asano, T.; Oka, Y. Type I phosphatidylinositol-4-phosphate 5-kinases. Cloning of the third isoform and deletion/substitution analysis of members of this novel lipid kinase family. J. Biol. Chem. 1998, 273, 8741–8748. [CrossRef] [PubMed]
138. Loijens, J.C.; Boronenkov, I.V.; Parker, G.J.; Anderson, R.A. The phosphatidylinositol 4-phosphate 5-kinase family. Adv. Enzym. Regul. 1996, 36, 115–140. [CrossRef]
139. van den Bout, I.; Divecha, N. PIP5K-driven PtdIns(4,5)P2 synthesis: Regulation and cellular functions. J. Cell Sci. 2009, 122, 3837–3850. [CrossRef]
140. Tolias, K.F.; Rameh, L.E.; Ishihara, H.; Shibasaki, Y.; Chen, J.; Prestwich, G.D.; Cantley, L.C.; Carpenter, C.L. Type I Phosphatidylinositol-4-phosphate 5-Kinases Synthesize the Novel Lipids Phosphatidylinositol 3,5-Bisphosphate and Phosphatidylinositol 5-Phosphate. J. Biol. Chem. 1998, 273, 18040–18046. [CrossRef]
141. Liu, Y.; Bankaitis, V.A. Phosphoinositide phosphatases in cell biology and disease. Prog. Lipid Res. 2010, 49, 201–217. [CrossRef]
142. Ooms, L.M.; Horan, K.A.; Rahman, P.; Seaton, G.; Gurung, R.; Kethesparan, D.S.; Mitchell, C.A. The role of the inositol polyphosphate 5-phosphatases in cellular function and human disease. Biochem. J. 2009, 419, 29–49. [CrossRef]
143. Dyson, J.M.; Fedele, C.G.; Davies, E.M.; Becanovic, J.; Mitchell, C.A. Phosphoinositide Phosphatases: Just as Important as the Kinases. Phosphoinosit. I Enzym. Synth. Degrad. 2012, 58, 215–279. [CrossRef]
144. Martelli, A.M.; Gilmour, R.S.; Bertagnolo, V.; Neri, L.M. Nuclear localization and signaling activity of phosphoinositidase C b in Swiss 3T3 cells. Nature 1992, 358, 242–245. [CrossRef] [PubMed]
145. Song, M.S.; Salmena, L.; Pandolfi, P.P. The functions and regulation of the PTEN tumour suppressor. Nat. Rev. Mol. Cell Biol. 2012, 13, 283–296. [CrossRef] [PubMed]
146. Hollander, M.C.; Blumenthal, G.M.; Dennis, P.A. PTEN loss in the continuum of common cancers, rare syndromes and mouse models. Nat. Rev. Cancer 2011, 11, 289–301. [CrossRef]
147. Lindsay, Y.; McCoull, D.; Davidson, L.; Leslie, N.R.; Fairservice, A.; Gray, A.; Lucocq, J.; Downes, C.P. Localization of agonistsensitive PtdIns(3,4,5)P3 reveals a nuclear pool that is insensitive to PTEN expression. J. Cell Sci. 2006, 119, 5160–5168. [CrossRef]
148. Edimo, W.E.; Vanderwinden, J.M.; Erneux, C. SHIP2 signalling at the plasma membrane, in the nucleus and at focal contacts. Adv. Biol. Regul. 2013, 53, 28–37. [CrossRef]
149. Foukas, L.C.; Berenjeno, I.M.; Gray, A.; Khwaja, A.; Vanhaesebroeck, B. Activity of any class IA PI3K isoform can sustain cell proliferation and survival. Proc. Natl. Acad. Sci. USA 2010, 107, 11381–11386. [CrossRef]
150. Davis, W.J.; Lehmann, P.Z.; Li, W. Nuclear PI3K signaling in cell growth and tumorigenesis. Front. Cell Dev. Biol. 2015, 3, 24. [CrossRef]
151. Resnick, A.C.; Snowman, A.M.; Kang, B.N.; Hurt, K.J.; Snyder, S.H.; Saiardi, A. Inositol polyphosphate multikinase is a nuclear PI3-kinase with transcriptional regulatory activity. Proc. Natl. Acad. Sci. USA 2005, 102, 12783–12788. [CrossRef]
152. Blind, R.D.; Suzawa, M.; Ingraham, H.A. Direct Modification and Activation of a Nuclear Receptor–PIP2 Complex by the Inositol Lipid Kinase IPMK. Sci. Signal. 2012, 5, ra44. [CrossRef] [PubMed]
