1000/1000
Hot
Most Recent

Fatty Acid esters of Hydroxy Fatty Acids (FAHFAs) are ester derivatives of fatty acids (FAs) with hydroxy fatty acids (HFAs) carrying the hydroxyl group at different positions. This novel class of endogenously generated lipids, discovered in 2014, has attracted the interest, because such lipids have been found to exhibit anti-diabetic and anti-inflammatory effects. The great number of possible combinations of FAs with HFAs together with the great number of possible positional isomers makes the analysis of FAHFAs very challenging. FAHFAs have been detected and quantified in a variety of human and animal tissues, as well as in plant tissues and foods. Analytical methodologies, either targeted or untargeted, may highly contribute in studying FAHFA levels under various biomedical conditions, and in understanding their role in physiological and pathological conditions.
Fatty acid esters of hydroxy fatty acids (FAHFAs) constitute a class of recently identified novel lipids exhibiting anti-diabetic and anti-inflammatory effects. Due to their high biological significance, a tremendous effort has been devoted to the development of analytical methods for the detection and quantitation of FAHFAs during the last five years. The analysis of FAHFAs is very challenging due to the great number of possible regio-isomers arising from the great number of possible combinations of FAs with HFAs, and the low abundancies of FAHFAs in biological samples.
During the last two decades, a variety of endogenously generated lipids have been recognized as biomolecules that dynamically affect human physiology and pathophysiology [1]. In 2014, Kahn, Saghatelian and co-workers reported the discovery of a previously unrecognized class of endogenous mammalian lipids exhibiting anti-diabetic and anti-inflammatory effects [2]. Employing a quantitative mass spectrometry (MS) lipidomics approach, they compared a big set of ions in adipose-selective overexpression of Glut4 (AG4OX) and wild type (WT) mice and they observed a cluster of ions in AG4OX mice that was elevated >16-fold. The measured accurate mass of these ions enabled them to calculate the molecular formulas, indicating that all were members of a single lipid class containing a unique signature of four oxygen atoms. However, such formulas did not correspond to any metabolite known at that time in metabolite databases, including LIPIDMAPS [3]. Fragmentation studies suggested that these lipid species consisted of hydroxy fatty acids (HFAs) acylated by fatty acids, and thus these lipids were named as fatty acid esters of hydroxy fatty acids (FAHFAs). Continuing with a targeted MS approach, they indeed identified in mouse serum 16 FAHFA family members that consisted of four fatty acids and four hydroxy fatty acids in different combinations [2].
Since that time, several branched FAHFA families have been identified in serum, mammal tissues, plants and foods. Each one consists of multiple regio-isomers, in which the hydroxy group participating in the ester bond is at different positions (e.g., 5- or 9-). The general structure of FAHFAs is depicted in Figure 1. It should be noticed that in 1982, Nicolaidis and Ruth reported in human and steer meibum an unusual group of high molecular weight fatty acid derivatives, which were identified as (O-acyl)-ω-hydroxy fatty acids [4]. These lipids differ from those identified and studied during the last six years, because the ester group is at the ω-position, making them linear and not branched.
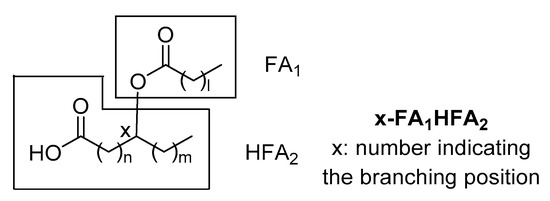
Figure 1. General structure of fatty acid esters of hydroxy fatty acids (FAHFAs).
During the last five years, a tremendous effort has been devoted to the development of analytical methods for the detection and quantitation of FAHFAs in biological samples as well as in plants and foods. The great number of possible combinations of FAs with HFAs together with the great number of possible regio-isomers makes the analysis of FAHFAs very challenging due to overlapping of the various regio-isomers in the liquid chromatography (LC) conditions.
The major FAHFAs that have been detected are a combination of palmitic acid (PA), stearic acid (SA), oleic acid (OA), or palmitoleic acid (PO) with their corresponding hydroxy fatty acids (HPA, HSA, HOA, HPO respectively), providing PAHPA, OAHPA, PAHOA, OAHOA, PAHSA and OAHSA [2][5] (Figure 2). A variety of regio-isomers have been identified with the ester linkage at position C5, C7, C8, C9, C10, C11, C12 and C13. Although most of the FAHFAs contain saturated lipid chains, also polyunsaturated FA-derived FAHFAs have been uncovered, including omega-3 polyunsaturated-derived and linoleic-derived FAHFAs [6][7]. Samples of the major FAHFAs are commercially available by various companies. In addition, methodologies for their synthesis, either in racemic or chiral form, have been developed [2][8][9][10][11].
