1000/1000
Hot
Most Recent

As the direct regulatory role of p53 and some of its isoform proteins are becoming established in modulating gene expression in cancer research, another aspect of this mode of gene regulation that has captured significant interest over the years is the mechanistic interplay between p53 and micro-RNA transcriptional regulation. The input of this into modulating gene expression for some of the cathepsin family members has been viewed as carrying noticeable importance based on their biological effects during normal cellular homeostasis and cancer progression.
The tumor suppressor gene TP53 is mutated at a high frequency in a whole range of malignant diseases and has therefore been intensely researched for many years [1]. As is to be expected, the number of molecular networks that it has been shown to fundamentally regulate have also grown with great diversity and include aspects of DNA repair [2], cell senescence [3], angiogenesis [4], apoptosis [5][6] and cell cycle regulation [7]. While the main role of p53 in most of these processes are through its being able to directly regulate gene expression upon DNA binding, it can also mediate this through interacting with other transcription factors and regulators [8]. In some of its genetically mutated forms (mut-p53), p53 can take on the properties of a protein that is oncogenic, while some mutated derivatives can simply be inactive at the genetic or protein level [9]. Similarly, one key contributing factor originates from p53 being expressed as isoform proteins arising from the use of alternative promoters, translation initiation sites and mRNA splicing sites and which can act individually or in concert in modulating gene expression (Figure 1) [10][11].
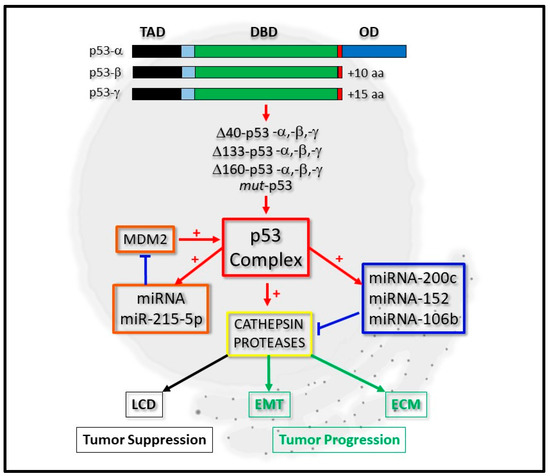
Figure 1. Integrative regulation of cathepsin proteases by p53 and micro-RNA expression. P53-alpha (p53-α) can be expressed as p53-beta (p53-β) or p53-gamma (p53-γ) isoform proteins, which lack the oligomerization domain (OD). Derivatives of these, which lack the complete Transactivation domain (TAD), but retain the DNA-binding domain (DBD), can also be expressed as Δ40-p53, Δ133-p53, Δ160-p53 or mut-p53 forms. The p53 complex can be regulated by micro-RNA (miRNA) expression through a positive feedback loop by positively regulating miRNA-215-5p, which negatively regulates MDM2 protein levels (orange boxes). It can transcriptionally regulate cathepsin protease expression directly or indirectly through directly regulating the expression of miRNA-200c, for example (blue box). Cathepsin protease expression (yellow box) contributes to lysosomal-mediated cell death (LCD) as a tumor suppressor (black boxes) or cell differentiation through Epithelial-Mesenchymal Transition (EMT) and the Extracellular Matrix (ECM) during tumor progression (green boxes).
From a regulatory perspective, p53 protein levels are kept to a minimum, through its polyubiquitination and destabilization by MDM2 and the proteasomal degradation pathway [9][10][11]. However, upon treating mammalian cells with oxidative stress or cytotoxic agents, nuclear p53 can become stabilized and modulate gene expression of proteins central to mediating cell arrest, DNA repair or apoptosis [12][13][14][15]. Additionally, post-translational modifications can also regulate p53 activity that mechanistically contribute to its cytoplasmic translocation, and where it can mediate mitochondria- or lysosomal-mediated cell death [16][17].
