1000/1000
Hot
Most Recent

In humans, the ability to digest milk lactose is conferred by a β-galactosidase enzyme called lactase-phlorizin hydrolase (LPH). The LPH enzyme is encoded by the lactase (LCT) gene, located on the chromosome 2q21. Exclusively expressed in the small intestine, in the apical part of microvilli within the brush border membrane of enterocytes, the LPH enzyme reaches the highest levels of activity during the nursing period. After the weaning phase, however, in the majority of humans, the activity of LPH declines rapidly because of a decrease in the levels of the enzyme, and this trait is known as lactase non-persistence (LNP). As a consequence of LNP, the majority of humans are unable to digest lactose during adulthood, and some suffer clinical complications when they consume it. It has been estimated that approximately two-thirds of humans are LNP worldwide. In the remaining third, however, there are individuals with the ability to digest milk and other lactose-rich dairy products during adulthood. This trait is known as lactase persistence (LP), and is particularly common in descendants from populations that have traditionally practiced cattle domestication.
The frequency of LP is high in northern European populations, decreases across Southern Europe and the Middle East, and is low in non-pastoralist Asian and African communities[1][2][3][4]. Notably, LP is also common in pastoralist populations from Africa. It is hypothesized that natural selection has elicited a prime role in determining the current frequencies of LP in different human communities since the development of cattle domestication in the Middle East and North Africa around ~7500–9000 years before present (BP). Although LP and LNP frequencies have been widely studied worldwide, there is a lack of a work collecting all newly available information published to date[5][6][7][8][9][10][11][12][13][14][15][16]; especially the new frequency data reported for American populations. Given the great interest of this research topic, an online graphic resource gathering all reported lactase frequency data (both at the phenotypic and genetics levels) would be of much utility for the scientific community. With this purpose, interactive maps and other geographic information system (GIS) tools have been extensively employed with great success in other evolutionary biology areas[17]. Interactive mapping involves using maps that allow zooming in and out, panning around, identifying specific features, and querying underlying data, such as by topic or an accurate indicator (e.g., the allele frequency of a certain LP variant in a particular sub-population or ethnicity), generating reports, and other means of using or visualizing information in a map. This allows a more attractive way of presenting large sets of data that would not be sufficiently well exploited in static plots or large tables. Moreover, the online format of these resources allows easy access for researchers and constant update.
On the other hand, although great efforts have been devoted to elucidating the exact molecular mechanisms responsible for the natural decline in LPH activity after weaning (LNP), some gaps remain to be fulfilled[18]. Neither is there a unique, accepted theory about the evolutionary origin of LP and what was the advantage that it conferred to carriers, and that favoured such a transmission across generations.
Anguita-Ruiz A et al. (2020) have recently published a review on the genetics of Lactase phenotypes. The aims of the review were: (1) to gather and summarize all available information on LNP and LP genetic mechanisms and evolutionary adaptation theories, and (2) to create online interactive world maps, including all LP phenotype and genotype frequency data reported to date. The work constitutes the first interactive, and most updated, online resource for studying and exploring the phenotypic and genetic data on lactase phenotypes (http://bionit.ugr.es/pages/investigacion/software/bioinformatics-methods-software). The review is intended for general clinicians, nutritionists, dietitians, paediatricians, evolutionary biology researchers, geneticists, and any researcher working on the topic.
Bellow, the main conclusions of the work can be found:
Lactase phenotypes (LP and LNP) in humans present a complex genetic basis that has been widely investigated during the last decades. Although great advances have been achieved, the exact molecular mechanisms underlying the appearance of LNP after the weaning phase remain not fully understood. Probably, the combined action of transcription factors and epigenetics alterations on the LCT gene is the most plausible explanation. In the present review, we have summarized all proposed explanatory theories and gathered them in a figure (Figure 1). The mechanisms responsible for LP, on the other hand, are known in better detail, and respond to the existence of cis-element mutations mapping a gene other than LCT; the regulatory enhancer region MCM6. Particularly, there have been a total of twenty-three SNPs within the MCM6 associated with LP so far in human populations. These variants seem to have arisen during the same period but independently in different human populations, the reason why LP has become a textbook example of convergent regulatory evolution and gene-culture co-evolution.