153. Edimo, W.E.; Derua, R.; Janssens, V.; Nakamura, T.; Vanderwinden, J.M.; Waelkens, E.; Erneux, C. Evidence of SHIP2 Ser132 phosphorylation, its nuclear localization and stability. Biochem. J. 2011, 439, 391–404. [CrossRef]
154. Chung, J.-H.; Eng, C. Nuclear-Cytoplasmic Partitioning of Phosphatase and Tensin Homologue Deleted on Chromosome 10 (PTEN) Differentially Regulates the Cell Cycle and Apoptosis. Cancer Res. 2005, 65, 8096–8100. [CrossRef] [PubMed]
155. Liu, J.-L.; Sheng, X.; Hortobagyi, Z.K.; Mao, Z.; Gallick, G.E.; Yung, W.K.A. Nuclear PTEN-Mediated Growth Suppression Is Independent of Akt Down-Regulation. Mol. Cell. Biol. 2005, 25, 6211–6224. [CrossRef]
156. Allis, C.D.; Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet. 2016, 17, 487–500. [CrossRef] [PubMed]
157. Lee, D.Y.; Hayes, J.J.; Pruss, D.;Wolffe, A.P. A positive role for histone acetylation in transcription factor access to nucleosomal DNA. Cell 1993, 72, 73–84. [CrossRef]
158. Biel, M.;Wascholowski, V.; Giannis, A. Epigenetics—An Epicenter of Gene Regulation: Histones and Histone-Modifying Enzymes. Angew. Chem. Int. Ed. 2005, 44, 3186–3216. [CrossRef]
159. McKay, L.M.; Carpenter, B.; Roberts, S.G. Regulation of the Wilms’ tumour suppressor protein transcriptional activation domain. Oncogene 1999, 18, 6546–6554. [CrossRef]
160. Roberts, S.G. Transcriptional regulation by WT1 in development. Curr. Opin. Genet. Dev. 2005, 15, 542–547. [CrossRef]
161. Carpenter, B.; Hill, K.J.; Charalambous, M.; Wagner, K.J.; Lahiri, D.; James, D.I.; Andersen, J.S.; Schumacher, V.; Royer-Pokora, B.; Mann, M.; et al. BASP1 Is a Transcriptional Cosuppressor for the Wilms’ Tumor Suppressor Protein WT1. Mol. Cell. Biol. 2004, 24, 537–549. [CrossRef]
162. Toska, E.; Campbell, H.A.; Shandilya, J.; Goodfellow, S.J.; Shore, P.; Medler, K.F.; Roberts, S.G. Repression of Transcription by WT1-BASP1 Requires the Myristoylation of BASP1 and the PIP2-Dependent Recruitment of Histone Deacetylase. Cell Rep. 2012, 2, 462–469. [CrossRef] [PubMed]
163. Cheng, M.K.; Shearn, A. The Direct Interaction Between ASH2, a Drosophila Trithorax Group Protein, and SKTL, a Nuclear Phosphatidylinositol 4-Phosphate 5-Kinase, Implies a Role for Phosphatidylinositol 4,5-Bisphosphate in Maintaining Transcriptionally Active Chromatin. Genetics 2004, 167, 1213–1223. [CrossRef]
164. Visa, N.; Percipalle, P. Nuclear Functions of Actin. Cold Spring Harb. Perspect. Biol. 2010, 2, a000620. [CrossRef]
165. Rando, O.J.; Zhao, K.; Janmey, P.; Crabtree, G.R. Phosphatidylinositol-dependent actin filament binding by the SWI/SNF-like BAF chromatin remodeling complex. Proc. Natl. Acad. Sci. USA 2002, 99, 2824–2829. [CrossRef]
166. Batista, N.J.; Desai, S.G.; Perez, A.M.; Finkelstein, A.; Radigan, R.; Singh, M.; Landman, A.; Drittel, B.; Abramov, D.; Ahsan, M.; et al. The Molecular and Cellular Basis of Hutchinson–Gilford Progeria Syndrome and Potential Treatments. Genes 2023, 14, 602. [CrossRef] [PubMed]
167. Yu, H.; Fukami, K.; Watanabe, Y.; Ozaki, C.; Takenawa, T. Phosphatidylinositol 4,5-bisphosphate reverses the inhibition of RNA transcription caused by histone H1. JBIC J. Biol. Inorg. Chem. 1998, 251, 281–287. [CrossRef]