With over 14000 micro-RNAs annotated from the human genome that can regulate as much as 30% of all mRNAs expressed intracellularly, it is interesting to note that over 46% micro-RNA promoters have been reported to contain putative p53 binding sites [12][13]. While this highlights a potential direct link between p53 protein activation and micro-RNA expression, another important and direct role for the p53 protein in miRNA processing has also emerged. Here, p53 (or transcriptionally inactive p53) was revealed to be a central regulator of micro-RNA processing, through its ability to modulate the maturation of the micro-RNAs and their accessibility to mature mRNA messengers through its association with the protein Drosha [14] and the RISC complex (reviewed in [15][16]). Of importance is the ubiquitin ligase MDM2, which is under micro-RNA-mediated control as seen through the inhibitory actions of miRNA-192, miRNA-194, miRNA-215, miRNA-143, miRNA-145, and miRNA-605 expression [17]. For example, loss of miRNA-215-5p expression can enhance expression of MDM2, which results in diminished p53 protein levels [18]. As reported therein, p53 also positively regulated miRNA-215-5p expression, highlighting the existence of a p53 positive feedback loop [18]. Similarly, another good example of p53 regulation, through a miRNA acting on upstream activators of p53, occurs through miRNA-34, which acts by down-regulating the expression of the SIRT1 and HDAC intermediates that negatively-regulate p53 through its deacetylation (reviewed in [19]). While the actions of such micro-RNAs may give rise to enhanced levels of active p53 protein at the transcription and translation levels indirectly, p53 transcripts can also be directly targeted by miRNA-25 and miRNA-125b expression (reviewed in [16]).
The cathepsin proteases are a family of proteins that are developing greater importance due to them being intimately linked to tumor progression [20][21] and suppression [22]. During cancer progression, not only do they modulate the extracellular matrix and permit the dispersal of tumor cells following tumor growth, some of them also modulate the trans-differentiation of cells through the process of Epithelial Mesenchymal Transition (EMT) [21][23]. Simultaneously, the transcriptional regulation of cathepsins by p53 is also an area of research that is gaining much attention [22][24], particularly as lysosomes become more prone to lysis by lysosomorphic and cytotoxic agents upon cathepsin over-expression [25][26] and through p53 directly modulating lysosomal-mediated cell death [27][28], (Figure 1).
Consequently, the scientific interests revolving around the regulatory axis shared by all forms of p53, micro-RNAs and the cathepsins have captured the attention of many basic researchers, with a view to defining their co-regulatory relationships in greater depth (Figure 1).
The p53 protein was first described over 30 years ago and its biological significance since then has had a significant amount of input into many of the p53-related paradigms that have been developed in many aspects of cancer cell biology. During this time, the TP53 gene has also revealed itself to encode a number of important p53 isoforms proteins [10][11], which have set many precedents while laying a number of very strong foundations for the characterization of the subsequently discovered p53 somatic mutations with relative ease [9]. For simplicity, the p53 isoforms can be categorized into two groups (Figure 1). The first group contains the p53-α, p53-β and p53-γ forms (which respectively encode WT-p53 (wild-type p53) and isoforms lacking the carboxyl-terminal Oligomerization Domain (OD), which is replaced with 10–15 amino acid extensions formed through alternative splicing of the mRNA (Figure 1 and Table 1). While these are driven transcriptionally from the promoter upstream of the first exon [29], Δ40-p53 isoform derivatives can also arise through the alternative splicing of the p53 transcript and the use of the initiator AUG at codon 40 [30]. Additional p53 protein derivatives (lacking part of its amino-terminal) can also arise from transcripts being driven from a second promoter located between intron 1 and exon 5, giving rise to ΔN-terminal p53 isoforms which have a 133 and 160 amino acid deletion at the amino-terminal [30][31][32]. Broadly, the p53 derivatives lacking the amino termini can be categorized into the second group (Figure 1 and Table 1).
