Tuft cells are taste-like chemosensory cells found in the intestinal epithelium involved in the activation of group 2 innate lymphoid cells (ILC2). Although tuft cells in other tissues secrete the neurotransmitter acetylcholine (ACh), their function in the gut remains poorly understood. We investigated changes in the expression of genes and cell differentiation of the intestinal epithelium by stimulation with interleukin-4 (IL-4) or IL-13 in macaque intestinal organoids. This study is the first to demonstrate ACh upregulation by IL-4 induction in primates, suggesting that IL-4 plays a role in Paneth cell granule secretion via paracrine stimulation.
1. Introduction
Intestinal epithelial cells (IECs) exert multiple sensing mechanisms against nutrients, tastants, microorganisms, and chemicals. The tuft cell (or brush cell) is an atypical epithelial cell-type with a characteristic apical tuft of microvilli, and is found in a variety of organs, including the intestine. It is considered to be a taste-like chemosensory cell because these cells express type 2 taste cell markers, such as POU domain class 2 transcription factor 3 (POU2F3), alpha-gustducin (α-gust or GNAT3), and transient receptor potential cation channel subfamily M member 5 (TRPM5)
[1][2]. Recent studies in rodents have shown that intestinal tuft cells activate group 2 innate lymphoid cells (ILC2) by secreting interleukin-25 (IL-25), followed by parasite infection
[3][4][5]. Activated ILC2 secretes Th2 cytokines, such as IL-13 and IL-4, and promotes the hyperplasia of both tuft and goblet cells, leading to the mucous secretion and activation of intestinal motility, known as the “weep and sweep” response
[5]. Other physiological functions of tuft cells in the gut are also proposed for functions such as to sense bitter compounds and succinate that will guide hyperplasia of tuft cells
[6][7][8][9].
Although biosynthetic enzymes of leukotrienes, prostaglandins, and acetylcholine (ACh) have been reported to be expressed in human intestinal epithelial cells
[10], the mechanism by which these molecules are released remains unclear. While tuft cells in the trachea and airways release ACh by noxious or bitter stimuli to exclude these hazardous chemicals
[11], the function of ACh from intestinal tuft cells remains elusive.
In the present study, we investigated the response of the intestinal epithelium to Th2 cytokines using a macaque organoid system. Our data indicate that ACh accumulation in tuft cells correlates with the upregulation of Choline acetyltransferase (ChAT) transcripts triggered by Th2 cytokines. Furthermore, we propose that ACh secreted by tuft cells acts in a paracrine manner to positively regulate granule secretion from Paneth cells.
2.1. Transcriptome Analysis of Macaque Intestinal Organoids after 72 h Culture in Differentiation Medium
We tested whether modified media could be used to culture macaque intestinal organoids. The organoids were maintained with the improved media using IGF-1 and FGF-2 (IF medium) (
Figure 1). As a result, morphological changes were observed compared with cultures using the conventional medium with p38i. We observed that the conventional method using p38i had a lower proliferation potential than the modified media using IGF-1 and FGF-2, especially when organoids were passaged. Next, we examined whether cell differentiation could be induced by differentiation (DI) medium or DI medium with IL-4 (IL-4 medium), as described in the Materials and Methods (
Figure 1a,b). Immunofluorescent studies showed that doublecortin-like kinase 1 (DCLK1) and serotonin (5HT), which are tuft cells and enterochromaffin cell markers [
13,
14,
15], respectively, were rarely detected in IF medium, while the DI medium or IL-4 medium upregulated the expression of both. IL-4 medium had a significant effect on DCLK1 expression (
Figure 1d). As the efficiency of cell differentiation into tuft cells by IL-4 induction was maintained at the same level as in our previous report[
12], the IF conditions were used for subsequent experiments.
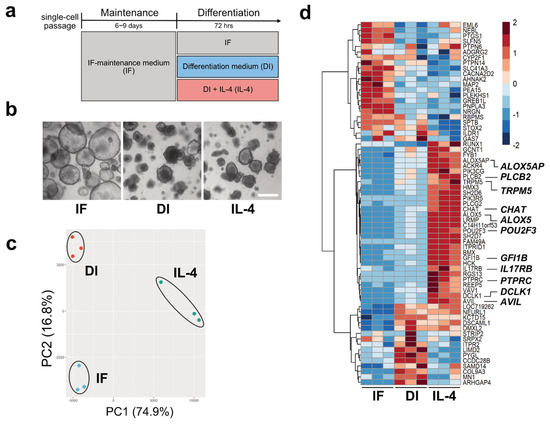
Figure 1. Tuft cell induction and verification. (a). The strategy of inducing tuft cells by changing culture media. (b) Representative images of the intestinal organoids cultured in three different media for 72 h. The morphology of the organoids cultured in differentiation (DI) or interleukin (IL)-4 media changed, becoming more shrunken or budding in shape, while organoids cultured under the IF media maintained a ball-like structure. Scale bar, 500 µm. (c). Principal component analysis (PCA) for the transcriptome profiles of the organoids cultured in control (IF), DI, and IL-4 medium. The scatter plots based on PCA scores of PC1 and PC2 represent the similarity among each biological replicate in three groups. (d). A heatmap of 67 DEGs for the tuft cell in the organoids cultured in IF, DI, or IL-4 medium. Colors indicate the relative expression levels based on row-wise z-score of transformed TPM values.
