Heavy metal pollution is a severe concern worldwide, owing to its harmful effects on ecosystems. Phytoremediation has been applied to remove heavy metals from water, soils, and sediments by using plants and associated microorganisms to restore contaminated sites. The Typha genus is one of the most important genera used in phytoremediation strategies because of its rapid growth rate, high biomass production, and the accumulation of heavy metals in its roots. Plant growth-promoting rhizobacteria have attracted much attention because they exert biochemical activities that improve plant growth, tolerance, and the accumulation of heavy metals in plant tissues. Because of their beneficial effects on plants, some studies have identified bacterial communities associated with the roots of Typha species growing in the presence of heavy metals.
- heavy metal tolerance
- bacterial diversity
- phytoremediation
- root
1. Bacteria Associated with the Rhizosphere of Typha spp.
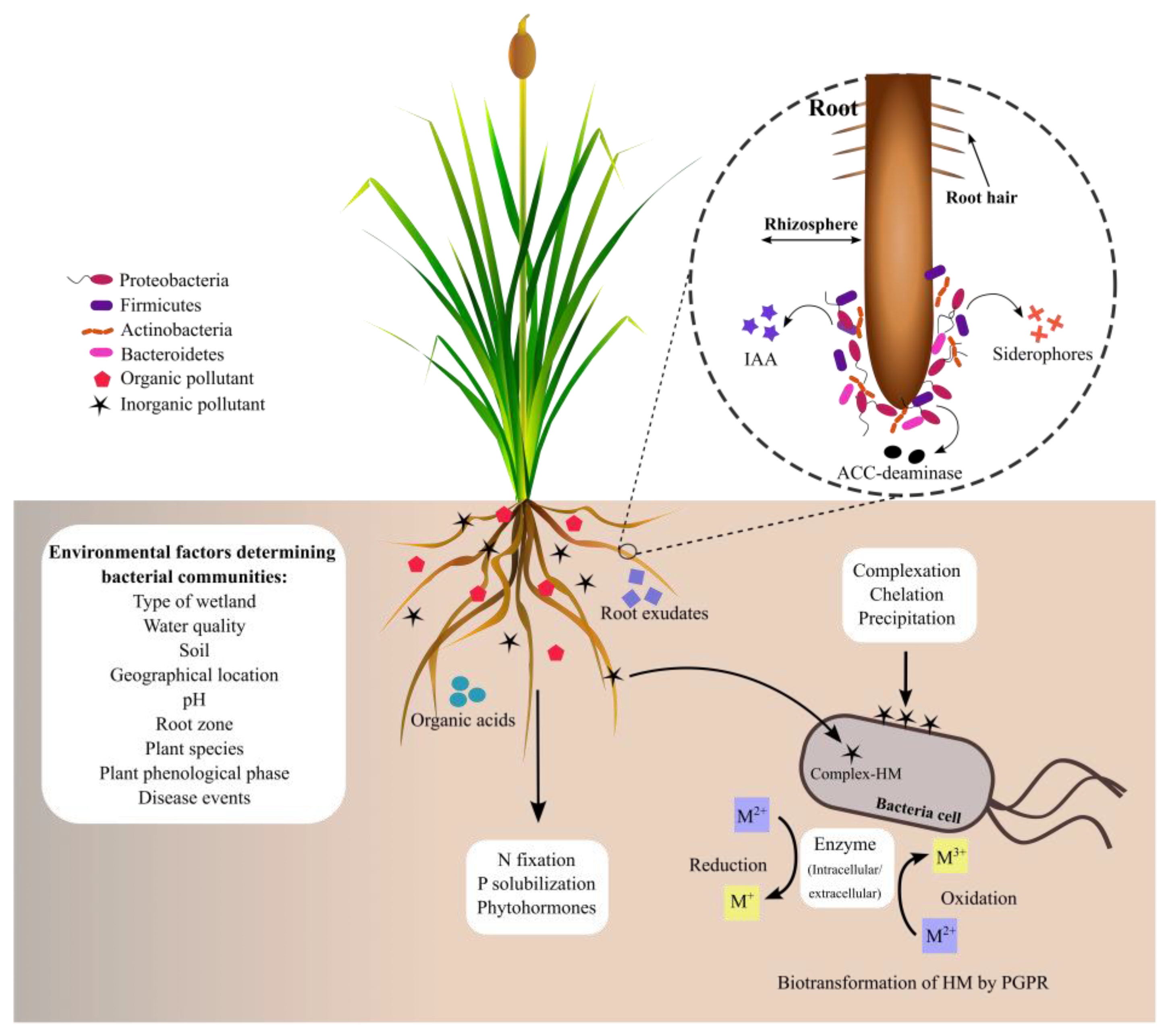
2. Bacterial Communities Associated with Typha Roots in Natural Environments