Table 1. p53 isoform proteins.
|
p53 Isoform |
Amino Acids |
Protein (kD) |
Reference |
|
p53-α |
1-393 |
53 |
[33] |
|
p53-β |
1-331+10 |
47 |
[10] |
|
p53-γ |
1-331+15 |
48 |
[29] |
|
Δ40-p53-α |
40-393 |
47 |
|
|
Δ40-p53-β |
40-331+10 |
42 |
[29] |
|
Δ40-p53-γ |
40-331+15 |
42 |
[29] |
|
Δ133-p53-α |
133-393 |
35 |
[29] |
|
Δ133-p53-β |
133-331+10 |
29 |
[29] |
|
Δ133-p53-γ |
133-331+15 |
29 |
[29] |
|
Δ160-p53-α |
161-393 |
31 |
[32] |
|
Δ160-p53-β |
161-331+10 |
26 |
|
|
Δ160-p53-γ |
161-331+15 |
26 |
Biologically, all of the p53 isoforms exhibit diverse degrees of dominant-inhibitory effects for trans-activating gene expression through their abilities to form homo-tetramers or hetero-tetramers with WT-p53 [30][31][34]. This is based upon some of the isoforms lacking the OD, the full trans-activating domain (TAD) and showing varying degrees of protein stability and transcriptional activity based on the presence or absence of key phosphorylation sites, such as Ser-46 [35][36][37][38] and the carboxyl-terminal MDM2-specific ubiquitination sites [34][39]. Importantly, their biochemical characterization has indeed helped in offering an insight into how the p53 proteins arising from somatic mutations within the TP53 gene may differ biochemically in comparison to WT-p53 (or p53-α). Such mutations can be broadly described as a gain of function (GOF) or a loss of function (LOF) and the most commonest of them are the R175, G245, R248, R249, R273 and R282 mutants (collectively known as mut-p53) and which make up around 30% of all mutations found within the TP53 gene [40][41][42].
More specifically, the characterization of such p53 mutants has offered some excellent mechanistic insights into how certain micro-RNAs are regulated transcriptionally, especially in the context of cancer progression. For example, as far back as 2011, Chang et al. (2011) reported that miRNA-200c expression could be down-regulated upon the expression of a number of mut-p53 derivatives in 106 patient samples and MCF12A BC cells, which correlated significantly with tumor grade [43]. More recently, the expression of mut-p53 has also been linked to decreased miRNA-200c expression in human osteosarcoma cells by Tamura et al. (2015, [44]) and Alam et al. (2017, [45]) who identified the R280K mut-p53 protein as being responsible for this [45]. Here, increased expression levels of Moesin in MCF7 1001 BC cells were reported, as a significant contributing factor to carcinogenesis.
Collectively, the existence of such a high number of p53 isoform proteins can potentially offer a number of alternative mechanisms for how the TP53 gene can exert its biological effects. Consequently, their importance in being able to regulate tumor suppressive miRNA expression, either exclusively or with WT-p53, is being viewed as mechanistically significant during tumor initiation or progression.
The family of cathepsin proteases is composed of aspartate proteases (D, E), serine proteases (A, G) and the cysteine proteases (B, C, F, H, K, L, O, S, V, Z/X, W) [20]. Collectively, they are expressed as inactive zymogens, which have the capability to become auto-activated or trans-activated as they traffic from the endosome to reside within the lysosome, but can also be found in the nucleus [46]. Some of them are upregulated in expression, especially during cancer progression and can be secreted into the Extracellular Matrix (ECM) where they can modulate ECM components and contribute to malignancy [47][48]. Nevertheless, normally they are localized within the lysosome, from where they can leak into the cytoplasm and activate intermediates from the intrinsic apoptotic pathway as in the case of BID cleavage, causing the activation of apoptosis [49].
More recently, cathepsins L and D have been seen to reside in the nucleus where they can cleave the Histone H3 protein [50][51], CUX1 [52][53][54], TRPS1 [23] and enhance proliferation, induce EMT and increase the motility of cells. Consequently, a strong interest in how cathepsin expression is regulated has developed with the transcriptional regulation of cathepsins D and L having been linked to p53. Here, cathepsin D was expressed in a p53-dependent manner in U1752, Pa1 and ML1 leukemia cell manner and p53 was reported to bind to two p53 consensus sequences within the cathepsin D promoter [22]. Similarly, p53 could bind the promoter region of cathepsin L and the expression of which could also be driven by mut-p53 expression in glioblastoma cells [24]. Being mindful of these observations, there are justifiable reasons for why the scope of research here needs to be broadened in order to ascertain how cathepsins may be regulated in the absence and presence of p53 (or its isoforms and mut-p53 derivatives), and whether such events can still permit the cathepsins to drive tumor progression.