168. Gelato, K.A.; Tauber, M.; Ong, M.S.; Winter, S.; Hiragami-Hamada, K.; Sindlinger, J.; Lemak, A.; Bultsma, Y.; Houliston, S.; Schwarzer, D.; et al. Accessibility of Different Histone H3-Binding Domains of UHRF1 Is Allosterically Regulated by Phosphatidylinositol 5-Phosphate. Mol. Cell 2014, 54, 905–919. [CrossRef]
169. Stijf-Bultsma, Y.; Sommer, L.; Tauber, M.; Baalbaki, M.; Giardoglou, P.; Jones, D.R.; Gelato, K.A.; van Pelt, J.; Shah, Z.; Rahnamoun, H.; et al. The Basal Transcription Complex Component TAF3 Transduces Changes in Nuclear Phosphoinositides into Transcriptional Output. Mol. Cell 2015, 58, 453–467. [CrossRef] [PubMed]
170. Peña, P.V.; Davrazou, F.; Shi, X.;Walter, K.L.; Verkhusha, V.V.; Gozani, O.; Zhao, R.; Kutateladze, T.G. Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 2006, 442, 100–103. [CrossRef]
171. Li, H.; Ilin, S.;Wang,W.; Duncan, E.M.;Wysocka, J.; Allis, C.D.; Patel, D.J. Molecular basis for site-specific read-out of histone H3K4me3 by the BPTF PHD finger of NURF. Nature 2006, 442, 91–95. [CrossRef]
172. Feng, X.; Hara, Y.; Riabowol, K. Different HATS of the ING1 gene family. Trends Cell Biol. 2002, 12, 532–538. [CrossRef]
173. Jones, D.R.; Bultsma, Y.; Keune,W.-J.; Halstead, J.R.; Elouarrat, D.; Mohammed, S.; Heck, A.J.; D’Santos, C.S.; Divecha, N. Nuclear PtdIns5P as a Transducer of Stress Signaling: An In Vivo Role for PIP4Kbeta. Mol. Cell 2006, 23, 685–695. [CrossRef]
174. Bua, D.J.; Martin, G.M.; Binda, O.; Gozani, O. Nuclear phosphatidylinositol-5-phosphate regulates ING2 stability at discrete chromatin targets in response to DNA damage. Sci. Rep. 2013, 3, 2137. [CrossRef]
175. Reed, S.M.; Quelle, D.E. p53 Acetylation: Regulation and Consequences. Cancers 2014, 7, 30–69. [CrossRef]
176. Nagashima, M.; Shiseki, M.; Miura, K.; Hagiwara, K.; Linke, S.P.; Pedeux, R.;Wang, X.W.; Yokota, J.; Riabowol, K.; Harris, C.C. DNA damage-inducible gene p33ING2 negatively regulates cell proliferation through acetylation of p53. Proc. Natl. Acad. Sci. USA 2001, 98, 9671–9676. [CrossRef]
177. Liu, Z.; Scannell, D.R.; Eisen, M.B.; Tjian, R. Control of Embryonic Stem Cell Lineage Commitment by Core Promoter Factor, TAF3. Cell 2011, 146, 720–731. [CrossRef]
178. Deato, M.D.E.; Marr, M.T.; Sottero, T.; Inouye, C.; Hu, P.; Tjian, R. MyoD Targets TAF3/TRF3 to Activate Myogenin Transcription. Mol. Cell 2008, 32, 96–105. [CrossRef]
179. Yao, J.; Fetter, R.D.; Hu, P.; Betzig, E.; Tjian, R. Subnuclear segregation of genes and core promoter factors in myogenesis. Genes Dev. 2011, 25, 569–580. [CrossRef] [PubMed]
180. Mandal, P.; Eswara, K.; Yerkesh, Z.; Kharchenko, V.; Zandarashvili, L.; Szczepski, K.; Bensaddek, D.; Jaremko, Ł.; Black, B.E.; Fischle, W. Molecular basis of hUHRF1 allosteric activation for synergistic histone modification binding by PI5P. Sci. Adv. 2022, 8, eabl9461. [CrossRef] [PubMed]
181. Relav, L.; Doghman-Bouguerra, M.; Ruggiero, C.; Muzzi, J.C.D.; Figueiredo, B.C.; Lalli, E. Steroidogenic Factor 1, a Goldilocks Transcription Factor from Adrenocortical Organogenesis to Malignancy. Int. J. Mol. Sci. 2023, 24, 3585. [CrossRef] [PubMed]