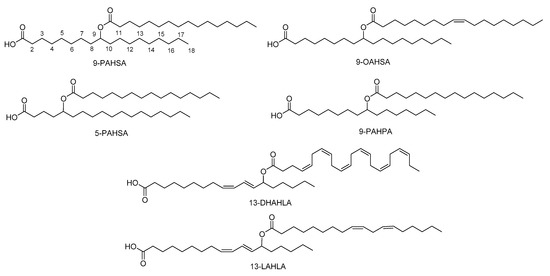
Figure 2. Representative structures of FAHFAs.
As described in the seminal work of Kahn and coworkers, the structure of PAHSA was elucidated by mass spectrometry studies [2]. The fragmentation of the precursor ion observed at m/z 537 generated ions with m/z 255, 281 and 299, which were attributed to palmitic acid (PA), octadecenoic acid and hydroxystearic acid (HSA), respectively (Figure 3). Since the ion with m/z 537 did not contain any double bond, octadecenoic acid resulted from fragmentation in the MS and, thus, PAHSA was proposed as an ester between PA and HSA. The ions observed at m/z 127 and 155, also recorded in an MS3 experiment [6], indicated the position of hydroxyl group at C-9 of stearic acid (Figure 3).
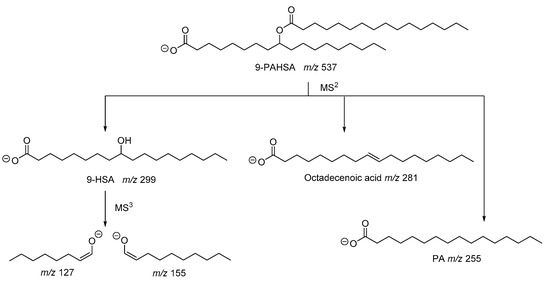
Figure 3. Fragmentation pathway of 9-PAHSA.
In 2015, α computer-generated library with 3267 tandem mass spectra (MS/MS) for 1089 FAHFA species was presented [12]. In silico electrospray ionization (ESI) MS/MS spectra in negative ion mode were modelled based on the reference spectra of 9-PAHSA obtained under 10, 20 and 40 V collision-induced dissociation (CID) voltages acquired with ultra-high performance liquid chromatography-quadrupole time-of-flight (UHPLC-QTOF) MS/MS profiling methods at 4 spectra/s. To generate the fragments with m/z 127 and 155, indicating the position of the hydroxyl group, MS/MS experiments with a longer acquisition time of 1 spectrum/s were required at a higher voltage (40 V).
Determination of the ester position in isomeric FAHFAs was studied by ion trap mass spectrometry [13]. The CID mass spectra of precursor [M–H]− ions for 5-PAHSA, 9-PAHSA, 12-PAHSA and 18-PAHSA showed the same three predominant product ions in varying ratios. However, the MS3 spectra acquired from 5-HSA, 9-HSA, 12-HSA and 18-HSA revealed distinct diagnostic product ions unique to each isomer in the region below m/z 240, allowing for pinpointing of the ester location.
The fragmentation pathway of the polyunsaturated 13-DHAHLA is shown is Figure 4. The precursor ion (m/z 605) generated the fragment with m/z 295, which subsequently gave rise to fragments with m/z 195 and m/z 179, indicating the position of hydroxyl group [6]. Similar fragments were produced from the fragmentation of 13-LAHLA [7].
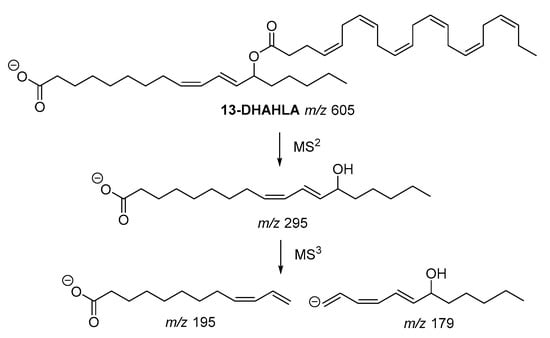
Figure 4. Fragmentation pathway of 13-DHAHLA.
MS studies play a pivotal role in the elucidation of the structure of these natural lipids. In particular, MSn fragmentation studies allow the determination of the branching position, discriminating the various positional isomers.