Abscisic acid (ABA) is a significant regulator of numerous flexible traits of plant developmental improvements, including embryo maturation, germination, seed dormancy, floral initiation, and root growth.
1. Introduction
Abscisic acid (ABA) is a plant hormone that helps plants respond to drought. Drought-responsive genes may be divided into two categories based on their ABA response: ABA-dependent and ABA-independent genes
[1]. Even though numerous drought-responsive genes are engaged in the ABA signaling system, most drought-induced genes do not react to ABA treatment, indicating the presence of ABA-independent drought-response pathways
[2]. Numerous genes are involved in response to drought and salinity stress; under such conditions, tolerance is triggered by osmotic stress, which liberates ABA
[3]. ABA-dependent and -independent mechanisms control osmotic stress-responsive gene expression
[4]. Plants’ stress response systems, for example, comprise both ABA-dependent and ABA-independent activities.
DREB2A/2B,
AREB1,
RD22BP1, and
MYC/MYB are the TFs that interact with their corresponding cis-acting elements, DRE/CRT, ABRE, and MYCRS/MYBRS, respectively, to regulate the ABA-responsive gene expression
[5].
DFREB1/CBF-type TFs are critical in water and salt stress tolerance in higher plants. These TFs regulate the expression of target genes by binding to
CRT/DRE sites in their promoters. Drought and salt stress, as well as exogenous ABA, stimulated
MbDREB1 expression
[6].
MbDREB1 promoter analysis identified an ABA-responsive element (ABRE) that induced an ICE1-like binding site, two MYB recognition sites, and three stress-inducible GT-1 boxes. ABA, drought, and salt treatments activated GUS activity in transgenic Arabidopsis
[6].
Conversely, both ABA-independent and ABA-dependent stress-induced genes (
COR15a and
rd29B, respectively) were upregulated in Arabidopsis overexpressing
MbDREB1. Both ABA-dependent and ABA-independent pathways used
MbDREB1 to activate plant tolerance to low temperature, drought, and salt stress
[6]. Salt and drought stress induced
PR-1,
PR-5,
RAB-18, and
RD-29A genes in plants pretreated with ABA
[7]. Both ABA-dependent and ABA-independent osmotic stress signaling first adjust constitutively expressed TFs, leading to the expression of early response transcriptional activators, activating downstream stress tolerance effector genes
[8]. TFs play diverse roles in controlling gene expression
[9] in plants, necessary for regulating biological processes, such as development and environmental stress responses
[10]. TFs are the key molecular switches that enable plants to withstand harsh conditions and direct the plant’s developmental process in response to abiotic stress
[11]. TFs play an essential role in crop improvements and are considered good candidates for improving tolerance to various abiotic stressors
[12]. TFs are considered the best genetic materials to breed and develop stress-tolerant crop varieties because of their role as master regulators of many stress response-related genes compared to manipulation in a single functional gene
[13]. For improving abiotic stress resilience in plants, Tripathi et al.
[14] discussed the contributions of new technologies such as DAP-seq, bulk, or single-cell ChIP-seq RNA-seq yeast 1-hybrid and CRISPR/Cas9. ChIP-seq is a method used to analyze protein interactions with DNA. ChIP-seq combines ChIP with massively parallel DNA sequencing to identify the binding sites of DNA-associated proteins
[14]. The chIP or yeast one-hybrid method has a role in identifying co-regulated genes that are strongly differentially expressed in response to the stress treatment and characterization of TFs that regulate many target genes
[14]. A high-throughput TF binding site discovery method using genomic DNA in vitro can quickly identify target genes that directly bind downstream transcription factors. The DAP-seq method is fast, inexpensive, and more easily scaled than the ChIP-seq method
[15]. For example, comparative transcriptomics informed by phylogenetic relationships uncovered lineage and extremophile-specific differences in ABA response. DAP-Seq was utilized to establish GRNs in each species for the entire ABA-AREB/ABF clade
[16]. The stress-inducible CRISPR/Cas9 is a robust, practical, and helpful approach for developing crop varieties resistant to climate change. It will be a helpful tool for capable and particular genome editing in different plants for several traits, including abiotic stresses
[17]. Therefore, some genes are targeted through genome editing based on CRISPR/Cas9 technology in different crops. Another yeast two-hybrid method is a well-established genetics-based system that uses yeast to display binary protein-protein interactions
[18]. Using such techniques, the PYL6 and MYC2 interact, and their interaction is enhanced in the presence of ABA
[19].