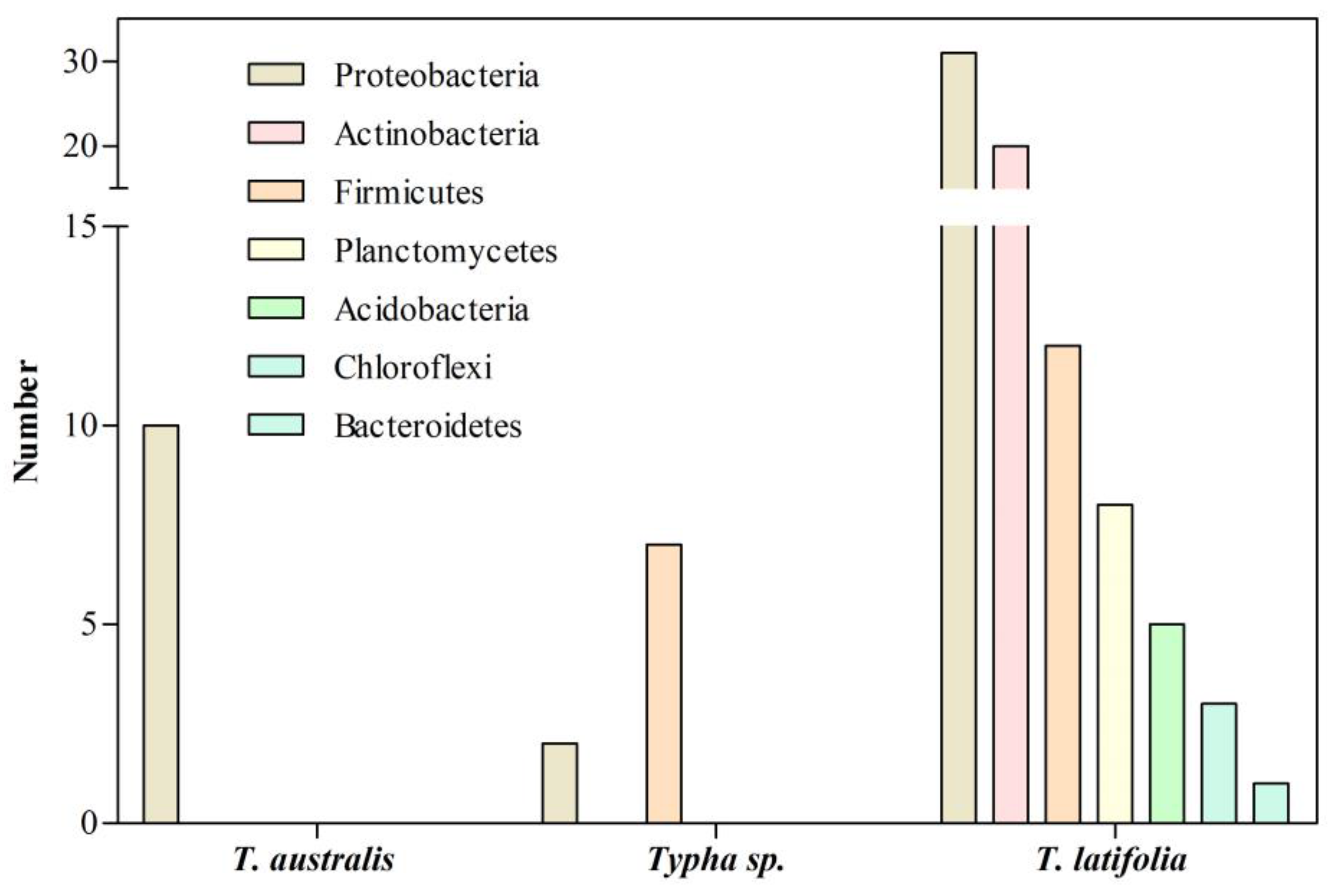
3. Bacterial Communities Associated with the Roots of Typha Exposed to HMs
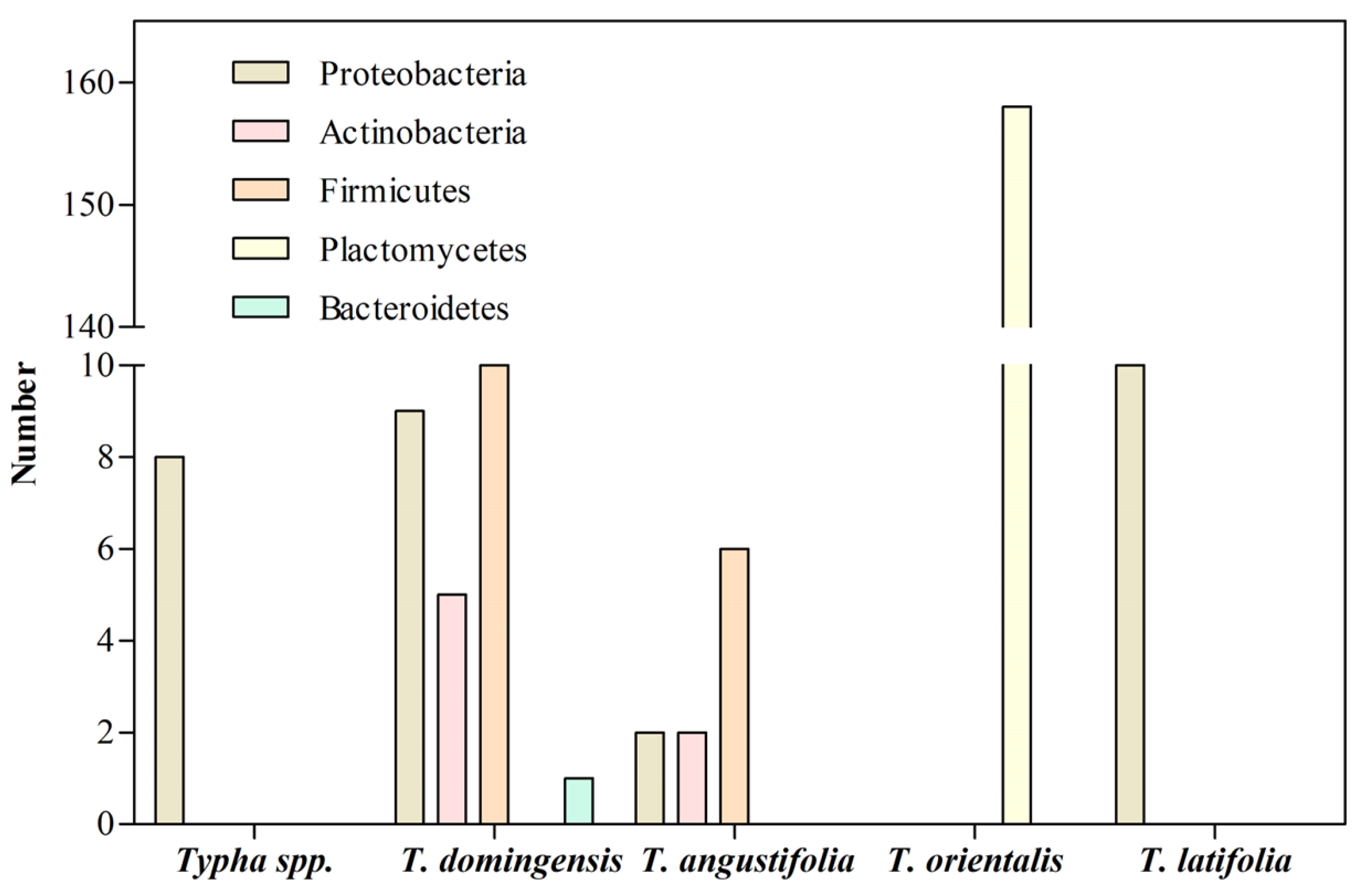
| Specie | Phylum | Metal | Site | References |
|---|---|---|---|---|
| Typha sp. | Proteobacteria | Cr | Wetland | [69][14] |
| T. domingensis | Proteobacteria Firmicutes Actinobacteria Bacteroidetes |
Cr, Ni, Fe | Pond and stream | [106][15] |
| T. angustifolia | Firmicutes Proteobacteria Actinobacteria |
Fe | Wetland | [64][16] |
| T. orientalis | Planctomycetes Uncultured bacterium |
Cu, Zn, Pb | Lake | [107][17] |
| T. latifolia | Proteobacteria | Cd | Contaminated site | [53][11] |
4. Other Microorganisms Associated with the Rhizosphere of Typha spp.
The rhizosphere is the habitat of many microorganisms and invertebrates, considered one of Earth’s most dynamic interfaces [104][9]. Although bacteria are the main inhabitants of the rhizosphere, the plant roots can be colonized by fungi, protozoa, rotifers, nematodes, and microarthropods, which