Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 2 by Lindsay Dong and Version 1 by Manuel Garcia-Herreros.
Genomic and phenotypic selection criteria have been crucial in dairy cattle. Udder health and milk production are important factors affecting productivity in dairy cattle. Furthermore, genomic and phenotypic selection are essential tools for increasing milk supply for human consumption, decreasing the use of antimicrobial products, improving animal health and welfare, and developing efficient dairy cattle production systems.
- genomic analysis
- phenotypic traits
- evaluation criteria
- milk production
- health
- longevity
- dairy cattle
1. Introduction
For decades, dairy cattle breeding has been focused on increasing milk production [1]. Many functional traits have been described as having negative genomic correlations with milk production as well as reductions in genetic merit for health and fitness [2]. However, one of the constant goals and challenges in high-producing dairy cattle has been to balance fertility, udder health, and metabolic diseases without affecting milk production or compromising animal welfare [1,2,3,4][1][2][3][4].
Genomic and phenotypic evaluation methods provide accurate and efficient evaluations of udder traits in dairy cattle, allowing farmers to make more informed breeding decisions for animal selection. Phenotypic evaluation methods use physical measurements and observations to evaluate udder traits, providing a more holistic evaluation of the cow’s overall health and productivity. On the other hand, genomic evaluation methods use DNA sequencing to identify specific genetic markers associated with desirable udder traits, providing a more direct and precise evaluation.
Combining genomic and phenotypic evaluation methods can provide a complete picture of a cow’s udder health and milk production potential, allowing for more effective breeding and selection decisions. Furthermore, using genomic and phenotypic evaluation methods can ultimately lead to improved udder health, milk production, and overall profitability for dairy farmers and enhanced quality of dairy products for consumers.
In the dairy cow, different factors, such as udder conformation and size and environmental conditions, contribute to variations in cow performance and functionality, mainly in longevity and productivity [5]. In recent years, dairy cattle selection has been based on female-derived traits such as milk production performance and linear conformation traits [1]. Several of these linear conformation traits are negatively correlated with other functional traits, such as milk production, which has led to a reduction in health, body condition, and adaptability-derived traits [6]. Over the last 10–20 years, other traits related to udder health, such as somatic cell counts (SCCs), productive life (longevity), and other reproductive-related traits, such as daughter fertility, have been incorporated [3]. These traits have become increasingly important in the dairy industry, and recently, genomic selection has created new opportunities for increased accuracy of the selection of such traits in very young animals [3,7][3][7]. In other ruminant species, highly efficient selection schemes were established based on pyramidal population management with core breeders at the top. The selection schemes were based on pedigree and official milk records, artificial insemination results, controlled mating, and the estimation of breeding values with the aim of accelerating genetic progress [1]. On the other hand, the constant participation of young bulls in testing and breeding programs has been described in different dairy herds in the U.S. [8]. For bull selection, udder conformation and the expected difference in milk production were considered the most essential traits. However, traditionally, the most important traits for cow selection were milk production followed by udder conformation, feet and legs, and fat percentage [8]. Nowadays, the need to preserve particular traits of economic interest for milk production and to ensure positive genetic progress is one of the main objectives [9,10,11][9][10][11]. Therefore, current selection programs require specific breeding plans that take into account milk production and functional traits, including those related to udder conformation and functionality [12].
Moreover, the heritability assessment and genetic correlations between traits and selection indices allow identifying the most suitable individuals for selection in dairy breeds [11,13][11][13]. Poor udder and teat conformation has been reported to reduce profitability in dairy herds [14]. Furthermore, this has an impact on the incidence of mastitis at calving and leads to decreased productivity during the lifetime of the cow [9]. Therefore, it is necessary to evaluate the effects of udder conformation on performance and future progeny.
2. Udder Evaluation Criteria
