1000/1000
Hot
Most Recent

Phylogenetically algae is regarded as polyphyletic as its origin cannot be traced back to single common hypothetical ancestor. However, genomic studies on algae suggest that algae evolved through endosymbiosis giving rise to at least eight to nine phyla over a period of time.
Algae diversity range from tiny unicellular microalgae to giant seaweeds which can grow over 50 m long, abundantly found in virtually every ecosystem [1]. These ancient organisms are defined as unicellular or multicellular photosynthetic organisms regarded as polyphyletic due to morphological resemblance with plants but do not share a common ancestor [2]. Similar to plants, algae are autotrophs, photosynthesize, and produce the same storage compounds and defense strategies. On the contrary, algae do not possess specialized root, stem, nor vascular bundles, and they lack diploid embryo stage. The characteristic features of algae include autotrophic chlorophyll-bearing thalloid plant body, lacks sterile tissue around its reproductive structures, and zygote development is by mitosis or meiosis but not via embryo formation [3]. Algae are not only diverse in their size but also in their ecological distribution, cellular biology, photosynthetic pigments, structural and reserve polysaccharides, and evolutionary origin of heterogeneous algae include prokaryotic and eukaryotic species [4].
Algae are polyphyletic as their origin cannot be traced back to single common hypothetical ancestor. The symbiogenic events believed to have occurred more than 1.5 billion years ago were the reason algae came into existence, and, because of so much time gap, it is difficult to track the key events that resulted in algae occurrence [5]. It is thought that they came into existence when a photosynthetic cyanobacteria invaded a unicellular eukaryote giving rise to double-membranous primary plastid. This is known as primary endosymbiosis and as a result of this endosymbiosis Green algae (Chlorophytes and other land plants), Red algae (Rhodophytes), and Glaucophytes came into existence [6]. Another secondary endosymbiosis occurred when other heterotrophic eukaryotes engulfed red and green algae, due to which many other algal species came into existence, forming three or four membranes of secondary plastid [7]. The molecular sequence analysis suggests that there exist around eight to nine major phyla (divisions) of algae. These are the cyanobacteria (chloroxybacteria) and the eukaryotic phyla Dinophyta (dinoflagellates), Glaucophyta, (glaucophytes), Cryptophyta (cryptomonads), Euglenophyta (euglenoids), Ochrophyta (a diverse array of tiny flagellates, chrysophyceans, diatoms, brown algae, and many other groups), Haptophyta (haptophytes), Rhodophyta (red algae), and Chlorophyta (green algae) [8].
There is a general consensus that first photosynthetic eukaryote emerged from a heterotrophic eukaryote which captured a coccoid cyanobacterium about 1.6 billion years ago. Gradually, the cyanobacterium was enslaved, and the endosymbiont’s genes transferred to the host nucleus as a new organelle called as primary plastid. This event is named as primary endosymbiosis [9]. The genome of cyanobacteria underwent evolution, resulting in either loss or transfer of genes to the host nucleus in a process termed as endosymbiotic gene transfer (EGT). As a result, only little portion of genome retained within primary plastid, including some genes encoding for photosynthesis [10]. The primary endosymbiosis gave rise to three groups, having primary plastids, forming a major group of autotrophic eukaryotes Archaeplastida: Glaucophytes, Red algae, and Green algae. Genomes of primary plastids consist of highly conserved genes employed extensively to study the evolution of diverse Archaeplastida groups [11].
The Red algal lineage or Rhodophyta consist of 5000–6000 multicellular eukaryotic marine algal species. Phylum Rhodophyta is comprised of two sub-phyla Rhodophytina and Cyanidiophytina with seven classes [12]. Rhodophyta have evolved with plastids lacking chlorophyll accessory pigments, rather, containing chlorophyll-a, phycocyanin, and phycoerytherin, so light is captured by phycobiliproteins [13]. Secondly, there is complete absence of centrioles and flagella [14]. The genome in the red algae are compact in nature and have high gene capacity. The mitochondrial genome in Florideophyceae are considered highly conserved despite the loss of 32 genes, which suggest endosymbiotic gene transfer [15]. Genomic and transcriptomic analysis of four different classes of Red algae (Compsopogonophyceae, Stylonematophyceae, Rhodellophyceae, and Porphyridiophyceae) reveals significant evolutionary insights. The proteins from the plastid genome of 37 red algae were translated and analyzed, which supported the idea that all four classes show monophyly and are deeply rooted in red algal phylogenetic tree [16].