Generally speaking, developing interests have revolved around how cathepsin genes may be regulated by specific micro-RNAs, and of importance here is how these may be linked to what is commonly known about p53 and cathepsin protease regulation. In the instance of cathepsin proteases, this area of research appears to be relatively undeveloped, and being mindful of there being around 15 cathepsin proteases (with the majority of them being linked to cancer development or progression [20]), reportedly only a few of them appear to be regulated by micro-RNAs that have a direct or indirect connection with p53. Moreover, the regulation of cathepsins in the context of p53 isoforms or mutant-derivatives thereof appear to be even less explored and is an important consideration in light of how quickly this area of p53 biology is expanding.
Consequently, in highlighting the nature of these developing integrative regulatory networks, the next section is devoted to reviewing, a) which micro-RNAs are regulated by (or regulate) p53, and b) how these micro-RNAs regulate cathepsin protease family members in the context of cancer, with a view to broadening our understanding of the regulatory interplay between p53 and cathepsin transcription. Broadly speaking, miRNA-200c, miRNA-152 and miRNA-106b appear to be the most characterized in this context, with others such as miRNA-29a (cathepsin K, [55]), miRNA-30 (cathepsin D, [56]), miRNA-25-3p (cathepsin K, [57]), miRNA-140 (cathepsin B, [58]), miRNA-483-5p (cathepsin K, [59]) and miRNA-506-3p (cathepsin K, [60]) being characterized to a lesser extent (Table 2).
Table 2. The developing networks between micro-RNA, cathepsin proteases and p53 expression.
|
Micro-RNA |
Cathepsin |
p53 isoform |
Cell Type |
Reference |
|
miRNA-200c |
L |
WT-p53-α |
A549 Lung |
|
|
miRNA-152 |
L |
WT-p53-α |
Gastrointestinal |
|
|
miRNA-106b |
A |
WT-p53-α |
Colorectal |
|
|
miRNA-140 |
B |
- |
Glioblastoma |
[58] |
|
miRNA-30 |
D |
- |
Macrophage |
[56] |
|
miRNA-25-3p |
K |
- |
Osteoblast |
[57] |
|
miRNA-483-5p |
K |
- |
PBMC |
[59] |
|
miRNA-506-3p |
K |
- |
Macrophage |
[60] |
|
miRNA-29a |
K |
- |
Osteoblast |
[55] |
The expression of micro-RNAs connected with cathepsin gene expression are highlighted in conjunction with specific p53 isoforms and cell types they have been collectively characterized in (WT-p53, wild-type p53; PBMC, Peripheral Blood Mononuclear Cells; -, unknown).
Based on the above, there are clear regulatory relationships that are emerging between p53, cathepsin and micro-RNA expression. While the focus of this article has so far been originated from defining the molecular roles that p53 and cathepsins share in disease progression, for completeness we would like to extend the importance of the above miRNAs within a clinical context. This has great significance through the common biological traits their downstream target gene products share with some of the cathepsin proteases, and therefore it is worth focusing on this through highlighting alternative transcripts (or proteins) that are targeted by these miRNAs. In addition to this, we would like to review the recent progress on how these micro-RNAs are being utilized in diagnostic and prognostic assays. For simplicity and consistency, we will keep the focus and the context as close to lung cancer (miRNA-200c), gastric cancer (miRNA-152) and colorectal cancer (miRNA-106b), as possible.