Udder evaluation has been a part of dairy cattle selection for centuries, with early breeders selecting cows based on visual inspection and manual palpation of the udder. In the early 1900s, standardized methods for udder evaluation were developed, including the linear score system and the udder index score. Dairy cattle breed associations have been developing classification systems since 1929, when the Holstein-Friesian Association of America introduced the first official system, intending to utilize these systems as phenotypic or genetic indicators of milk production ability, longevity, and udder health and milk production capacity over time [2,3,4,5,7,8,9][2][3][4][5][7][8][9]. In the mid-1900s, milk production testing became a common method for evaluating udder health and milk quality, with farmers tracking the amount and quality of milk produced by each cow. The establishment of modern livestock breeding programs dates back to the 1960s, when nucleus breeding initiatives emerged that utilized official pedigree data and milk production records to drive genetic progress [3]. While increasing milk production and improving quality (such as fat and protein content) continues to be the top priority for profitability, livestock breeders have also begun considering other important traits for selection. These may include mechanical milking ability, udder health, disease resistance (e.g., against mastitis and internal parasites), and even the nutritional value of milk (such as fatty acid composition) [1,15][1][15]. Given the importance of considering a range of traits beyond just milk production and quality, the dairy industry has increasingly focused on developing comprehensive selection criteria that consider factors such as udder health, disease resistance, and mechanical milking ability. One example of this is the Holstein Association of Canada’s efforts to collect data on linear conformation traits in dairy cattle from the 1970s to the 1980s as a means of improving selection criteria based on type-derived traits [16]. The phenotypic linear traits considered were general appearance, milk form, body capacity, rump, feet and legs, mammary system, fore udder, and rear udder [16,17][16][17]. On the other hand, nine descriptive traits were also considered: stature, size, style, brisket, floor, loin strength, dorsal width, and pin placement [12,17][12][17]. Heritability estimates for type traits in dairy cattle ranged from 0.06 (style) to 0.33 (size). Phenotypic correlations between type traits and calving interval were essentially zero. The highest beneficial negative genetic correlations with calving interval were for breast depth (−0.42), rear udder (−0.37), and capacity (−0.34), and the highest antagonistic positive correlations were for milk yield (0.43) and milk quality traits (0.38) [17]. In the late 1900s and early 2000s, genomic evaluation methods were developed, allowing for more accurate and efficient evaluations of udder traits. Today, a combination of visual inspection, manual palpation, milk production testing, and genomic and phenotypic evaluation methods are becoming widespread to evaluate udder traits in dairy cattle. Thus, current breeding programs based on the traditional quantitative approach have achieved appreciable genetic gains for milk production. They have implemented selection for other traits, including milk composition, udder morphology, somatic cell count, and mastitis resistance [1,18][1][18]. The implementation of novel selection strategies utilizing molecular-based data to enhance both conventional and high-value traits holds significant promise for advancing functional improvements in dairy breeds [1].3. Evaluation Criteria Based on Udder Phenotypic Traits
Phenotypic evaluation has long been a key component of dairy cattle breeding, allowing farmers and breeders to evaluate a cow’s physical traits and characteristics to make informed breeding decisions. While more recent advancements in genomic evaluation have provided new tools for evaluating an animal’s genetic potential, phenotypic evaluation remains a critical aspect of the breeding process, particularly when assessing traits that cannot be measured through DNA analysis. However, several intrinsic and extrinsic factors should be taken into account together with the phenotypic and genomic udder evaluation methods for dairy cattle selection (Figure 1). Understanding the criteria used to assess udder phenotypic traits is an essential aspect of dairy cattle breeding, as these traits are key indicators of a cow’s milk production potential and overall udder health. The selection criteria for productive animals based on udder conformation include several traits associated with the general linear conformation of females, such as stature, strength, milk production, teat diameter, hind legs (lateral view), rump angle, rump width, anterior udder attachment, posterior udder height, posterior udder arch, udder depth, suspensory ligament, and teat placement [12]. The selection criteria based on udder conformation traits could differ depending on the livestock species, breed, or crossbreed evaluated. In one study, Montbéliarde (MO) × Holstein (HO) and Viking Red (VR) × HO crossbred cows were compared with first-calving pure HO cows to evaluate the conformation for production and reproductive traits [24][19]. It was observed that crossbred cows had less udder-to-hock clearance than HO cows. In addition, crossbred cows had more distance between the front and rear teats and longer teats than HO cows [24,25][19][20]. However, the frequency of first lactation cows that were discarded because of udder conformation issues was uniformly low across all groups (<1%) [24][19].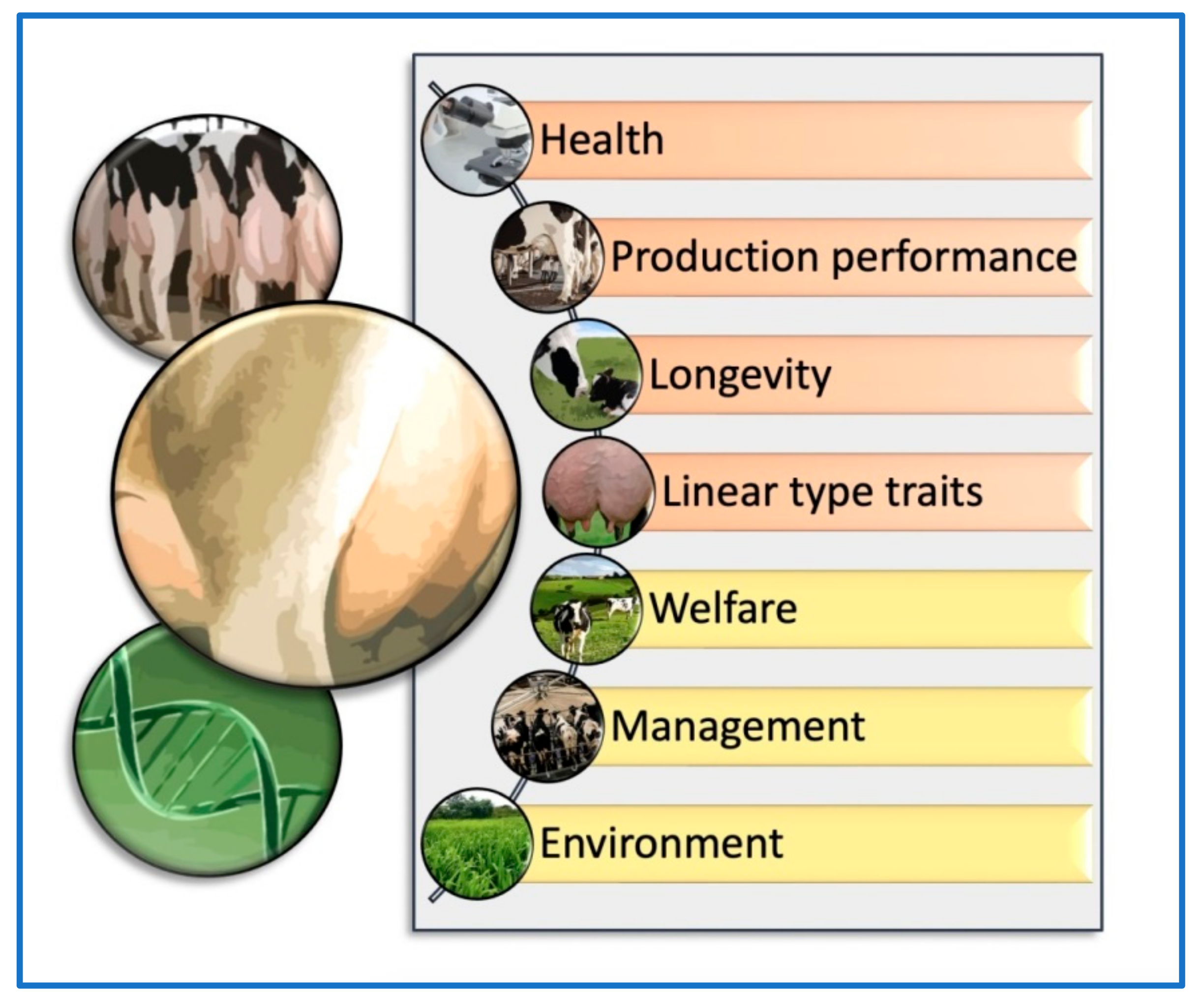

Figure 1. Brief scheme of the main elements that should be considered together with the phenotypic and genomic udder evaluation methods for dairy cattle selection. Orange: intrinsic components such as health, production, longevity, and linear (conformation) type traits. Yellow: extrinsic components such as welfare, management, and environment.