182. Krylova, I.N.; Sablin, E.P.; Moore, J.; Xu, R.X.;Waitt, G.M.; MacKay, J.A.; Juzumiene, D.; Bynum, J.M.; Madauss, K.; Montana, V.; et al. Structural Analyses Reveal Phosphatidyl Inositols as Ligands for the NR5 Orphan Receptors SF-1 and LRH-1. Cell 2005, 120, 343–355. [CrossRef] [PubMed]
183. Li, Y.; Choi, M.; Cavey, G.; Daugherty, J.; Suino, K.; Kovach, A.; Bingham, N.C.; Kliewer, S.A.; Xu, H. Crystallographic Identification and Functional Characterization of Phospholipids as Ligands for the Orphan Nuclear Receptor Steroidogenic Factor-1. Mol. Cell 2005, 17, 491–502. [CrossRef] [PubMed]
184. Wang, Y.-H.; Hariharan, A.; Bastianello, G.; Toyama, Y.; Shivashankar, G.V.; Foiani, M.; Sheetz, M.P. DNA damage causes rapid accumulation of phosphoinositides for ATR signaling. Nat. Commun. 2017, 8, 2118. [CrossRef]
185. Sobol, M.; Krausová, A.; Yildirim, S.; Kalasová, I.; Fáberová, V.; Vrkoslav, V.; Philimonenko, V.; Marášek, P.; Pastorek, L.; Cˇ apek, M.; et al. Nuclear phosphatidylinositol 4,5-bisphosphate islets contribute to efficient RNA polymerase II-dependent transcription. J. Cell Sci. 2018, 131, jcs211094. [CrossRef]
186. Gavgani, F.M.; Karlsson, T.; Tangen, I.L.; Morovicz, A.P.; Arnesen, V.S.; Turcu, D.C.; Ninzima, S.; Spang, K.; Krakstad, C.; Guillermet-Guibert, J.; et al. Nuclear upregulation of class I phosphoinositide 3-kinase p110 correlates with high 47S rRNA levels in cancer cells. J. Cell Sci. 2021, 134, jcs246090. [CrossRef] [PubMed]
187. Dyson, N.J. RB1: A prototype tumor suppressor and an enigma. Genes Dev. 2016, 30, 1492–1502. [CrossRef]
188. Kruiswijk, F.; Labuschagne, C.F.; Vousden, K.H. p53 in survival, death and metabolic health: A lifeguard with a licence to kill. Nat. Rev. Mol. Cell Biol. 2015, 16, 393–405. [CrossRef]
189. Divecha, N.; Roefs, M.; Los, A.; Halstead, J.; Bannister, A.; D’Santos, C. Type I PIPkinases Interact with and Are Regulated by the Retinoblastoma Susceptibility Gene Product—pRB. Curr. Biol. 2002, 12, 582–587. [CrossRef]
190. Los, A.P.; Vinke, F.P.; de Widt, J.; Topham, M.K.; van Blitterswijk,W.J.; Divecha, N. The Retinoblastoma Family Proteins Bind to and Activate Diacylglycerol Kinase . J. Biol. Chem. 2006, 281, 858–866. [CrossRef]
191. Los, A.P.; deWidt, J.; van Blitterswijk,W.J.; Divecha, N. Is there a role for diacylglycerol kinase in cell cycle regulation? Adv. Enzym. Regul. 2008, 48, 31–39. [CrossRef]
192. Choi, S.; Chen, M.; Cryns, V.L.; Anderson, R.A. A nuclear phosphoinositide kinase complex regulates p53. Nature 2019, 21, 462–475. [CrossRef] [PubMed]
193. Chen, M.; Choi, S.; Wen, T.; Chen, C.; Thapa, N.; Lee, J.H.; Cryns, V.L.; Anderson, R.A. A p53–phosphoinositide signalosome regulates nuclear AKT activation. Nature 2022, 24, 1099–1113. [CrossRef] [PubMed]
194. Ahmed, N.N.; Franke, T.F.; Bellacosa, A.; Datta, K.; Gonzalez-Portal, M.E.; Taguchi, T.; Testa, J.R.; Tsichlis, P.N. The proteins encoded by c-akt and v-akt differ in post-translational modification, subcellular localization and oncogenic potential. Oncogene 1993, 8, 1957–1963. [PubMed]
195. Neri, L.M.; Milani, D.; Bertolaso, L.; Stroscio, M.; Bertagnolo, V.; Capitani, S. Nuclear translocation of phosphatidylinositol 3-kinase in rat pheochromocytoma PC 12 cells after treatment with nerve growth factor. Cell. Mol. Biol. 1994, 40, 619–626.