are attracted by rhizodeposits, nutrients, exudates, border cells, and mucilage released by the plant root [100][4]. The presence of endophytic fungi has been reported in T. latifolia roots, including Aspergillus sp., Myrothecium sp., Phoma sp., Penicillium sp., Acremonium sp., and Fusarium sp., while in its rhizome Penicillium sp., Myrothecium sp., and Fusarium sp. have been found [173][36]. Furthermore, the efficiency of Aspergillus niger, Acremonium sp., Aureobasidium sp., Cephalosporium sp., and Fusarium sp., to degrade pollutants or absorb HMs has been demonstrated; thus, they are an option in mycoremediation [173,174][36][37]. On the other hand, a study conducted in Zhejiang Province, Eastern China, by Guan et al. [175][38] reported the presence of common fungal species including Pleosporales sp., Teratosphaeria microspora, Geotrichum candidum, Engyodontium album, Blastocladiales sp., uncultured soil fungus, Fusarium graminearum, and Cladosporium bruhnei in the rhizosphere of T. latifolia and T. orientalis. While Paramicrosporidium saccamoebae, Rhynchosporium secalis, Acremonium roseolum, and Cladosporium sphaerospermun were found in T. latifolia, and uncultured soil fungus in T. orientalis. Finally, arbuscular vesicular (AV) fungi are known to increase plant resistance to heavy metals, and this differs according to fungal species, plant, and environmental conditions [176][39].References
- Gao, T.; Shi, X.-Y. Taxonomic Structure and Function of Seed-Inhabiting Bacterial Microbiota from Common Reed (Phragmites australis) and Narrowleaf Cattail (Typha angustifolia L.). Arch. Microbiol. 2018, 200, 869–876.
- Li, Y.H.; Zhu, J.N.; Liu, Q.F.; Liu, Y.; Liu, M.; Liu, L.; Zhang, Q. Comparison of the Diversity of Root-Associated Bacteria in Phragmites australis and Typha angustifolia L. in Artificial Wetlands. World J. Microbiol. Biotechnol. 2013, 29, 1499–1508.
- Arroyo, P.; de Miera, L.E.S.; Ansola, G. Influence of Environmental Variables on the Structure and Composition of Soil Bacterial Communities in Natural and Constructed Wetlands. Sci. Total Environ. 2015, 506–507, 380–390.