Among the selection criteria taken into account in cattle, the direct measuring and correlated responses to a single trait for milk production could be a convenient method in dairy production systems [26][21]. For example, several producers use a selection line consisting of artificial insemination (AI) bulls selected for their high estimated transmission capacity for milk production associated with udder conformation [27][22]. On the other hand, several studies currently target the genetic evaluation of novel traits based on methods to predict selection accuracy. These methods are based on reference populations of cows using indicator traits that could increase the accuracy of genomic selection for important traits such as udder health [3].
4. Evaluation Criteria Based on Udder Genomic Traits
As genomic evaluation methods have become more prevalent in dairy cattle breeding, understanding the criteria used to evaluate udder genomic traits has become increasingly crucial for breeders seeking to make informed decisions about animal selection. The progress of genetic and genomic evaluation in dairy cattle is based on discovering new traits of interest for predicting selection accuracy. Thus, the genetic and genomic assessment aims to improve how using indicator traits could increase the accuracy of genomic selection [3]. To develop effective selection programs for new traits, it is necessary to create large databases based on highly reliable estimated genetic values (EGVs) [2,35,36][2][23][24].
When analyzing genetic data, a positive correlation was found between Somatic Cell Score (SCS) and milk quality and composition traits, highlighting the interconnected nature of these characteristics [11,20,23][11][25][26]. The genetic traits for milk composition were negatively correlated with the udder conformation (−0.40) and with rear muscling and udder volume (−0.28) [11]. These results were obtained by using the current selection index, which mainly focuses on dairy strength and could negatively influence all other traits [11].
All data based on genome sequences from previous generations provides comprehensive information on the polymorphic loci that characterize a given breed and differentiate it from others. Association analysis with imputed sequences, mainly when applied to multiple traits simultaneously, is a powerful approach to detecting candidate causal variants underlying complex phenotypes. Twelve quantitative trait loci (QTL) affecting different morphological characteristics of the mammary gland were detected in the German Fleckvieh cattle population. Most of the QTLs were located in non-coding regions of the genome but in close proximity to candidate genes that could be involved in mammary gland morphology (SP5, GC, NPFFR2, CRIM1, RXFP2, TBX5, RBM19, and ADAM12) [39][27]. In Holstein cattle, genetic analyses of udder conformation traits have been developed [19][28]. A genome-wide association has been described with five udder conformation traits, including anterior udder attachment, central suspensory ligament, posterior udder attachment height, posterior udder attachment width, and udder depth [19][28]. Heritability and standard errors for these five udder traits ranged from 0.04 ± 0.00 to 0.49 ± 0.03. In thise study, phenotypic data were measured on 1000 Holstein cows, and the GeneSeek Genomic Profiler (GGP) Bovine 100 K SNP chip was used to analyze genotypic data [19][28]. Numerous candidate genes were identified within 200 kb of significant SNPs. Among all significant SNPs associated with udder conformation traits, most of them were located within the following genes: Microsomal Glutathione S-Transferase 1 (MGST1), Microsomal Glutathione S-Transferase 2 (MGST2), Microtubule-associated scaffold protein 1 (MTUS1), LOC101903734, LOC112447118, Structural maintenance of chromosomes protein 5 and 6 (SMC5-SMC6), Parkin RBR E3 ubiquitin-protein ligase (PRKN), Syntaxin-Binding Protein 6 (STXBP6), glutamate ionotropic receptor delta-type subunit 2 (GRID2), E2F transcription factor 8 (E2F8), Cadherin 11 (CDH11), Forkhead Box P1 (FOXP1), Localisation factor complex 1 (SLF1), transmembrane protein 117 (TMEM117), SET binding factor 2 (SBF2), CGPVitamin D binding protein (GC), galectin 2 (LGALS2), adhesion G protein-coupled receptor B3 (ADGRB3), and Glutamate-Cysteine Ligase Catalytic Subunit (GCLC) as well as KEGG pathway genes. While one of the SNPs on Chr6 was close (50 kb) to the ubiquitin-conjugating enzyme E2 K gene (UBE2K), three SNPs on Chr21, Chr29, and Chr18 were located close (100 kb) to STXBP6, E2F8, and CDH11, respectively [19][28].
