Liquid biopsy, which surmounts the limitation of tissue biopsy, is evaluated as a potential tool for early cancer detection and monitoring. By sampling from blood, stool, urine, saliva, and other fluid samples, liquid biopsy provides a non-invasive and feasible cancer detection service. Compared with tissue biopsy, liquid biopsy is also more comprehensive to evaluate tumor heterogeneity since tumor sites can release aberrant signals into body fluid.
1. Introduction
When cells mutate, they could divide uncontrollably and eventually form cancer [1]. According to the World Health Organization, cancer accounts for nearly 10 million deaths in 2020. Unfortunately, this number is estimated to be still climbing in the following decades and will reach 27 million new cases in 2040 [2]. As the second factor of death, cancer accounts for one-sixth of deaths worldwide each year [3]. Therefore, fighting against cancer is a huge challenge for global public health. Early detection, followed by tailored site-specific treatment, plays an important role in the front-line cure of cancer and could reduce the eventual mortality of cancer patients [4][5][6].
Cancer is associated with mutated genes; and genetic analysis is increasingly applied in cancer diagnosis [7]. The traditional methods for genetic testing on cancer patients are sampling from tumor tissues. However, tumor tissue biopsy is limited by several drawbacks such as invasive acquisition, clinical complications, sample preservation, and tumor heterogeneity [8][9][10].
Liquid biopsy [7][11], which surmounts the limitation of tissue biopsy, is evaluated as a potential tool for early cancer detection and monitoring [12]. By sampling from blood, stool, urine, saliva, and other fluid samples, liquid biopsy provides a non-invasive and feasible cancer detection service [13][14][15][16]. Compared with tissue biopsy, liquid biopsy is also more comprehensive to evaluate tumor heterogeneity since tumor sites can release aberrant signals into body fluid [17][18]. Researchers paid significant attention to the different components from liquid biopsy which are associated with cancers [19][20][21][22][23].
As the possibility or severity of tumor in the body is relevant to the liquid biopsy components, accurate cancer prediction based on the characteristics of these components becomes a significant problem. The application of machine learning protocols has been widely studied in recent years, proving to be valuable in early cancer detection. Nevertheless, the required knowledge to implement these methods is high, posing an obstacle to researchers who are looking to get started on liquid biopsy analysis and early cancer detection.
2. Liquid Biopsy Components
During the formation and growth of primary tumors, cells undergo active release, necrosis, or apoptosis [24][25]. In these process, various components are released into the liquid, including circulating tumor cells, cell-free DNA, circulating tumor DNA, cell-free RNA, exosomes, and tumor educated platelets(TEPs) [26].
The presence of circulating Tumor Cells (CTCs) was firstly identified by Ashworth (Australia) in 1869 [27]. When Ashworth performed an autopsy on a metastatic breast cancer patient, cells similar to those from the primary tumor were found in the blood. CTCs are currently defined as the tumor cells that shed or migrate actively into the vessel from the primary tumor or metastatic sites and then circulate in the bloodstream [28]. The opinion of tumor self-seeding suggests that CTCs can recirculate back, resulting in the possibility of metastases, which is responsible for the majority of deaths associated with cancer [29][30]. As the access to peripheral blood circulation is a prerequisite for distant metastasis of tumors [31], detection of tumor cells in blood will indicate the possibility of distant metastasis of tumors [32].
CTC is isolated from peripheral blood, which can avoid invasive and complex biopsy procedures. The culture of tumor cell lines takes a long time and is homogeneous, which cannot accurately reflect the genetic diversity and the changing tumor microenvironment. In contrast, CTCs-derived xenografts can reflect the biological characteristics of cancer more accurately, providing a visual window for studying the dynamic evolution of cancer and allowing monitoring of the longitudinal evolution of tumors at the molecular level.
Platelets (also termed thrombocytes) are the second most abundant cell types in peripheral blood, existing as circulating anucleated cell fragments. The largest platelets are about 2–3 microns in diameter [33]. More recently, platelets are implicated a central role in the local and systemic responses to tumor growth [34][35]. Confrontation of platelets with tumor cells by transferring tumor-associated biomolecules (‘education’) is an emerging research field resulting in the term of tumor-educated platelets (TEPs).