- Kennedy, A.C.; de Luna, L.Z. Rhizosphere. In Encyclopedia of Soils in the Environment; Hillel, D., Ed.; Elsevier: Oxford, UK, 2005; pp. 399–406. ISBN 978-0-12-348530-4.
- Jha, P.N.; Kumar, A. Endophytic Colonization of Typha australis by a Plant Growth-promoting Bacterium Klebsiella oxytoca Strain GR-3. J. Appl. Microbiol. 2007, 103, 1311–1320.
- Ashkan, M.F.; Bleakley, B. Isolation, Characterization and Identification of Putative Bacterial Endophytes from Some Plants in Hot Springs, South Dakota. Int. J. Curr. Microbiol. Appl. Sci. 2017, 6, 756–767.
- Pietrangelo, L.; Bucci, A.; Maiuro, L.; Bulgarelli, D.; Naclerio, G. Unraveling the Composition of the Root-Associated Bacterial Microbiota of Phragmites australis and Typha latifolia. Front. Microbiol. 2018, 9, 1650.
- Lagos, L.; Maruyama, F.; Nannipieri, P.; Mora, M.L.; Ogram, A.; Jorquera, M.A. Current Overview on the Study of Bacteria in the Rhizosphere by Modern Molecular Techniques: A Mini-review. J. Soil Sci. Plant Nutr. 2015, 15, 504–523.
- Philippot, L.; Raaijmakers, J.M.; Lemanceau, P.; van der Putten, W.H. Going Back to the Roots: The Microbial Ecology of the Rhizosphere. Nat. Rev. Microbiol. 2013, 11, 789–799.
- Inouhe, M.; Huang, H.; Chaudhary, S.K.; Gupta, D.K. Heavy Metal Bindings and Their Interactions with Thiol Peptides and Other Biological Ligands in Plant Cells. In Metal Toxicity in Plants: Perception, Signaling and Remediation; Gupta, D.K., Sandalio, L.M., Eds.; Springer: Berlin/Heidelberg, Germany, 2012; pp. 1–21. ISBN 978-3-642-22081-4.
- Rolón-Cárdenas, G.A.; Arvizu-Gómez, J.L.; Pacheco-Aguilar, J.R.; Vázquez-Martínez, J.; Hernández-Morales, A. Cadmium-Tolerant Endophytic Pseudomonas rhodesiae Strains Isolated from Typha latifolia Modify the Root Architecture of Arabidopsis thaliana Col-0 in Presence and Absence of Cd. Braz. J. Microbiol. 2021, 52, 349–361.
- Ponce-Hernández, A.; Maldonado-Miranda, J.; Medellin-Castillo, N.; Alonso-Castro, A.; Carranza Alvarez, C. Phytoremediation Technology: Sustainable Solution for Cleaning Up of Recalcitrant Pollutants from Disturbed Environs. In Bioremediation and Biotechnology, Vol 3: Persistent and Recalcitrant Toxic Substances; Springer International Publishing: Cham, Switzerland, 2020; pp. 245–268. ISBN 978-3-030-46074-7.
- Wu, Y.; Ma, L.; Liu, Q.; Vestergård, M.; Topalovic, O.; Wang, Q.; Zhou, Q.; Huang, L.; Yang, X.; Feng, Y. The Plant-Growth Promoting Bacteria Promote Cadmium Uptake by Inducing a Hormonal Crosstalk and Lateral Root Formation in a Hyperaccumulator Plant Sedum alfredii. J. Hazard. Mater. 2020, 395, 122661.
- Pacheco-Aguilar, J.R.P.; Peña-Cabriales, J.J.P.; Maldonado-Vega, M.M. Identification and Characterization of Sulfur-Oxidizing Bacteria in an Artificial Wetland That Treats Wastewater From a Tannery. Int. J. Phytoremed. 2008, 10, 359–370.
- Shehzadi, M.; Fatima, K.; Imran, A.; Mirza, M.S.; Khan, Q.M.; Afzal, M. Ecology of Bacterial Endophytes Associated with Wetland Plants Growing in Textile Effluent for Pollutant-Degradation and Plant Growth-Promotion Potentials. Plant Biosyst. Int. J. Deal. All Asp. Plant Biol. 2016, 150, 1261–1270.