5. Evaluation Criteria Based on Genomic and Phenotypic Traits: Udder Health, Production, and Longevity
5.1. Genomic, Genotypic, and Phenotypic Udder Traits: Impact on Mammary Gland Health
The development of national health registration programs began several decades ago. Accordingly, the first reports of genetic results based on health traits in the United States started in 2004 [41][29]. A few years later, in 2007, Canada reported large amounts of data on mastitis incidence in cows [42,43][30][31]. Available data based on the mastitis incidence can be analyzed by multi-trait models using SCS data and other derived traits indicative of SCS in early lactation, partial SCS, test day SCS, anterior udder attachment, udder depth, and BCS to perform traditional and genomic evaluations for mastitis resistance [35,43,44][23][31][32]. Udder health traits associated with bacteriological testing of quarters maximize the information on infection; however, these tests are not practical on a population scale [45][33]. Nevertheless, several milk enzymes are possible indicators of tissue damage at the mammary gland level [45][33]. The heritabilities are around 0.2 for SCCs and 0.1 for other traits, reflecting genetic variations in the immunological defenses and their effect on the teat, phagocytosis, and immune response [45][33]. There is a positive relationship between somatic cell count (SCC) and milk yield at the genetic level but a strong negative relationship between them at the phenotypic level [45][33]. Genetic correlation refers to the degree to which the same genes affect two different traits, in this case, SCC and milk yield. Furthermore, it was observed that the CHL1 gene is up-regulated with stress levels and influences the mechanisms of the immune system [49][34]. Therefore, the activity interactions between the genes may fluctuate depending on environmental stressors. On the other hand, other authors report that CFAP69, STEAP2, and ITGB3BP genes located on chromosome 4 are related to the mechanisms of the mammary gland immune system [50,51][35][36]. The objective measures of mammary gland conformation and linear type scores are used to predict mastitis in experimental linear classification programs [1]. However, the relationships between conformation and mastitis are often inconsistent due to moderate or low correlations between mastitis indicators [46][37]. Selection to reduce the occurrence of cows with deep udders, especially low rear udders, open teats, teats that are set back, and also short and wide teats, can improve efforts to reduce the incidence of mastitis, along with better health control, therapeutic managing, and proper milking procedures [46][37]. There are factors associated with the distribution of clinical bovine mastitis between the hind and forequarters, in addition to risk factors related to certain aspects of lactation, udder conformation, and management practices [54,55,56][38][39][40]. Thus, programs to select sires whose progeny have the lowest SCCs should be carefully planned, taking into account the interpretation of SCCs as an immunological defense mechanism [11,27,57,58][11][22][41][42]. High SCCs decrease the likelihood of subsequent infections; however, there is a heritable variation in SCCs in dairy cattle. Therefore, using SCCs as a defense marker should be studied in greater depth [45][33]. Clinical mastitis in the hindquarters was found to be more common among primiparous cows, with a prevalence rate of 61.9% in affected cases [54][38].5.2. Genomic, Genotypic, and Phenotypic Udder Traits: Effects on Milk Production and Quality
Traditionally, dairy cattle have been selected for their ability to produce milk in quantity and quality [20][25]. The traditional approach to dairy cattle selection has changed, and secondary traits are now included in the selection indices, with less emphasis on milk production and more interest in milk quality and other non-productive traits [20,53,63][25][43][44]. Greater emphasis on non-productive traits is reflected in the industry’s desire to breed less productive but more functional dairy cattle [20][25]. The phenotypic and genomic traits related to milk yield, protein, and fat are included in the performance tests in young bulls [8,11][8][11]. Several linear-type traits have been added to the factor analysis. Heritability (h2) estimates for milk yield and milk quality-derived traits ranged from 0.125 to 0.219 [11,45][11][33]. Moreover, genetic estimates for milk yield and milk quality traits were negatively correlated with traits explaining udder conformation (−0.40) and rear muscularity [11]. The latter genetic traits also showed a negative correlation with udder volume (−0.28). Additionally, head typicality and rear leg traits were not correlated with milk yield and milk quality but were negatively correlated with meat-related traits such as rear muscularity (−0.32) [11]. The consequence of these results is that the use of the current selection index, which is mainly focused on milk production traits, may lead to a deterioration of all other traits. Therefore, more appropriate selection indices should consider an association between genetic and functional traits [11,23,37,57][11][26][41][45]. The selection and correlation of different traits for milk production based on the selection of AI bulls preferred for high transmission capacity is now possible [3,27,57][3][22][41]. The estimates regarding sire selection criteria may vary; however, the selection for milk production traits effectively increases milk production [8]. Nonetheless, in a research project evaluating the direct and correlated effects of single-trait selection on milk yield all selective breeding groups increased productivity, but also undesirable responses correlated with selection for milk production were detected [27][22]. Sometimes, bull sires from different genetic lines were selected by progeny testing based on the first lactation productivity and milk quality by associating traits for fat-corrected milk production, percentage of daughters discarded at the first lactation, and daughters’ udder conformation. However, one study showed no differences regarding milk yield among lines [57][41]. It has been observed by RNA sequencing that DDIT3, RPL23A, SESN2, and NR4A1 genes are significantly and differentially expressed between mammary glands of lactating Holstein cows with extremely high or low protein and fat percentages. Therefore, these four genes could affect milk production and composition traits [76][46]. In addition, the genes identified in another study, such as PGM1 and ARL4A, were related to milk production traits, lactose synthesis, glucose metabolism [77][47], and milk production or composition [78][48]. Linear evaluation plays a vital role in estimating the milk production of dairy cows; however, these evaluations, sometimes taken through subjective methods, can show variations [5,31][5][49]. On the other hand, objective methods for estimating milk productivity provide more accurate and reliable data but require more sophisticated technologies [5]. In one study, sire-derived type traits were analyzed in cows with low, medium, or high production performance during the first lactation [80][50]. The factors examined in the model included herd-year-season, age at calving, the month of calving, recording status interaction, change in herd size, and season.5.3. Genomic, Genotypic, and Phenotypic Udder Traits: Influence on True and Functional Longevity
