1000/1000
Hot
Most Recent

Ankyrin repeat (AR) domains are considered the most abundant repeat motif found in eukaryotic proteins. AR domains are predominantly known to mediate specific protein–protein interactions (PPIs) without necessarily recognizing specific primary sequences, nor requiring strict conformity within its own primary sequence.
Many proteins have evolved through gene duplication and recombination events to produce repetitive motifs in their primary sequences. These non-overlapping repeat regions, commonly referred to as tandem repeats, provide a high amount of sequence conservation that are generally thought to prevent deleterious residue substitutions that cause alterations to the global fold of the domain. Another tangible benefit of tandem repeats is that they can provide multiple binding sites for various intracellular proteins that can play an important role in protein structural integrity. Tandemly occurring repeats within the primary sequence display specific characteristics in their three-dimensional structure, forming an integrated assembly which allows for tandem repeat domain characterization [1]. The classification of tandem repeats is based upon the formation and specific localization of secondary structural elements such as α-helical bundles, β-hairpin loops, β-sheets and propellers, and horse shoe shapes [2][3][4]. Although identification of tandem repeats relies heavily on these characteristics, the size and diversity can greatly vary. Intriguingly, tandem repeat domains occur in 14% of all eukaryotic proteins and are three times as likely to occur in eukaryotic than prokaryotic proteins [5]. Four abundant classes of repeat domains exist that vary in their elongated structures that facilitate protein–protein interactions that include ankyrin repeats (AR), leucine rich repeats (LRR), armadillo repeats (ARM), and tetratricopeptide repeats (TPR) [6]. Each repeat domain acts as a scaffold for substrate proteins and their selectivity is dependent on the subtle differences in primary sequence within each repeated domain. It is unclear which residues are critical for the scaffolding structure and which are required for the overall function of the domain [7]. While protein–protein interactions are regulated through specific amino acid sequences or structural characteristics, variations within the surface exposed residues in AR domains enable specific protein binding [6].
AR domains were first discovered as a repeating sequence in Saccharomyces cerevisiae cell cycle regulators Swi6, cell division control protein 10 (Cdc10) and Notch in Drosophila melanogaster [8]. This ~33-amino acid long repeat subsequently was named after the cytoskeletal protein ANKYRIN, a 206 kDa protein that contains 24 tandem AR repeats [9]. Since its initial discovery, AR domains have been observed to be present in many eukaryotic proteins, making this domain potentially the most abundant repeat domain in the eukaryotic proteome [7]. To date, there are over 367,000 predicted AR domains found within 68,471 nonredundant proteins annotated in the SMART database [10][11]. With such prevalence of AR-containing proteins in eukaryotes coupled with AR domains acting as scaffolds to facilitate protein–protein interactions in the cell, it is speculated that the AR domain originated through evolutionary pressure events to provide the necessary function of facilitating the variety of signaling pathways eukaryotic organisms use to regulate cellular homeostasis [6].
Comparing AR-containing proteins has proven to be difficult as each protein has acquired various characteristics through multiple evolutionary events. This is largely due to conservation within AR domains relying on various residue types rather than requiring highly conserved residues at specific sites. Given that specific residue types influence the protein’s secondary structure coupled with AR domains not relying on specific residue conservation, it suggests the AR domain is defined primarily on its 3D structural fold rather than its functional support for AR-containing proteins. For example, highly conserved regions of AR domains exist between each repeat, whereas variation of hydrophobic residues can occur while maintaining the structural integrity of the domain [12]. Conserved motifs that influence α-helical and β-loop folds have recently been identified in AR domains has allowed for better AR domain identification and prediction from the primary sequence of a protein.
In comparison to naturally occurring AR domains, there is little deviation from its typical helix–loop–helix–β–hairpin/loop structure, which is supported through conservation of residue type. Inter-domain interactions tend to be short distances, which aids in the linear solenoid packing of the AR domain fold rather than a typical globular shape. Both hydrophobic interactions through non-polar regions of the inter and intra α-helices as well as hydrogen bonding through polar residues found near the N-terminus define and stabilize the AR domain’s structural integrity [6][13] (Figure 1A).
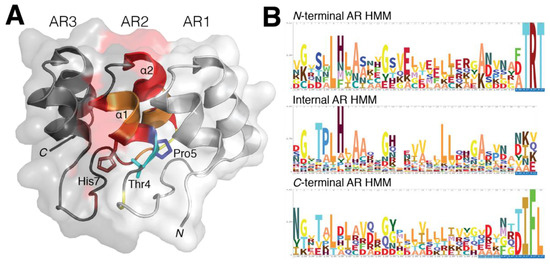
Figure 1. Structural conservation within ankyrin repeat domains. (A) Residues Thr4, Pro5 and His7 play a critical role in providing both the 90° and L-shape formation through hydrogen bonding. The elongated solenoid shape of an AR domain is predominantly dictated by these three residues while hydrophobic interactions in the core of the AR are required to stabilize the domain’s 3D fold. (B) The hidden Markov model (HMM) profile of the AR-containing proteins for N-terminal, internal and C-terminal repeats were analyzed to highlight the occurrence of residues in identified AR domain families. The classic G-X-TPLHLA motif was readily identified to have a strong probability of occurring in these AR-containing proteins, whereas in both N- and C-terminal repeats were observed to contain only portions of this motif. While conservation within AR domains is more prevalent within the internal and C-terminal repeats, the N-terminal AR still retains similar AR domain characteristics.
AR domain-containing proteins are ubiquitous and play essential roles in numerous biological processes that can also influence the onset of various diseases and disorders. Many AR domain-containing proteins have been identified in numerous occasions to support the ubiquitin signaling pathway and deubiquitylation through sequence alignment. Being essential scaffolds to mediate protein–protein interactions, it is intriguing that this domain is not functionally driven but rather dependent on the structural characteristics of the AR domain. Understanding how these AR-domain containing proteins and their role in ubiquitin signaling on the molecular level is a major challenge in our present understanding of their function(s) and activity. Expanded biochemical and biophysical examinations on the novel mechanisms used for AR-domain protein recruitment are needed. By experimentally identifying and validating potential substrates and interactors for these AR-domain proteins, we will be able to improve our knowledge of how these proteins work and we will be able to better assess if these proteins could serve as novel drug targets or as biomarkers for disease.