- Saha, C.; Mukherjee, G.; Agarwal-Banka, P.; Seal, A. A Consortium of Non-Rhizobial Endophytic Microbes from Typha angustifolia Functions as Probiotic in Rice and Improves Nitrogen Metabolism. Plant Biol. 2016, 18, 938–946.
- Zhou, X.; Zhang, J.; Wen, C. Community Composition and Abundance of Anammox Bacteria in Cattail Rhizosphere Sediments at Three Phenological Stages. Curr. Microbiol. 2017, 74, 1349–1357.
- Chellaiah, E.R. Cadmium (Heavy Metals) Bioremediation by Pseudomonas Aeruginosa: A Minireview. Appl. Water Sci. 2018, 8, 154.
- Fakhar, A.; Gul, B.; Gurmani, A.R.; Khan, S.M.; Ali, S.; Sultan, T.; Chaudhary, H.J.; Rafique, M.; Rizwan, M. Heavy Metal Remediation and Resistance Mechanism of Aeromonas, Bacillus, and Pseudomonas: A Review. Crit. Rev. Environ. Sci. Technol. 2022, 52, 1868–1914.
- Renu, S.; Sarim, K.M.; Singh, D.P.; Sahu, U.; Bhoyar, M.S.; Sahu, A.; Kaur, B.; Gupta, A.; Mandal, A.; Thakur, J.K.; et al. Deciphering Cadmium (Cd) Tolerance in Newly Isolated Bacterial Strain, Ochrobactrum intermedium BB12, and Its Role in Alleviation of Cd Stress in Spinach Plant (Spinacia oleracea L.). Front. Microbiol. 2022, 12, 758144.
- Pandey, S.; Ghosh, P.K.; Ghosh, S.; De, T.K.; Maiti, T.K. Role of Heavy Metal Resistant Ochrobactrum sp. and Bacillus spp. Strains in Bioremediation of a Rice Cultivar and Their PGPR like Activities. J. Microbiol. 2013, 51, 11–17.
- Faisal, M.; Hasnain, S. Beneficial Role of Hydrophytes in Removing Cr(VI) from Wastewater in Association with Chromate-Reducing Bacterial Strains Ochrobactrum intermedium and Brevibacterium. Int. J. Phytoremed. 2005, 7, 271–277.
- Kavita, B.; Keharia, H. Reduction of Hexavalent Chromium by Ochrobactrum intermedium BCR400 Isolated from a Chromium-Contaminated Soil. 3 Biotech 2012, 2, 79.
- Ozdemir, G.; Ozturk, T.; Ceyhan, N.; Isler, R.; Cosar, T. Heavy Metal Biosorption by Biomass of Ochrobactrum anthropi Producing Exopolysaccharide in Activated Sludge. Bioresour. Technol. 2003, 90, 71–74.
- Bhattacharya, A.; Gupta, A. Evaluation of Acinetobacter sp. B9 for Cr (VI) Resistance and Detoxification with Potential Application in Bioremediation of Heavy-Metals-Rich Industrial Wastewater. Environ. Sci. Pollut. Res. 2013, 20, 6628–6637.
- Pang, B.; Lv, L.; Pang, C.; Ye, F.; Shang, C. Optimization of Growth Conditions of Acinetobacter sp. Cr1 for Removal of Heavy Metal Cr Using Central Composite Design. Curr. Microbiol. 2021, 78, 316–322.
- Zakaria, Z.A.; Zakaria, Z.; Surif, S.; Ahmad, W.A. Hexavalent Chromium Reduction by Acinetobacter haemolyticus Isolated from Heavy-Metal Contaminated Wastewater. J. Hazard. Mater. 2007, 146, 30–38.
- Ndeddy Aka, R.J.; Babalola, O.O. Effect of Bacterial Inoculation of Strains of Pseudomonas aeruginosa, Alcaligenes feacalis and Bacillus subtilis on Germination, Growth and Heavy Metal (Cd, Cr, and Ni) Uptake of Brassica juncea. Int. J. Phytoremed. 2016, 18, 200–209.