Longevity-related traits have garnered growing interest among various milk-producing species, including sheep, cattle, goats, and other animals, as efforts to enhance the longevity of these animals continue to gain importance. Therefore, understanding the impact of genomic, genotypic, and phenotypic udder traits is crucial in determining dairy cows’ true and functional longevity. Functional longevity is defined as the number of days between first calving and culling, that is to say, the length of cows´ productive life [81][51]. Functional longevity is an economically important trait to increase the profitability of dairy management [82][52]. In different bovine dairy herds, the reasons for culling cows can be voluntary (mainly because of low productivity) or involuntary (mainly because of health and low fertility) [81,83][51][53]. The longevity-derived traits include (i) true longevity (all reasons for culling, including productivity) and (ii) functional longevity (all reasons for culling except productivity) [82][52]. Conformation traits and functional longevity have been related by survival analysis (Cox proportional hazards models) in first-lactation Holstein cows [81][51]. The dairy character had the strongest correlation between a composite trait and functional longevity, followed by the udder final score [81,84][51][54]. A more accurate methodology for improving animal longevity is necessary to decrease the involuntary culling rates rather than extending traits that influence the herd life [82][52]. Therefore, the proportional hazard model is helpful for assessing genetic fitness for the traits influencing herd life. However, the differences between estimates made with the proportional hazards models and those made with linear animal models for one or more traits are unclear. Productive traits, udder traits, and leg and hoof traits are genetically correlated with longevity; consequently, these traits are used to assess longevity indirectly [12,16][12][16]. The reliability of genetic fitness-related estimates for longevity is increased by combining direct and indirect estimates [82][52]. Therefore, these genetic correlations should be reviewed periodically in different dairy cattle production systems as they vary according to the year of birth [82][52]. In light of this, QTLs affecting economically relevant traits were studied for eight US Holstein cattle genetic lines [85][55]. A marker on chromosome 14 associated with differences in fat yield, fat percentage, and milk yield was observed in two genetic lines. Other markers located on chromosomes 16 and 20 were related to differences in udder depth and anterior udder attachment, respectively. A marker on chromosome 27 was associated with a difference in milkability index. These additional markers complete the quantitative trait locus mapping to identify QTLs affecting economically important traits in a selected commercial Holstein population [85][55]. The relationship between different conformation traits and functional longevity is essential in dairy cows and has been evaluated for years using survival analyses [80,81][50][51]. As mentioned, the highest correlations among descriptive traits were observed for longevity-udder attachment and longevity-udder depth [26,35,43,46,59][21][23][31][37][56]. However, functional longevity decreased with decreasing body condition in dairy cows [81][51]. Other relevant factors affecting longevity could be related to chronic stress in dairy cattle.5.4. Relationship between Genomic, Genotypic, and Phenotypic Udder Traits: Health, Production, and Longevity
The intricate interplay between genomic, genotypic, and phenotypic udder traits is fundamental to understanding the complex relationship between health, production, and longevity in dairy cattle. The dairy cattle sector is significantly impacted by the economic costs associated with the high incidence and prevalence of clinical and subclinical mastitis, including expenses related to treatment, production losses, and reduced animal welfare [59,93][56][57]. The large databases generated have allowed for assessing the incidence of this health problem and investigating the genetic background of clinical mastitis and its relationships with other udder-derived traits of interest for the dairy industry [59][56]. There is persistent controversy about low milk SCCs and susceptibility to mastitis [94][58]. However, high SCCs in milk may indicate inflammation or infection of the mammary gland [94][58]. Other studies involved molecular genome mapping results which provided information on quantitative trait loci (QTL) related to mastitis resistance and provided a better understanding of the genetic relevance of the traits [37][45]. Many countries have implemented selection programs based on a linear decrease in SCCs for increasing mastitis resistance [37][45]. Improving the selection accuracy for mastitis resistance includes advances in modeling, an optimal combination of mastitis-related traits, and associated udder predictors [37][45]. In addition, the definition of the overall breeding objective that includes udder-related conformational and functional traits and the inclusion of molecular-based information is now available from QTL [37,96][45][59]. These cutting-edge studies will lead to a better understanding of the genetic background of mastitis resistance and allow for more accurate selection, improving udder health, animal welfare, and profitability in the future modern dairy industry [59,97][56][60]. Within the domain of conformation traits, a study was conducted to assess the impact of udder morphological characteristics on milk production in Bos indicus cows [104,105][61][62]. First, the udder diameter and height, teat length and diameter, and milk production were measured, and finally, the study determined the values of udder morphological characteristics in local zebu cows. The results showed that the udder size was highly and positively correlated with milk production. These findings will be instrumental in genetic improvement programs for zebu cows [106][63]. Dairy cattle offer an attractive model for investigating the genes responsible for the substantial diversity observed both within and between mammalian species concerning their milk volume, protein, and fat composition, highlighting the potential significance of genomic traits. Many phenotypes for these traits and the complete genome sequence of the key founders of modern dairy cattle populations are available. Association tests were conducted on Holstein and Jersey cattle with exceptional phenotypes to identify variants within the target regions, while gene expression data were analyzed to pinpoint potential candidate genes such as BTRC, MGST1, SLC37A1, STAT5A, STAT5B, PAEP, VDR, CSF2RB, MUC1, NCF4, and GHDC associated with milk production [108][64]. In Bos taurus, 141 and five novel genes related to milk production and SCS have been identified, respectively. These novel genes were also found to be functionally related to heat tolerance (e.g., SLC45A2, IRAG1, and LOC101902172), longevity (e.g., SYT10 and LOC101903327), and fertility [109][65].References
- Carta, A.; Casu, S.; Salaris, S. Invited Review: Current State of Genetic Improvement in Dairy Sheep. J. Dairy Sci. 2009, 92, 5814–5833.
- Egger-Danner, C.; Cole, J.B.; Pryce, J.E.; Gengler, N.; Heringstad, B.; Bradley, A.; Stock, K.F. Invited Review: Overview of New Traits and Phenotyping Strategies in Dairy Cattle with a Focus on Functional Traits. Animal 2014, 9, 191–207.
- Chesnais, J.P.; Cooper, T.A.; Wiggans, G.R.; Sargolzaei, M.; Pryce, J.E.; Miglior, F. Using Genomics to Enhance Selection of Novel Traits in North American Dairy Cattle. J. Dairy Sci. 2016, 99, 2413–2427.
- Gutiérrez-Reinoso, M.A.; Aponte, P.M.; Cabezas, J.; Rodriguez-Alvarez, L.; Garcia-Herreros, M. Genomic Evaluation of Primiparous High-Producing Dairy Cows: Inbreeding Effects on Genotypic and Phenotypic Production-Reproductive Traits. Animals 2020, 10, 1704.