As far back as 2013, the importance of miRNA-200c in the regulation of disease progression has positively been gaining greater momentum. For example, the loss of miRNA-200c within the lungs [66] was seen to correlate with NSCLC cells showing an invasive and chemo-resistant phenotype [67], while positive expression of it could sensitize cells to chemotherapeutic [68] and radiotherapeutic [69] agents. As reported by Cortez et al. (2014), such findings could be extended and they reported the expression of miRNA-200c enhanced radio-sensitivity of cells in a xenograft lung cancer model through miRNA-200c expression inducing the oxidative stress response by its regulation of oxidative response genes [70]. Similarly, Shi et al. (2013) showed that A549 cells could be radio-sensitized upon miRNA-200c expression [69], while Kopp et al. (2013) showed that miRNA-200c could target K-Ras expression and that it could inhibit tumor progression and therapeutic resistance in a panel of BC cell lines [71]. Additional tumor suppressive effects have also been reported and which showed miRNA-200c expression to decrease NCCLC and A549 migration or invasiveness. MiRNA-200c was also reported to target USP25 [72], ZEB1 [73] or ZEB2 [74], and had the effect of modulating cell migration and differentiation of cells. Similarly, miRNA-200c expression was also correlated with reduced cell migration of H23 cells through enhanced E-cadherin expression [75]. Conversely, miRNA-200c was also seen to function by inducing cell death through the apoptotic pathway. For example, Bai et al. (2014) showed that miRNA-200c expression targeted the RECK gene and induced the apoptotic death of H460 lung cells, which was enhanced in the presence of Reservatol stimulation [76]. Generally, the functional role of positive miRNA-200c expression appears to be one that minimizes tumor progression and is mechanistically linked to the suppression of genes that have an oncogenic effect (Table 3).
Table 3. Elevated (+) or reduced (−) miRNA-200c levels are shown, as are their target genes, their biological effects and whether these factors can sensitize cells to certain therapeutic agents. The cell types indicate the types of cells characterized. BC, breast cancer; NSCLC, non-small cell lung cancer.
|
micro-RNA |
Target |
Negative Effect |
Sensitizing Agent |
Cell Type |
Reference |
|
200c (+) |
VEGF, VEGFR2 |
Angiogenesis, Cell Migration |
Radiation |
A549 |
[69] |
|
200c (+) |
PRDX2, SENS1, GABPA/Nrf2 |
Oxidative Response |
Radiation |
A549, H460, H1299 |
[70] |
|
200c (+) |
K-Ras |
Proliferation, Cell cycle |
− |
Lung and BC cell lines |
[71] |
|
200c (+) |
USP25 |
Cell Migration EMT |
− |
NSCLC cell lines |
[72] |
|
200c (−) |
ZEB1 |
Cell Migration |
Gefitinib |
PC-9-ZD |
[73] |
|
200c (−) |
ZEB2 |
EMT |
− |
A-549 |
[74] |
|
200c (+) |
Possibly E-cadherin |
Cell Migration |
− |
H23, A549, HCC-44 |
[75] |
|
200c (+) |
Possibly RECK |
Proliferation |
Reservatol |
H-460 |
[76] |
Simultaneously, a number of excellent studies have also published how miRNA evaluation in cells can be successfully utilized as a diagnostic and prognostic tool. For example, Tejero et al. (2014) reported that miRNA-200c could be a good biomarker for overall survival (OS) during the early stages of NSCLC adenocarcinoma [75]. Here, qRT-PCR was used to evaluate 155 resected patient tumor samples for miRNA-200c expression and their findings complimented with functional studies using H23, HCC44 and A549 cell lines. Elevated miRNA-200c expression in early stage NSCLC was significantly correlated with a decrease in OS [75]. Similarly, Kim et al. (2014) reported miRNA-200c expression to be significantly up-regulated and correlated with tumor size, lymphovascular invasion and poor OS [77]. Other publications supporting such trends have also been recently reported through the extensive use of meta-analyses to help define the diagnostic potential of miRNA-200c expression. For example, Shao et al. (2015) correlated high levels of circulating miRNA-200c with a poor OS and PFS (in advanced disease) and low miRNA-200c levels with poor survival during early stages of disease [78]. Here, 18 published studies were analyzed and the regulation of EMT (or MET) by miRNA-200c was seen as a possible cause. Teng et al. (2016) identified circulating and tissue-derived miRNA-200c as a potential diagnostic and prognostic marker for epithelial ovarian cancer (EOC) [79]. Si et al. (2017) analyzed 110 resected tumor samples from NSCLC patients for quantification of miRNA-200c, the expression of which was associated with positive lymph node metastasis, TNM classification and a reduced 5 year disease-free survival rate [80]. More recently, the use of miRNA as biomarkers for responsiveness to chemotherapeutics have also gained some attention as reported by Li et al. (2017). Here, the findings from 46 published articles showed that low expression levels of miRNA-200c (or IHC negative staining) was a good predictor for responsiveness to chemo- or radio-therapy in esophageal cancer [81]. Moreover, Zheng et al. (2017) used a meta-analysis from 60 reported studies to highlight that increased miRNA-200c expression correlated with poor prognosis in gastrointestinal cancer (GIC) patients [82], while increased miRNA-200c expression offered a better OS for ovarian cancer (OC) patients, as reported by Shi et al. (2018) [83], (Table 4).