- Sodhi, K.K.; Kumar, M.; Singh, D.K. Multi-Metal Resistance and Potential of Alcaligenes sp. MMA for the Removal of Heavy Metals. SN Appl. Sci. 2020, 2, 1885.
- Liu, W.; Wang, Q.; Wang, B.; Hou, J.; Luo, Y.; Tang, C.; Franks, A.E. Plant Growth-Promoting Rhizobacteria Enhance the Growth and Cd Uptake of Sedum plumbizincicola in a Cd-Contaminated Soil. J. Soils Sediments 2015, 15, 1191–1199.
- Salam, L.B.; Shomope, H.; Ummi, Z.; Bukar, F. Mercury Contamination Imposes Structural Shift on the Microbial Community of an Agricultural Soil. Bull. Natl. Res. Cent. 2019, 43, 163.
- Li, D.; Chen, J.; Zhang, X.; Shi, W.; Li, J. Structural and Functional Characteristics of Soil Microbial Communities in Response to Different Ecological Risk Levels of Heavy Metals. Front. Microbiol. 2022, 13, 1072389.
- Zhang, J.; Shi, Q.; Fan, S.; Zhang, Y.; Zhang, M.; Zhang, J. Distinction between Cr and Other Heavy-Metal-Resistant Bacteria Involved in C/N Cycling in Contaminated Soils of Copper Producing Sites. J. Hazard. Mater. 2021, 402, 123454.
- Rolón-Cárdenas, G.A.; Martínez-Martínez, J.G.; Arvizu-Gómez, J.L.; Soria-Guerra, R.E.; Alfaro-De la Torre, M.C.; Alatorre-Cobos, F.; Rubio-Santiago, J.; González-Balderas, R.D.M.; Carranza-Álvarez, C.; Macías-Pérez, J.R.; et al. Enhanced Cd-Accumulation in Typha latifolia by Interaction with Pseudomonas rhodesiae GRC140 under Axenic Hydroponic Conditions. Plants 2022, 11, 1447.
- Rubio-Santiago, J.; Hernández-Morales, A.; Rolón-Cárdenas, G.A.; Arvizu-Gómez, J.L.; Soria-Guerra, R.E.; Carranza-Álvarez, C.; Rubio-Salazar, J.E.; Rosales-Loredo, S.; Pacheco-Aguilar, J.R.; Macías-Pérez, J.R.; et al. Characterization of Endophytic Bacteria Isolated from Typha latifolia and Their Effect in Plants Exposed to Either Pb or Cd. Plants 2023, 12, 498.
- Khan, A.A.H. Endophytic Fungi and Their Impact on Agroecosystems. In Medicinal Plants: Biodiversity, Sustainable Utilization and Conservation; Khasim, S.M., Long, C., Thammasiri, K., Lutken, H., Eds.; Springer: Singapore, 2020; pp. 443–499. ISBN 9789811516368.
- Yadav, A.; Goyal, D.; Prasad, M.; Singh, T.B.; Shrivastav, P.; Ali, A.; Dantu, P.K. Bioremediation of Toxic Pollutants: Features, Strategies, and Applications. In Contaminants in Agriculture: Sources, Impacts and Management; Naeem, M., Ansari, A.A., Gill, S.S., Eds.; Springer International Publishing: Cham, Switzerland, 2020; pp. 361–383. ISBN 978-3-030-41552-5.
- Guan, M.; Pan, X.-C.; Wang, S.; Wei, X.-L.; Zhang, C.-B.; Wang, J.; Liu, W.-L.; Liu, S.-Y.; Chang, J. Comparison of Fungal Communities among Ten Macrophyte Rhizospheres. Fungal Biol. 2018, 122, 867–874.
- Cheng, S. Effects of Heavy Metals on Plants and Resistance Mechanisms. Environ. Sci. Pollut. Res. 2003, 10, 256–264.