- Flower, F.C.; Weary, D.M. Gait Assessment in Dairy Cattle. Animal 2009, 3, 87–95.
- Jacobs, J.A.; Siegford, J.M. Lactating Dairy Cows Adapt Quickly to Being Milked by an Automatic Milking System. J. Dairy Sci. 2012, 95, 1575–1584.
- Cecchinato, A.; Macciotta, N.P.P.; Mele, M.; Tagliapietra, F.; Schiavon, S.; Bittante, G.; Pegolo, S. Genetic and Genomic Analyses of Latent Variables Related to the Milk Fatty Acid Profile, Milk Composition, and Udder Health in Dairy Cattle. J. Dairy Sci. 2019, 102, 5254–5265.
- Shanks, R.D.; Rooney, K.A.; Hutjens, M.F. Breeding Practices on Illinois Holstein Farms. J. Dairy Sci. 1983, 66, 1209–1217.
- Beard, J.K.; Musgrave, J.A.; Funston, R.N.; Travis Mulliniks, J. The Effect of Cow Udder Score on Cow/Calf Performance in the Nebraska Sandhills. Transl. Anim. Sci. 2019, 3, 14–19.
- Gutiérrez-Reinoso, M.A.; Aponte, P.M.; García-Herreros, M. A Review of Inbreeding Depression in Dairy Cattle: Current Status, Emerging Control Strategies, and Future Prospects. J. Dairy Res. 2022, 89, 3–12.
- Mancin, E.; Sartori, C.; Guzzo, N.; Tuliozi, B.; Mantovani, R. Selection Response Due to Different Combination of Antagonistic Milk, Beef, and Morphological Traits in the Alpine Grey Cattle Breed. Animals 2021, 11, 1340.
- Castañeda-Bustos, V.J.; Montaldo, H.H.; Valencia-Posadas, M.; Shepard, L.; Pérez-Elizalde, S.; Hernández-Mendo, O.; Torres-Hernández, G. Linear and Nonlinear Genetic Relationships between Type Traits and Productive Life in US Dairy Goats. J. Dairy Sci. 2017, 100, 1232–1245.
- Gutierrez-Reinoso, M.A.; Aponte, P.M.; Garcia-Herreros, M. Genomic Analysis, Progress and Future Perspectives in Dairy Cattle Selection: A Review. Animals 2021, 11, 599.
- Hazel, A.R.; Heins, B.J.; Hansen, L.B. Health Treatment Cost, Stillbirth, Survival, and Conformation of Viking Red-, Montbéliarde-, and Holstein-Sired Crossbred Cows Compared with Pure Holstein Cows during Their First 3 Lactations. J. Dairy Sci. 2020, 103, 10917–10939.
- Heimes, A.; Brodhagen, J.; Weikard, R.; Hammon, H.M.; Meyerholz, M.M.; Petzl, W.; Zerbe, H.; Engelmann, S.; Schmicke, M.; Hoedemaker, M.; et al. Characterization of Functional Traits with Focus on Udder Health in Heifers with Divergent Paternally Inherited Haplotypes on BTA18. BMC Vet. Res. 2019, 15, 241.
- Hansen, L.B.; Cole, J.B.; Marx, G.D.; Seykora, A.J. Productive Life and Reasons for Disposal of Holstein Cows Selected for Large versus Small Body Size. J. Dairy Sci. 1999, 82, 795–801.
- Dadati, E.; Kennedy, B.W.; Burnside, E.B. Relationships between Conformation and Calving Interval in Holstein Cows. J. Dairy Sci. 1986, 69, 3112–3119.
- Strandberg, E.; Shook, G.E. Genetic and Economic Responses to Breeding Programs That Consider Mastitis. J. Dairy Sci. 1989, 72, 2136–2142.
- Hazel, A.R.; Heins, B.J.; Hansen, L.B. Fertility, Survival, and Conformation of Montbéliarde × Holstein and Viking Red × Holstein Crossbred Cows Compared with Pure Holstein Cows during First Lactation in 8 Commercial Dairy Herds. J. Dairy Sci. 2017, 100, 9447–9458.
- Wiggans, G.R.; Hubbard, S.M. Genetic Evaluation of Yield and Type Traits of Dairy Goats in the United States. J. Dairy Sci. 2001, 84, 69–73.
- Casu, S.; Pernazza, I.; Carta, A. Feasibility of a Linear Scoring Method of Udder Morphology for the Selection Scheme of Sardinian Sheep. J. Dairy Sci. 2006, 89, 2200–2209.
- Kelm, S.C.; Freeman, A.E.; Brundage, A.L.; Pearson, R.E.; Martin, T.G.; McGilliard, L.D.; Hansen, L.B.; Young, C.W.; Voelker, H.H.; Shook, G.E.; et al. Direct and Correlated Responses to Selection for Milk Yield: Results and Conclusions of Regional Project NC-2, “Improvement of Dairy Cattle through Breeding, with Emphasis on Selection”. J. Dairy Sci. 2000, 83, 2721–2732.
- Miglior, F.; Koeck, A.; Jamrozik, J.; Schenkel, F.; Kelton, D.; Kistemaker, G.; Van Doormaal, B. Index for Mastitis Resistance and Use of BHBA for Evaluation of Health Traits in Canadian Holsteins. Interbull Bull. 2014, 48, 73–78.
- Parker Gaddis, K.L.; Cole, J.B.; Clay, J.S.; Maltecca, C. Genomic Selection for Producer-Recorded Health Event Data in US Dairy Cattle. J. Dairy Sci. 2014, 97, 3190–3199.
- Lucy, M.C. Non-Lactational Traits of Importance in Dairy Cows and Applications for Emerging Biotechnologies. N. Z. Vet. J. 2005, 53, 406–415.
- Parker Gaddis, K.L.; VanRaden, P.M.; Cole, J.B.; Norman, H.D.; Nicolazzi, E.; Dürr, J.W. Symposium Review: Development, Implementation, and Perspectives of Health Evaluations in the United States. J. Dairy Sci. 2020, 103, 5354–5365.