Table 4. Elevated (+) or reduced (−) miRNA-200c levels are shown, as are the cancer types, source of materials the miRNA was detected from and the patient cohort size. NSCLC, non-small cell lung cancer; EOC, epithelial ovarian cancer; GIC, gastrointestinal cancer; esophageal cancer (ES); OC, ovarian cancer. The negative or positive use of the technique in diagnostic or prognostic evaluation of patients are denoted by − or +, respectively.
|
micro-RNA |
Cancer type |
Source |
Cohort Size |
Diagnostic |
Prognosis |
Reference |
|
200c (+) |
NSCLC |
Tissue |
155 |
− |
Reduced |
[75] |
|
200c (+) |
NSCLC |
Tissue |
72 |
− |
Reduced |
[77] |
|
200c (−) |
varied |
Tissue/Blood |
18 studies |
− |
Poor OS and PFS |
[78] |
|
200c (+) |
EOC |
Tissue/Plasma |
14 studies |
+ |
+ |
[79] |
|
200c (+) |
NSCLC |
Tissue |
110 |
− |
Reduced |
[80] |
|
200c (−) |
EC |
Tissue |
46 studies |
− |
+ |
[81] |
|
200c (+/−) |
GIC |
Tissue/Blood |
60 studies |
− |
+ |
[82] |
|
200c (+) |
OC |
Tissue/Blood |
15 studies |
− |
+ |
[83] |
The expression of miRNA-152 has been evaluated in a number of cancers associated with the gastrointestinal tract over the last 10 years with some very clear findings on which target genes may be regulated by miRNA-152 and what role they may play during cancer progression. For example, Chen et al. (2010) analyzed 101 gastric cancer (GC) and colorectal cancer (CRC) tissue samples and reported a decrease in miRNA-152 expression, which correlated with an increased tumor size and advanced pT stage in GIC, and inversely correlated with cholecystokinin B receptor protein expression in GC [84]. Other target genes for miRNA-152 include PIK3CA in breast cancer (BC) [85] or PIK3R3 in CRC [86], EPAS1 in Paclitaxel-resistant BC cells [87], CD151 in GC [88], IGF-1R and IRS1 in BC [89], B7-H1 in GC [90], CDK8 in hepatocellular carcinoma (HCC) [91], p27 in bone marrow cells [92], SOS1 in Glioblastoma (GBM) [93] cells and KLF4 in colon cancer (CC) cells [94], (Table 5).
Table 5. Elevated (+) or reduced (−) miRNA-152 levels are shown, as are their target genes, their biological effects and whether these factors can sensitize cells to certain therapeutic agents. The cell types indicate the types of cells characterized. BC, breast cancer; GC, gastric cancer; CRC, colorectal cancer; BM, bone marrow; GBM, glioblastoma; HCC, hepatocellular carcinoma; CC, colon cancer.
