1000/1000
Hot
Most Recent

Tetrahydrofuran (THF) is widely used as a precursor for polymer syntheses and a versatile solvent in industries. THF is an environmental hazard and carcinogenic to humans. We characterized the THF degradation potential of a number of THF-degrading bacteria reported before and a new isolated filamentous fungus Pseudallescheria boydii ZM01. Two different microbial THF degradation pathways have been proposed here. In addition, The initial key metabolic intermediate 2-hydroxytetrahydrofuran was detected and identified by gas chromatography (GC) analyses for the first time during the THF degradation process.
Tetrahydrofuran (THF) is a colorless, highly soluble, and volatile heterocyclic compound that is widely used as an important basic feedstock for chemical syntheses and is an excellent solvent for lacquers and printing inks [1][2]. With the increasing consumption of THF, large amounts of industrial wastewater containing THF have been released into groundwater, rivers, and lakes [3][4]. Furthermore, increased attention has been paid to the carcinogenic effects of THF-polluted groundwater on human health after its toxicological effects were thoroughly demonstrated by a variety of assays in vivo. Various studies in female mice have demonstrated that THF can enhance tumor formation by inducing cell proliferation, can cause nervous system dysfunction, and can induce DNA damage [5][6][7]. Considering the advantages of microbial degradation, including the safety of disposal and cost effectiveness, the key to refractory organic pollutant treatment will be the exploitation of microbes from the natural environment with highly efficient pollutant-degrading activities [8][9]. THF is also classified as a refractory pollutant due to the high bond energy obstruction of C–O (360 KJ mol−1) [10]. A number of bacteria, including gram-positive Rhodococcus sp., Pseudonocardia sp., and Mycobacterium sp. and gram-negative Flavobacterium sp. and Pseudomonas sp. have been reported to aerobically grow on THF [11][12][13][14][15]. Although many THF-degrading bacteria have been reported, most of them cannot utilize THF at concentrations higher than 35 mM, and the lag phase is prolonged with increased initial substrate concentrations. The widely reported THF-degrading strain Rhodococcus ruber YYL showed the highest THF tolerance concentration of up to 200 mM among all the reported THF-degrading strains. Strain YYL can completely degrade 6 mM THF within 32 h without a lag phase, whereas the lag phase increased to more than 96 h when the initial concentration of THF was increased to 100 mM [16]. R. aetherivorans M8 exhibits an obvious lag phase of 5 days in the presence of 15 mM THF, which was prolonged to more than 15 d when the THF concentration was increased up to 30 mM [17]. Pseudomonas oleovorans DT4, which has a superior maximum THF degradation rate compared to the other reported THF-degrading strains, cannot mineralize THF completely in the presence of THF concentrations higher than 30 mM, with the degradation ratio decreasing to 45% with 40 mM THF [1]. Therefore, growth inhibition and incomplete degradation are major issues with respect to the ability of bacteria to efficiently degrade THF when under the stress of high substrate concentrations.
Filamentous fungi are involved in multiple pathways associated with wastewater treatment, such as organic compound biodegradation, the formation of sludge flocs, settleability of the treated sludge, and sludge detoxification [18]. Filamentous fungi grow as thread-like hyphae and reproduce by sporulation [19]. The hyphae can immobilize sludge particles, resulting in the formation of pellets, which leads to microbial aggregation as natural flocculants [20][21]. Notably, filamentous fungi can also improve the dewaterability and filterability of sludge by modifying its porosity structure [22]. Many studies have shown that compounds that are highly toxic toward bacteria may be harmless to fungi, such as antibiotics [21]. In addition, the high contents of functional groups in the cell walls of fungi, such as amino, amide, and phosphate groups, promote the ability of fungi to absorb low concentrations of heavy metals as micronutrient sources and to resist heavy metal stress [23]. The ability of filamentous fungi to form mycelial networks, to undergo detoxification, and to maintain stress resistance and richness in terms of biodiversity enhances their potential application in bioremediation [24][25]. To date, only three fungi, including Aureobasidium pullulans, Cordyceps sinensis [26], and Graphium sp. ATCC 58400 [27] have been reported to grow on THF, and the THF degradation characteristics of fungi have rarely been evaluated. Thus, studies on the THF degradation characteristics of THF-degrading fungi is important for developing bioremediation technologies for THF-containing wastewater.