- Pausch, H.; Emmerling, R.; Schwarzenbacher, H.; Fries, R. A Multi-Trait Meta-Analysis with Imputed Sequence Variants Reveals Twelve QTL for Mammary Gland Morphology in Fleckvieh Cattle. Genet. Sel. Evol. 2016, 48, 14.
- Nazar, M.; Abdalla, I.M.; Chen, Z.; Ullah, N.; Liang, Y.; Chu, S.; Xu, T.; Mao, Y.; Yang, Z.; Lu, X. Genome-Wide Association Study for Udder Conformation Traits in Chinese Holstein Cattle. Animals 2022, 12, 2542.
- Zwald, N.R.; Weigel, K.A.; Chang, Y.M.; Welper, R.D.; Clay, J.S. Genetic Selection for Health Traits Using Producer-Recorded Data. II. Genetic Correlations, Disease Probabilities, and Relationships with Existing Traits. J. Dairy Sci. 2004, 87, 4295–4302.
- Heringstad, B.; Klemetsdal, G.; Steine, T. Selection Responses for Disease Resistance in Two Selection Experiments with Norwegian Red Cows. J. Dairy Sci. 2007, 90, 2419–2426.
- Jamrozik, J.; Koeck, A.; Miglior, F.; Kistemaker, G.; Schenkel, F.; Kelton, D.; Doormaal, B. Van Genetic and Genomic Evaluation of Mastitis Resistance in Canada. Interbull Bull. 2013, 47.
- Koeck, A.; Miglior, F.; Kelton, D.F.; Schenkel, F.S. Alternative Somatic Cell Count Traits to Improve Mastitis Resistance in Canadian Holsteins. J. Dairy Sci. 2012, 95, 432–439.
- Miller, R.H. Traits for Sire Selection Related to Udder Health and Management. J. Dairy Sci. 1984, 67, 459–471.
- Wagner, P.; Yin, T.; Brügemann, K.; Engel, P.; Weimann, C.; Schlez, K.; König, S. Genome-Wide Associations for Microscopic Differential Somatic Cell Count and Specific Mastitis Pathogens in Holstein Cows in Compost-Bedded Pack and Cubicle Farming Systems. Animals 2021, 11, 1839.
- Yang, C.; Zhao, X.; Cui, N.; Liang, Y. Cadherins Associate with Distinct Stem Cell-Related Transcription Factors to Coordinate the Maintenance of Stemness in Triple-Negative Breast Cancer. Stem Cells Int. 2017, 2017, 5091541.
- Li, Y.; Han, B.; Liu, L.; Zhao, F.; Liang, W.; Jiang, J.; Yang, Y.; Ma, Z.; Sun, D. Genetic Association of DDIT3, RPL23A, SESN2 and NR4A1 Genes with Milk Yield and Composition in Dairy Cattle. Anim. Genet. 2019, 50, 123–135.
- Thomas, C.L.; Vinson, W.E.; Pearson, R.E.; Dickinson, F.N.; Johnson, L.P. Relationships between Linear Type Scores, Objective Type Measures, and Indicators of Mastitis. J. Dairy Sci. 1984, 67, 1281–1292.
- Lancelot, R.; Paye, B.; Lescourret, F. Factors Affecting the Distribution of Clinical Mastitis among Udder Quarters in French Dairy Cows. Vet. Res. 1997, 28, 45–53.
- Bhutto, A.L.; Murray, R.D.; Woldehiwet, Z. Udder Shape and Teat-End Lesions as Potential Risk Factors for High Somatic Cell Counts and Intra-Mammary Infections in Dairy Cows. Vet. J. 2010, 183, 63–67.
- Guarín, J.F.; Baumberger, C.; Ruegg, P.L. Anatomical Characteristics of Teats and Premilking Bacterial Counts of Teat Skin Swabs of Primiparous Cows Exposed to Different Types of Bedding. J. Dairy Sci. 2017, 100, 1436–1444.
- Miller, R.H.; Pearson, R.E.; Rothschild, M.F.; Fulton, L.A. Comparison of Single and Multiple-Trait Selected Sires. Response in Mastitis Traits. J. Dairy Sci. 1981, 64, 832–837.
- Boettcher, P.J.; Dekkers, J.C.M.; Kolstad, B.W. Development of an Udder Health Index for Sire Selection Based on Somatic Cell Score, Udder Conformation, and Milking Speed. J. Dairy Sci. 1998, 81, 1157–1168.
- Van Eetvelde, M.; Verdru, K.; de Jong, G.; van Pelt, M.L.; Meesters, M.; Opsomer, G. Researching 100 t Cows: An Innovative Approach to Identify Intrinsic Cows Factors Associated with a High Lifetime Milk Production. Prev. Vet. Med. 2021, 193, 105392.
- Bharti, P.; Bhakat, C.; Pankaj, P.K.; Bhat, S.A.; Arul Prakash, M.; Thul, M.R.; Puhle Japheth, K. Relationship of Udder and Teat Conformation with Intra-Mammary Infection in Crossbred Cows under Hot-Humid Climate. Vet. World 2015, 8, 898–901.
- Rupp, R.; Boichard, D. Genetics of Resistance to Mastitis in Dairy Cattle. Vet. Res. 2003, 34, 671–688.
- Li, T.; Gao, J.; Zhao, X.; Ma, Y. Digital Gene Expression Analyses of Mammary Glands from Meat Ewes Naturally Infected with Clinical Mastitis. R. Soc. Open Sci. 2019, 6, 181604.
- Lemay, D.G.; Hovey, R.C.; Hartono, S.R.; Hinde, K.; Smilowitz, J.T.; Ventimiglia, F.; Schmidt, K.A.; Lee, J.W.S.; Islas-Trejo, A.; Silva, P.I.; et al. Sequencing the Transcriptome of Milk Production: Milk Trumps Mammary Tissue. BMC Genom. 2013, 14, 872.