Green algae are termed as Chlorophyceae, and they are characterized by the presence of double membranous chloroplasts, stacked thylakoids, and photosynthetic pigments: chlorophyll-a and b [17]. Within their plastids they have starch molecules as polysaccharide reserve, a characteristic similar to green plants. The Chlamydomonas reinhardtii mutants showed starch biosynthesis similar to starch biosynthesized by maize endosperm [18]. Chlorophyceae contains five lineages: Chlamydomonadales, Chaetopeltidales, Oedogoniales, Sphaerpleales, and Chaetophorales [19]. They emerged about 470 million years ago, which marks one of the important evolutions of time, adapted all the photic zones of Earth, and paved way for the evolution of other life [20].
The glaucophytes (glaucocystophytes) are a group of freshwater microalgae which contain blue-green plastids often called as cyanelles. In five genera of glaucophytes, about thirteen species have been described. Glaucophyte plastids are similar to cyanobacteria, and different from plastids of other algae, in having a thin peptidoglycan wall and contain only phycobilins and chlorophyll a [21]. Glaucophyte is a poorly studied genera probably due to their rareness in nature and restricted habitat. Genomic data and a better understanding of the phylogenetic position of glaucophytes will provide valuable insights into the endosymbiotic origin and evolution of plastids in eukaryotes [22].
A phylogenomic study revealed a sister-group relationship between green or red algae and glaucophytes with a bootstrap value ≥ 90% [23]. The comparison of genome of diatom with other organism revealed that many genes present in its genome were obtained by the horizontal gene transfer (HGT) from bacteria. Moreover, viruses as gene transfer agents are of considerable importance in the marine environments [24]. Along with achieving new genes with HGT and EGT, gene loss, fusion and duplication have also added to size of algal genome. There is a great difference in the genome size among eukaryotes because of genome organization, like gene distribution. For example, Ostreococcus tauri, contain 8166 genes located on 20 chromosomes and spanning a genome size of 1.54 kb/gene [25].
Algae outside Archaeplastida have their plastids originating from acquisition of primary plastids during secondary endosymbiosis. This gave rise to haptophytes, cryptophytes, dinoflagellates, heterokonts, and other photosynthetic eukaryotes [26]. The existence of secondary endosymbiosis was first indicated by the occurrence of more than two envelope membranes around the secondary plastids. As evident from the presence of four membrane bounding plastids in chlorarachniophytes, cryptomonads, ochrophytes, and haptophytes, dinoflagellates and euglenoids appear to have lost one of these membranes possessing three [27]. Moreover, nucelomorph (remnant of endosymbiont nucleus) can be found within chlorarachniophytes and cryptophytes [28]. The process of algal evolution through endosymbiosis is depicted in Figure 1.
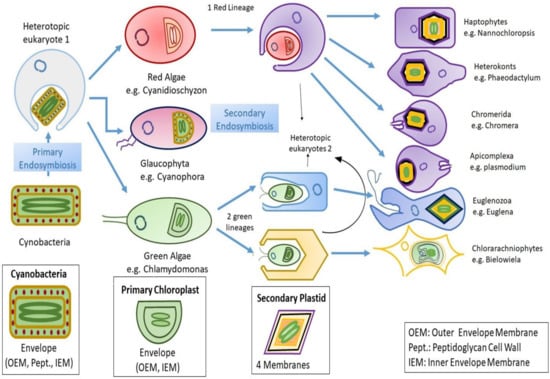
Figure 1. Diagrammatic representation of endosymbiosis: Primary endosymbiosis occurs when a heterotopic eukaryote engulfs a prokaryote (gram negative cyanobacteria), which leads to emergence of three lineages bearing primary chloroplast with two membranes that it vertically inherited from the cyanobacteria. Later on, the independent secondary endosymbiosis of green algae (eukaryote) by two unknown heterotopic eukaryotes that leads to emergence of Euglenozoa and Chlorarachinoophyte. Red algae also go through similar mechanism of secondary endosymbiosis. The consequent secondary plastid contains four membranes, out of which two are the ones that were inherited from the cyanobacteria during primary endosymbiosis, and the other two are still unknown.
To add to the complexity of algal evolution, there seems to exist tertiary endosymbiosis too. Some dinoflagellates have gained their plastid either by replacing the red-alga-derived plastid with a green-alga-derived plastid, or by capturing secondary endosymbiotic cryptophyte, haptophyte or heterokont alga in tertiary endosymbiosis [29]. Genomic data is rapidly accumulating to help us further understand the evolutionary process of photosynthetic eukaryotes, which appears to be a complex and haphazard process.