|
micro-RNA |
Target |
Negative Effect |
Sensitizing Agent |
Cell Type |
Reference |
|
152 (−) |
PIK3CA |
Cell Proliferation |
− |
HCC1806 |
[85] |
|
152 (−) |
PIK3R3 |
Cell Proliferation Migration |
− |
CRC cell lines |
[86] |
|
152 (−) |
EPAS |
Apoptosis |
Paclitaxel |
BC cell lines |
[87] |
|
152 (−) |
CD151 |
Proliferation Migration |
− |
GC Tissues |
[88] |
|
152 (−) |
IGF-1R |
Proliferation Angiogenesis |
− |
BC Tissues |
[89] |
|
152 (−) |
IRS1 |
Proliferation Angiogenesis |
− |
BC Tissues |
[89] |
|
152 (−) |
B7-H1 |
T-cell Proliferation |
− |
GC cell lines |
[90] |
|
152 (−) |
CDK8 |
Proliferation Apoptosis |
− |
HCC cell lines |
[91] |
|
152 (+) |
p27 |
Proliferation |
− |
BM cells, K562 |
[92] |
|
152 (−) |
SOS1 |
Proliferation Apoptosis |
Cisplatin |
GBM cell lines |
[93] |
|
152 (+) |
KLF4 |
Proliferation |
− |
CC cell lines |
[94] |
At the clinical level, Safrinzo et al. (2013) showed stage I-IIIA NSCLC patient plasma samples to contain decreased miRNA-152 expression levels, which correlated with decreased DFS for lung squamous cell carcinoma prevalence (SCC) [95]. Li et al. (2016) reported a decrease in expression of miRNA-152 in CRC tissues which inversely correlated with TNM staging and lymph node metastases [86], while Wang et al. (2017) observed a decrease in miRNA-152 expression in GC patients [90] and Ge et al. (2017) showed that miRNA-152-3p could target PIK3CA in BC as a tumor suppressor [85]. You et al. (2018) analyzed 15 GC tissues and confirmed that miRNA-152-3p expression was reduced and could directly target PIK3CA in SGC-7901 cells [96]. Alternatively, Matin et al. (2018) profiled 372 patient plasma samples collected before, during and after treatments for PC and elevated miRNA-152-3p levels reported, while (interestingly) low levels of miRNA-152-3p expression were observed in prostate cancer (PC) samples [97]. Such findings indeed highlight the power of miRNA-152 quantification as a diagnostic marker for PC (as seen for lung cancer, CRC and BC [98]). In CML, miRNA-152-3p expression was elevated in bone marrow (BM) samples and upon expression of miRNA-152-3p in K562 cells, proliferation was decreased and apoptosis levels were enhanced through targeting the p27 (CDKN1B) gene [92]. From the analysis of 89 HCC tumor samples, Yin et al. (2019) showed that miRNA-152-3p levels were decreased and which correlated with tumor volume and TNM staging [91]. Moreover, Wang et al. (2017) saw that decreased miRNA-152 expression was related to poor OS and DFS in GC, which could be used as an independent risk factor for the prediction of HCC prognosis [90]. More recently, Li et al. (2019) diagnosed early stage I-II BC by screening 106 plasma samples and tissues for miRNA-152-3p expression and reported it to be decreased, which correlated with ER-positive and PR-positive patients [99]. Finally, Song et al. (2020) observed reduced levels of miRNA-152-3p in a study of 30 invasive BC samples, which correlated with a poor prognosis [87] and the overexpression of which could sensitize chemo-resistant BC cells to Paclitaxel-mediated cell death (Table 6).
Table 6. Elevated (+) or reduced (−) miRNA-152 levels are shown, as are the cancer types, source of materials the miRNA was detected from and the patient cohort size. The negative or positive use of the technique in diagnostic or prognostic evaluation of patients are denoted by - or +, respectively. NSCLC, non-small lung cancer cells; CRC, colorectal cancer; PC, prostate cancer; BC, breast cancer; GC, gastric cancer; CML, chronic myelogenous leukemia; HCC, hepatocellular carcinoma.
|
micro-RNA |
Cancer type |
Source |
Cohort size |
Diagnostic |
Prognosis |
Reference |
|
152 (−) |
CRC |
Tissue |
28 |
+/− |
− |
[86] |
|
152 (−) |
BC invasive |
Tissue |
30 |
− |
Poor |
[87] |
|
152 (−) |
GC |
Tissues |
42 |
− |
− |
[90] |
|
152 (+) |
CML |
Bone Marrow |
40 |
- |
- |
[90] |
|
152 (−) |
HCC |
Tissue |
89 |
− |
+/− |
[91] |
|
152 (−) |
Stage I-IIIA NSCLC |
Plasma |
52 |
− |
Reduced DFS |
[95] |
|
152 (−) |
PC, lung, CRC, BC |
Plasma |
204 |
+ |
− |
[98] |
|
152 (−) |
BC stage I-II |
Plasma |
106 |
+ |
− |
[99] |
While miRNA-106b expression has indeed emerged as having a tangible biological effect in most cancer cell systems, the outcomes from such studies at this moment have offered mixed results and appears to be an area of research development. Cai et al. (2011) reported that miRNA-106b could target RB expression in laryngeal carcinoma [100] and ATG16L1 expression in Crohn’s Disease samples [101][102]. Additionally, all three micro-RNAs from the miRNA-106b-25 cluster were seen to target PTEN expression [103][104] and increased miRNA-106b expression recorded in CRC tissues which could target DLC-1 (while enhancing EMT, [105]) and FAT4 in CRC tissues or cell lines [106] (Table 7).