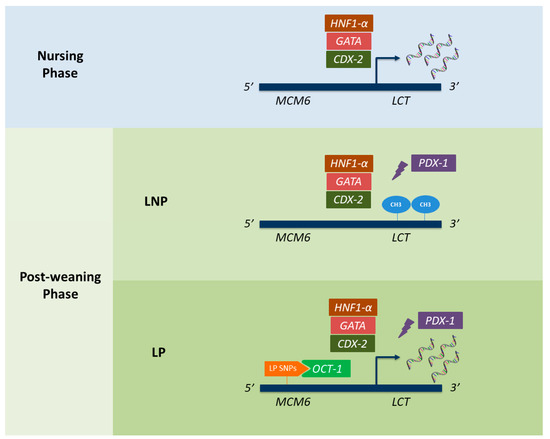
Figure 1. Genetic mechanisms underlying lactose persistence (LP) and lactase non-persistence (LNP) in humans. Rectangular shapes represent all the transcription factors currently known to interact with the lactase gene (LCT) promoter. While CDX-2, HNF1-α, GATA, and OCT-1 are known to promote LCT expression, the PDX-1 has been described as a transcriptional repressor. Oval shapes (named as “CH3”) refer to the appearance of methylations within the LCT region, which have also been described as repressing LCT expression. Finally, the different LP-associated alleles described in MCM6, and responsible for binding to OCT-1, are represented in orange. CDX-2: caudal type homeobox 2; HNF1-α: hepatocyte nuclear factor 1α; OCT-1: octamer-binding protein 1; PDX-1: pancreatic and duodenal homeobox 1.
The distribution of lactase phenotypes, and their associated SNPs across human populations, has also been widely studied though not recently reviewed. All available information has always been presented in the form of static world maps or large-dimension tables, so that it would benefit from the newly available visualization tools, such as interactive world maps. In the present review, besides gathering all information available to date for LP phenotype and genetic frequencies, we have also created two online interactive resources (http://bionit.ugr.es/pages/investigacion/software/bioinformatics-methods-software). As fully expandable and interactive graphics, our tools further constitute an upgrade over previously published static world maps, allowing users a personalized data exploration at the same time as accessing complete reports by population or ethnicity. Despite the great availability of data, we have detected that more studies are needed, especially for certain Central and South America regions, for which LP phenotype/genetics data lack entirely. Likewise, there is also a need for more genetics studies on the frequency of genetic variants other than the −14010:G>C (especially in non-Caucasian populations). Finally, and due to the existence of certain studies pointing to epigenetics as a plausible contributor to lactase phenotypes, it would also be interesting to carry out more population studies focused on the distribution of LCT epigenetics signatures associated with LP/LNP.
Concerning evolutionary genetics, LP is one of the strongest examples of positive selection found in the human genome. According to the latest studies, the evolutionary advantage conferred by LP alleles mainly lies in the lifetime access to nutrient-rich milk in populations that have traditionally practiced cattle domestication. However, there are still some discrepancies unexplained, as is the case of several ancient pastoral populations with a very low or null frequency of LP. As possible explanations, variability in milk fermentation practices and population admixture events have been proposed. On this matter, more nutritional anthropological studies investigating the amount, type, and seasonality of milk and dairy products consumed, and the perception of these foods in traditional populations would be helpful in understanding why some populations took up drinking fresh milk, whereas others mostly transformed it.
Human gut microbiome is, in a way, an extension of the human genome; as thousands of metabolic processes performed by members of the microbial community directly influence host physiology, including their host’s ability to utilize lactose and other carbohydrates. Hence, further studies of utilization of lactose by the human microbiome are needed to explain discrepancies found in LP/NLP phenotypes.
Research works like the present review lay the foundations for the study of the genetic bases of lactase phenotypes in humans, and represent a new paradigm in the way of visualizing genetic and phenotype data at the population level.