Two different microbial THF degradation pathways have been proposed in previous studies, but some metabolites remain speculative, such as 2-hydroxytetrahydrofuran. In the oxidation pathway, THF is inferred to be initially oxidized to 2-hydroxytetrahydrofuran, and 2-hydroxytetrahydrofuran is then oxidized to γ-butyrolactone before being transformed to 4-hydroxybutyrate. 2-hydroxytetrahydrofuran is a cyclic hemiacetal that can also autonomously form its tautomer 4-hydroxybutyraldehyde, which can be directly converted to 4-hydroxybutyrate. The metabolite 4-hydroxybutyrate, which is common to both metabolic pathways, is further oxidized to succinate and then thoroughly mineralized in the tricarboxylic acid cycle [17][28][29]. Several THF degradation intermediates in the proposed THF degradation pathway have been detected and verified in previous studies. Small amounts of γ-butyrolactone were detected as an intermediate in Graphium sp. ATCC 58400, Mycobacterium vaccae JOB-5, and Rhodococcus sp. DTB after incubation with THF [27][29][30]. In addition, the derivatized product of 4-hydroxybutyrate was finally detected and confirmed by gas chromatography-mass spectrometry (GC–MS) in the resting cell reaction of R. aetherivorans M8, which was initiated by the addition of 5 mM THF [17]. However, the initial key monooxygenation metabolite in both degradation pathways, 2-hydroxytetrahydrofuran, has not been detected and confirmed during the THF degradation process in all reported THF-degrading strains thus far [31][32].
The detection of the THF concentration was conducted as described in a previous study [33]. The THF concentration was measured by GC-2014C gas chromatography equipped with a flame ionization detector (FID) and an AOC-20i Auto injector (SHIMADZU, Shanghai, China). All samples were centrifuged at 10,000 rpm for 2 min, and the supernatant was subsequently collected after being filtered through a 0.45-μm filter for pretreatment. Samples at different time points corresponding to the growth curve were taken to detect the intermediate metabolites. The detection program was as follows: the initial temperature was set at 60 °C, gradually increased to 160 °C at a rate of 20 °C min−1, and then held at 160 °C for 5 min. The reference standards, such as THF, 2-hydroxytetrahydrofuran, and γ-butyrolactone, were appropriately prepared at different concentrations to generate standard curves for use in intermediate identification and concentration calculations.
To identify the intermediate metabolites of THF produced by strain ZM01, the intermediate metabolites were concentrated by organic solvent extraction. To this end, 20 mL of ethyl acetate was added directly to 70 mL of fermentation liquid, and the mixture was vigorously shaken for 20 min to completely extract the metabolites. The extraction phase was then evaporated to dryness and dissolved in 1 mL of ethyl acetate for GC-MS analysis. The final concentrated metabolic intermediates were identified using a GC-MS instrument (Agilent 7890B-7000C, Shanghai, China) equipped with an HP-5ms capillary column (30 m × 0.25 mm × 0.25 μm). The column oven temperature was maintained at 40 °C for 3 min and then increased to 120 °C for 2 min at a rate of 10 °C min−1. The eluted compounds were ionized by electron impact (70 eV) after passing through a GC-MS interface maintained at 250 °C. Ions with masses ranging from 10 to 400 atomic mass units were scanned at 1-s intervals. Compounds were identified by comparing their fragmentation patterns with those of the authentic standards and the mass spectra library (National Institutes of Standards Technology, Gaithersburg, MD, USA).
A filamentous fungus capable of growing in MSM supplemented with THF as the sole carbon source was successfully isolated from activated sludge by enrichment cultivation and designated strain ZM01. Strain ZM01 could grow as white thread-like hyphae and could develop conidia growing directly on vegetative hyphae after cultivation for 7 d at 30 °C. Most conidia were usually brown, thick-walled, globose to subglobose. The shape of conidiogenous cells was cylindrical, and no yellow diffusible pigment was produced during the cultivation of strain ZM01 on PDA at 30 °C. The microscopic characteristics of strain ZM01 were consistent with the typical microscopic features of Pseudallescheria sp. Based on phylogenetic analysis of the ITS region sequences (GenBank Accession No. MT754398) (Figure 1), strain ZM01 was observed to be closely related to P. boydii (GenBank Accession No. AY213683) with a high degree of similarity (99%). Thus, on the basis of morphological characteristics and phylogenetic relationships analysis, we identified the isolated THF-degrading fungus as Pseudallescheria boydii strain ZM01. To the best of our knowledge, this is the first report showing that THF can be metabolized by a Pseudallescheria sp. strain.
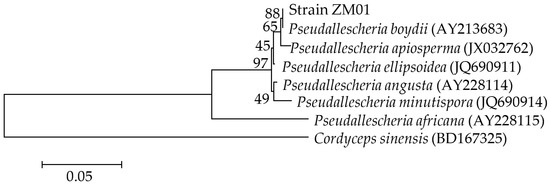
Figure 1. The phylogenetic dendrogram for strain ZM01 and closely related Pseudallescheria strains based on the internal transcribed spacer (ITS) region sequences: The representative tetrahydrofuran (THF)-degrading fungus C. sinensis is used as the root. Numbers after the names of organisms are accession numbers of published sequences. Numbers adjacent to the branches indicate the bootstrap values based on 1000 replicates.