196. Nguyen, T.L.X.; Choi, J.W.; Lee, S.B.; Ye, K.; Woo, S.-D.; Lee, K.-H.; Ahn, J.-Y. Akt phosphorylation is essential for nuclear translocation and retention in NGF-stimulated PC12 cells. Biochem. Biophys. Res. Commun. 2006, 349, 789–798. [CrossRef]
197. Stracker, T.H.; Petrini, J.H.J. The MRE11 complex: Starting from the ends. Nat. Rev. Mol. Cell Biol. 2011, 12, 90–103. [CrossRef]
198. Branzei, D.; Foiani, M. Regulation of DNA repair throughout the cell cycle. Nat. Rev. Mol. Cell Biol. 2008, 9, 297–308. [CrossRef]
199. Jones, D.R.; Foulger, R.; Keune, W.; Bultsma, Y.; Divecha, N. PtdIns5 P is an oxidative stress-induced second messenger that regulates PKB activation. FASEB J. 2012, 27, 1644–1656. [CrossRef]
200. Keune, W.-J.; Jones, D.R.; Bultsma, Y.; Sommer, L.; Zhou, X.Z.; Lu, K.P.; Divecha, N. Regulation of Phosphatidylinositol-5-Phosphate Signaling by Pin1 Determines Sensitivity to Oxidative Stress. Sci. Signal. 2012, 5, ra86. [CrossRef]
201. Bunce, M.W.; Boronenkov, I.V.; Anderson, R.A. Coordinated Activation of the Nuclear Ubiquitin Ligase Cul3-SPOP by the Generation of Phosphatidylinositol 5-Phosphate. J. Biol. Chem. 2008, 283, 8678–8686. [CrossRef]
202. Xu, L.; Wei, Y.; Reboul, J.; Vaglio, P.; Shin, T.-H.; Vidal, M.; Elledge, S.J.; Harper, J.W. BTB proteins are substrate-specific adaptors in an SCF-like modular ubiquitin ligase containing CUL-3. Nature 2003, 425, 316–321. [CrossRef]
203. Geyer, R.;Wee, S.; Anderson, S.; Yates, J.;Wolf, D.A. BTB/POZ Domain Proteins Are Putative Substrate Adaptors for Cullin 3 Ubiquitin Ligases. Mol. Cell 2003, 12, 783–790. [CrossRef] [PubMed]
204. Li, X.-M.;Wu, H.-L.; Xia, Q.-D.; Zhou, P.;Wang, S.-G.; Yu, X.; Hu, J. Novel insights into the SPOP E3 ubiquitin ligase: From the regulation of molecular mechanisms to tumorigenesis. Biomed. Pharmacother. 2022, 149, 112882. [CrossRef] [PubMed]
205. Okada, M.; Jang, S.-W.; Ye, K. Akt phosphorylation and nuclear phosphoinositide association mediate mRNA export and cell proliferation activities by ALY. Proc. Natl. Acad. Sci. USA 2008, 105, 8649–8654. [CrossRef]
206. Mohan, N.; AP, S.; Francis, N.; Anderson, R.; Laishram, R.S. Phosphorylation regulates the Star-PAP-PIPKI interaction and directs specificity toward mRNA targets. Nucleic Acids Res. 2015, 43, 7005–7020. [CrossRef]
207. Gonzales, M.L.; Mellman, D.L.; Anderson, R.A. CKI Is Associated with and Phosphorylates Star-PAP and Is Also Required for Expression of Select Star-PAP Target Messenger RNAs. J. Biol. Chem. 2008, 283, 12665–12673. [CrossRef]
208. Zhang, L.; Zhang, Y.; Chen, Y.; Gholamalamdari, O.; Wang, Y.; Ma, J.; Belmont, A.S. TSA-seq reveals a largely conserved genome organization relative to nuclear speckles with small position changes tightly correlated with gene expression changes. Genome Res. 2020, 31, 251–264. [CrossRef] [PubMed]
209. Xu, A.; Suh, P.-G.; Marmy-Conus, N.; Pearson, R.B.; Seok, O.Y.; Cocco, L.; Gilmour, R.S. Phosphorylation of Nuclear Phospholipase C 1 by Extracellular Signal-Regulated Kinase Mediates the Mitogenic Action of Insulin-Like Growth Factor I. Mol. Cell. Biol. 2001, 21, 2981–2990. [CrossRef]
210. Xu, A.; Wang, Y.; Xu, L.Y.; Gilmour, R.S. Protein Kinase C -mediated Negative Feedback Regulation Is Responsible for the Termination of Insulin-like Growth Factor I-induced Activation of Nuclear Phospholipase C 1 in Swiss 3T3 Cells. J. Biol. Chem. 2001, 276, 14980–14986. [CrossRef]