3. Machine Learning Algorithms and Clinical Application in Early Cancer Detection Based on Liquid Biopsy
Several machine learning algorithms are used to detect cancer based on the characteristics extracted from liquid biopsy. An overview of all relevant papers are listed in the
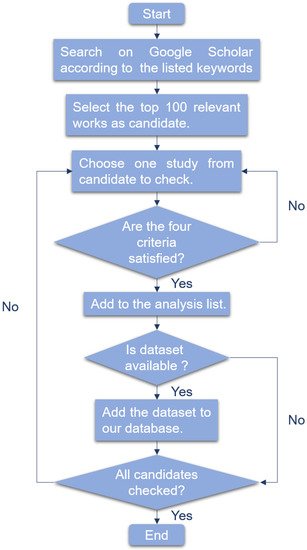
supplementary document (Table: Summary of related publications) with the direct URL of dataset if available. This section discusses and reviews the publications of the most commonly used algorithms for early cancer detection in recent 10 years. As this systematic survey aims to report wide studies related to early cancer detection based on liquid biopsy incorporating machine learning algorithms, over 400 papers were searched using the following keywords: (liquid biopsy OR exosome OR circulating tumor cell OR circulating tumor DNA OR cell free DNA OR microRNA OR tumor educated platelet) AND (cancer OR carcinoma OR adenocarcinoma OR tumor OR malignancy OR malignant disease) AND (svm OR support vector machine). We searched four extensively used machine learning algorithms by replacing the last keyword. For each algorithm, we checked the top 100 relevant publications in recent 10 years according to the following four criteria.
Figure 1 is the workflow of select publications.
Figure 1. Workflow of search adn select publications.
-
The research is about liquid biopsy.
-
The research is about cancer detection.
-
The research utilized corresponding machine learning method.
-
For several models compared, we only consider the model which performs best.
4. Discussion
From the perspective of machine learning, we find out that even simple machine learning algorithms such as linear models can lead to a high-quality performance for liquid biopsy-based diagnosis for several common cancer types. However, there is no perfect model that performs the best on all datasets. Besides, the performance of machine learning models is diverse under different hyperparameter settings. To ensure the stability, we recommend Bayesian optimization for hyperparameter tuning after considering performance and runtime. With a hyperparameter optimization strategy, the machine learning model is adaptive to different datasets.
In addition, among all the machine learning models, the most popular and widely used are conventional algorithms. This is partly due to the barriers between biology and computer science; it is also partly due to the dataset size limitation. In the current data amount context, the traditional machine learning model such as linear models, support vector machine and random forest are still dominant in early cancer detection for their training speed and robustness on small dataset. We hope that the all-sided review of machine learning procedures and corresponding code demos presented in this survey can act as a reference guide. Definitely, advanced machine learning algorithms could also be applied for exploring latent biomarkers and the complicated relationship in order to further improve the performance. However, model generalization and complexity have to be balanced in a fair manner.
As limited with the sample size and the interpretability of deep learning models, deep learning was not popular in liquid biopsy cancer detection. From the related studies in the past several years, we can observe that, with the increased data amount from the liquid biopsy, deep learning methods are likely to outperform conventional machine learning methods. However, there are also concerns. The first concern is that deep learning is vulnerable to overfitting. Therefore, regularization, dropout, and early stopping are utilized to prevent neural networks from overfitting. Besides, the birth of batch normalization improves the model baselines and speeds up all structures
[36]. Due to the variance shift conflict between dropout and batch normalization, these two methods are not recommended to be adopted simultaneously at bottlenecks except for high-dimentional data. Another concern is the black-box nature of deep learning
[37]. Since the hidden layers between input and output layers are complex, it is difficult to extract the most important features and match them with the biological explanation. The explainable framework design is vital to introduce machine learning models into clinical application
[38]. In general, the technique for explaining predictions can be categorized into backpropagation-based methods and perturbation-based methods
[39]. The recent successes of explainable framework
[38][39][40][41][42] do shed light on its promising ability. Therefore, we are still optimistic with its development in cancer detection in the future.
From the perspective of liquid biopsy components, we find out that machine learning is extensively used for single-omics analysis. However, a single type of circulating biomarker seldom fully reveals the essence of tumor occurrence. Therefore, multi-omics detection is another promising direction for early cancer detection and treatment monitoring. The exploration competence of machine learning can enable the capability to figure out the complex causal relationships between different molecular measurements. Therefore, the integration of machine learning methods and multi-omics, including genomics, epigenomics, transcriptomics, proteomics, metabolomics, and microbiomics, provides unprecedented opportunities to understand the underlying mechanism of tumor occurrence and early detection.