Table 7. Elevated (+) miRNA-106b levels are shown, as are their target genes, their biological effects and whether these factors can sensitize cells to certain therapeutic agents. The cell types indicate the types of cells characterized. CD, Crohn’s Disease; CRC, colorectal cancer.
|
micro-RNA |
Target |
Positive Effect |
Sensitizing Agent |
Cell Type |
Reference |
|
106b (+) |
RB |
Reduced Cell Arrest |
− |
Laryngeal carcinoma HEP2G+T1U212 |
[100] |
|
106b (+) |
ATG16L1 |
Decreased Autophagy |
− |
CD |
|
|
106b (+) |
PTEN |
Tumor Initiation Stemness |
Radiation |
CRC cell lines |
[104] |
|
106b (+) |
p21 (indirectly) |
Tumor Initiation Stemness |
Radiation |
CRC cell lines |
[104] |
|
106b (+) |
DLC-1 |
EMT |
− |
CRC Tissues CRC cell lines |
[105] |
|
106b (+) |
FAT4 |
Viability Angiogenesis Migration |
− |
CRC Tissues CRC cell lines |
[106] |
Based on the growing importance of utilizing miRNA expression within the clinic, their quantification for the diagnosis and prognosis of patients has moved in a positive direction. In the instance of miRNA-106b a number of excellent studies have significantly shaped this area and are worth mentioning.
As far back as 2010, Wang et al. (2010) analyzed CRC samples using qRT-PCR and found miRNA-106b to be up-regulated [107] as confirmed thereafter in colorectal cancer stromal tissues as well [108]. Subsequently, Wang et al. (2015) found miRNA-106b expression to be increased in 180 CRC patients, which correlated with a longer OS but were not seen as being statistically significant [109]. Similarly, Zhang et al. (2015) analyzed 95 CRC patient samples and miRNA-106b expression correlated with a shorter OS or DFS and which had significant reliability as an independent prognostic factor for CRC [110]. In the context of RCCC, Gu et al. (2015) performed a meta-analysis on 27 studies analyzing the expression of miRNA-106b, and (unlike CRC) reported that a decreased miRNA-106b was associated with a poor prognosis [111]. More recently, high exosomal miRNA-106b levels from the serum have been reported to correlate with a high TNM stage, a larger tumor volume and a poor prognosis [105].
Collectively, such findings support the notion that the use of miRNA-106b as a prognostic marker is unreliable, based on inconsistencies reported from a number of studies correlating miRNA expression levels with tumor grade (Table 8).
Table 8. Elevated (+) or reduced (−) miRNA-106b levels are shown, as are the cancer types, source of materials the miRNA was detected from and the patient cohort size. The negative or positive use of the technique in diagnostic or prognostic evaluation of patients are denoted by − or +, respectively. Exo, exosomal; RCCC, renal clear cell carcinoma; CC, colon cancer; CRC, colorectal cancer; OS, overall survival, DFS, disease-free survival; *, not statistically significant.
In summary, relatively good progress is being made in defining target genes for the above specific miRNAs, which may help to offer a broader perspective on how other genes of importance may synergize with cathepsin regulation in disease progression. Moreover, additional insights are also emerging into how such micro-RNAs can be utilized as reliable diagnostic and prognostic markers to possibly compliment on-going efforts with other biomarkers of importance, such as p53 and cathepsin expression. Additionally, from the above studies, oncogenic micro-RNAs are also emerging to play an important regulatory role in disease progression, and do have the potential to be targeted for therapeutic purposes using small molecule-inhibitors or -degraders (as reviewed in [112]) or through targeting specific upstream transcription regulatory signaling pathways.