- Raschia, M.A.; Nani, J.P.; Maizon, D.O.; Beribe, M.J.; Amadio, A.F.; Poli, M.A. Single Nucleotide Polymorphisms in Candidate Genes Associated with Milk Yield in Argentinean Holstein and Holstein x Jersey Cows. J. Anim. Sci. Technol. 2018, 60, 31.
- Veerkamp, R.F.; Gerritsen, C.L.M.; Koenen, E.P.C.; Hamoen, A.; De Jong, G. Evaluation of Classifiers That Score Linear Type Traits and Body Condition Score Using Common Sires. J. Dairy Sci. 2002, 85, 976–983.
- Boettcher, P.J.; Jairath, L.K.; Koots, K.R.; Dekkers, J.C.M. Effects of Interactions between Type and Milk Production on Survival Traits of Canadian Holsteins. J. Dairy Sci. 1997, 80, 2984–2995.
- Zavadilová, L.; Němcová, E.; Štípková, M. Effect of Type Traits on Functional Longevity of Czech Holstein Cows Estimated from a Cox Proportional Hazards Model. J. Dairy Sci. 2011, 94, 4090–4099.
- Sasaki, O. Estimation of Genetic Parameters for Longevity Traits in Dairy Cattle: A Review with Focus on the Characteristics of Analytical Models. Anim. Sci. J. 2013, 84, 449–460.
- Forabosco, F.; Groen, A.F.; Bozzi, R.; Van Arendonk, J.A.M.; Filippini, F.; Boettcher, P.; Bijma, P. Phenotypic Relationships between Longevity, Type Traits, and Production in Chianina Beef Cattle. J. Anim. Sci. 2004, 82, 1572–1580.
- Hocking, P.M.; McAllister, A.J.; Wolynetz, M.S.; Batra, T.R.; Lee, A.J.; Lin, C.Y.; Roy, G.L.; Vesely, J.A.; Wauthy, J.M.; Winter, K.A. Factors Affecting Length of Herdlife in Purebred and Crossbred Dairy Cattle. J. Dairy Sci. 1988, 71, 1011–1024.
- Ashwell, M.S.; Van Tassell, C.P.; Sonstegard, T.S. A Genome Scan to Identify Quantitative Trait Loci Affecting Economically Important Traits in a US Holstein Population. J. Dairy Sci. 2001, 84, 2535–2542.
- Martin, P.; Barkema, H.W.; Brito, L.F.; Narayana, S.G.; Miglior, F. Symposium Review: Novel Strategies to Genetically Improve Mastitis Resistance in Dairy Cattle. J. Dairy Sci. 2018, 101, 2724–2736.
- Malchiodi, F.; Jamrozik, J.; Christen, A.M.; Fleming, A.; Kistemaker, G.J.; Richardson, C.; Daniel, V.; Kelton, D.F.; Schenkel, F.S.; Miglior, F. Symposium Review: Multiple-Trait Single-Step Genomic Evaluation for Hoof Health. J. Dairy Sci. 2020, 103, 5346–5353.
- Rainard, P.; Foucras, G.; Boichard, D.; Rupp, R. Invited Review: Low Milk Somatic Cell Count and Susceptibility to Mastitis. J. Dairy Sci. 2018, 101, 6703–6714.
- Cole, J.B.; Wiggans, G.R.; Ma, L.; Sonstegard, T.S.; Lawlor, T.J.; Crooker, B.A.; Van Tassell, C.P.; Yang, J.; Wang, S.; Matukumalli, L.K.; et al. Genome-Wide Association Analysis of Thirty One Production, Health, Reproduction and Body Conformation Traits in Contemporary U.S. Holstein Cows. BMC Genom. 2011, 12, 408.
- Sahana, G.; Lund, M.S.; Andersson-Eklund, L.; Hastings, N.; Fernandez, A.; Iso-Touru, T.; Thomsen, B.; Viitala, S.; Sørensen, P.; Williams, J.L.; et al. Fine-Mapping QTL for Mastitis Resistance on BTA9 in Three Nordic Red Cattle Breeds. Anim. Genet. 2008, 39, 354–362.
- Riley, D.G.; Sanders, J.O.; Knutson, R.E.; Lunt, D.K. Comparison of F1 Bos Indicus x Hereford Cows in Central Texas: II. Udder, Mouth, Longevity, and Lifetime Productivity. J. Anim. Sci. 2001, 79, 1439–1449.
- Tolleson, M.W.; Gill, C.A.; Herring, A.D.; Riggs, P.K.; Sawyer, J.E.; Sanders, J.O.; Riley, D.G. Association of Udder Traits with Single Nucleotide Polymorphisms in Crossbred Bos Indicus- Bos Taurus Cows. J. Anim. Sci. 2017, 95, 2399–2407.
- Mingoas, K.J.P.; Awah-Ndukum, J.; Dakyang, H.; Zoli, P.A. Effects of Body Conformation and Udder Morphology on Milk Yield of Zebu Cows in North Region of Cameroon. Vet. World 2017, 10, 901.
- Raven, L.A.; Cocks, B.G.; Kemper, K.E.; Chamberlain, A.J.; Vander Jagt, C.J.; Goddard, M.E.; Hayes, B.J. Targeted Imputation of Sequence Variants and Gene Expression Profiling Identifies Twelve Candidate Genes Associated with Lactation Volume, Composition and Calving Interval in Dairy Cattle. Mamm. Genome 2016, 27, 81–97.
- Buaban, S.; Lengnudum, K.; Boonkum, W.; Phakdeedindan, P. Genome-Wide Association Study on Milk Production and Somatic Cell Score for Thai Dairy Cattle Using Weighted Single-Step Approach with Random Regression Test-Day Model. J. Dairy Sci. 2022, 105, 468–494.
More