Both drought and salinity are among the most severe abiotic factors restricting plant growth and yield. ABA-dependent (drought-inducible genes were clarified upon their induction by exogenous ABA, which means two different systems in stress-inducible gene expression; later, ABA-dependent genes are regulated by endogenous ABA based on mutant analyses) ABA-independent pathways regulate numerous genes that function in drought response. Many signaling molecules, such as ABA, ROS, H2O2, NO, Ca2+, PAs, and others, have been well known and revealed in plant signaling perception and transduction pathways. Many drought- and salinity-responsive genes are involved in ABA-signaling pathways, such as ABA-inducible (ABA-inducible genes were genes induced by exogenous ABA treatment in the early phase of research; now, ABA-inducible genes are regulated by endogenous ABA, as well), ABA-sensitivity (ABA-sensitivity is phenotypes related to ABA sensitive responses during germination, stress response, and so on; various mutants and ecotypes have been reported to be ABA-sensitive ones based on their responses to ABA), and ABA-mediated (ABA-mediated genes are genes regulated by ABA signaling pathways in stress responses and plant growth). However, several drought- and salinity-induced genes do not respond to ABA signaling, showing that ABA-independent signaling pathways also regulate the response to drought and salinity stress.
2. TFs Regulatory Network in Response to Drought and Salinity Stress
A transcription factor is a protein that binds to DNA and regulates gene expression by promoting or suppressing transcription. The function of TFs is to regulate turn on and off genes and ensure their expression in the suitable cell at the right time and the right amount throughout the life of the cell and the organism
[20]. Transcription factors are modular in structure and contain two domains. The first is the DNA-binding domain, which attaches to specific DNA sequences (enhancer or promoter) adjacent to regulated genes. DNA sequences that bind transcription factors are often referred to as response elements
[21]. The second is the activation domain, which contains binding sites for other proteins such as transcription coregulatory. These binding sites are frequently referred to as activation functions, transactivation domains, or trans-activating domains but do not mix with the topologically associating domain
[20][21]. Generally, a stress signal transduction pathway comprises the following key steps: (1) signal perception, (2) signal transduction, and (3) stress response. The first step in activating a signaling cascade for drought and salinity stress recognizes stress signal via receptors located on the membrane of the plant cell
[22]. After recognition, these sensors transmit the signal downstream through phytohormones and second messengers such as Ca
2+ and ROS
[23]. The second messengers, such as ROS, trigger another set of ROS-modulated PKs and PPs, including MAPK cascades, CDPKs, CBLs, CIPK, and many other PKs, as well PPs such as some PP2Cs (
Figure 1)
[22]. ABA is the principal hormone involved in the coordination of abiotic stress in plants
[24]. The different stress tolerance responsive TFs usually function independently. However, there is a possibility that some level of cross-link occurs between these TFs (
Figure 1). The last step is the expression of functional genes involved in different functions such as stomatal closure, oxidative damage, leaf senescence, or indirectly regulating regulatory genes contributing to signaling cascades and transcriptional regulation of gene expression
[22][25]. These abilities allow them to be excellent candidate genes for genetic manipulation of complex stress tolerance traits
[26]. To date, based on genome-wide analysis, a great deal of TFs belonging to different families, such as AP2/ERF, bHLH, bZIP, DREB, GATA, HD-Zip, Homeo-box, MADS-box, MYB, NAC, Tri-helix, WHIRLY, WOX, WRKY, YABBY, and zinc finger, and so on, have been identified in different plant species
[27][28].
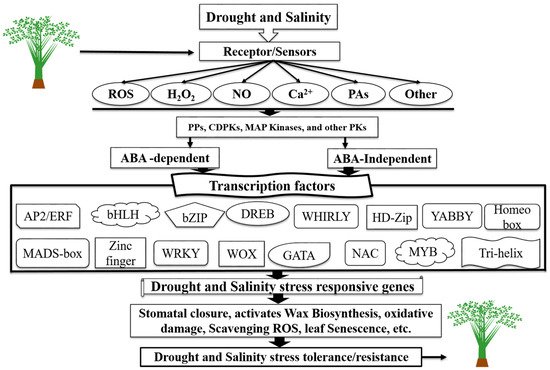
Figure 1. Schematic diagram of TFs as key components in transcriptional regulatory networks during drought and salinity stress-signaling pathways in different crops/plants. A diagrammatic representation of gene expression and drought and salinity stress signal perception in plants via ABA-independent and ABA-dependent pathways (modified from Khan et al.
[22]).