To observe transcriptional changes under three different culture conditions (IF, DI, and DI with IL-4), we performed RNA-Seq analysis. Principal component analysis (PCA) showed that biological replicates from the same culture conditions fell into the same cluster, indicating that clusters represented distinct groups (Figure 1c).
Using volcano plot analysis, we identified 1197 genes that were significantly upregulated and 734 genes that were downregulated by IL-4 (FC ≥ |1.5| and FDR ≤ 0.01). The representative genes within the upregulated genes were Mucin 2 (MUC2), Trefoil Factor 3 (TFF3), Defensin Alpha 6 (DEFA6), Lysozyme (LYZ), Arachidonate 5-Lipoxygenase (ALOX5), and CHAT. Leucine-rich repeat G-protein-coupled receptor (LGR5) was identified as one of the downregulated genes. Next, to gain further insights into the transcriptional changes in each group, transcriptome data were subjected to heatmap analysis. We compared the gene expression patterns of 88 genes reported as tuft cell markers [
16]. As a result, 30 genes, including POU2F3, TRPM5, DCLK1, Phospholipase C-beta-2 (PLCb2), Phospholipase C-gamma-2 (PLCg2), Interleukin-17 receptor B (IL17RB), ALOX5, and CHAT, were found to be abundantly expressed under IL-4-induced conditions compared with IF or DI conditions (
Figure 1d). In contrast, we failed to detect GNAT3 and IL-25, which are tuft cell markers in mouse [
17], owing to their low transcript levels.
To substantiate transcriptional regulation by Th2 cytokines, IL-13 was also added to the culture medium, and the expression of tuft cell marker proteins and transcripts was analyzed by immunohistochemistry and RNA-Seq. The results showed that both protein and gene expression patterns were similar to those stimulated by IL-4, indicating that IL-13 also induces tuft cell differentiation.
2.2. Confirmation of RNA-Seq Results by Semi-Quantitative RT-PCR
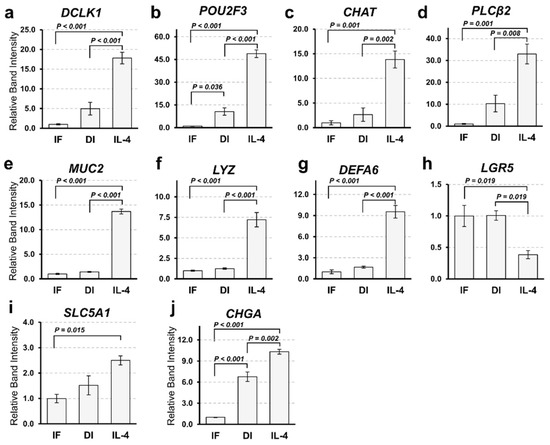
To confirm the results of RNA-Seq, we performed a semi-quantitative reverse transcription polymerase chain reaction (RT-PCR). The expression patterns of tuft cell markers (DCLK1, POU2F3, CHAT, and PLCb2) were comparable to the results obtained from RNA-Seq (Figure 2a–d). The expression of a goblet cell marker (MUC2) and Paneth cell markers (LYZ and DEFA6) was significantly increased by IL-4 induction, while that of a stem cell marker (LGR5) was decreased (Figure 2e–h). MUC2- and DEFA6-positive cells were also increased in organoids induced by IL-4. However, we failed to detect any change in the expression of the enterocyte marker (Solute Carrier Family 5 Member 1; SLC5A1) between the DI and IL-4 groups (Figure 2i). Although the expression of enteroendocrine cell marker (Chromogranin A; CHGA) appeared to increase in the IL-4 group (Figure 2j), this change was marginal in the RNA-Seq analysis (data not shown).
Figure 2. Gene expression analysis of intestinal markers by semi-quantitative reverse transcription polymerase chain reaction (RT-PCR). (a–j). Comparison of the expression marker genes under three different culture conditions. The mRNA expression of organoids cultured in IF, DI, or IL-4 media was subjected to semi-quantitative analysis. Marker genes of tuft cells (a–c), goblet cells (e), Paneth cells (f–g), intestinal stem cells (h), enterocytes (i), and enteroendocrine cells (j) were amplified by RT-PCR using gene specific primers. The expression level of each gene was examined by band intensity after electrophoresis. Data are shown as means ± SEM (N = 3); p-values are shown in each figure, Tukey’s honest significant difference (HSD), analysis of variance (ANOVA) p < 0.001 (a–h), p = 0.016 (i), and p = 0.012 (j).
2.3. ChAT and Ach Are Upregulated by IL-4
RNA-Seq analysis revealed that
ChAT expression was upregulated when induced by IL-4 (
Figure 3a). Immunohistochemical analysis using macaque intestinal tissues and organoid cultures showed that ChAT and DCLK1 were co-expressed in tuft cells (
Figure 3b). ChAT immunoreactivity was not observed in the 5HT-positive enteroendocrine cells. As ChAT is a key enzyme responsible for the biosynthesis of ACh
[12], we speculated that macaque organoids synthesize ACh. Therefore, we attempted to quantify ACh in organoids by HPLC/MS analysis (
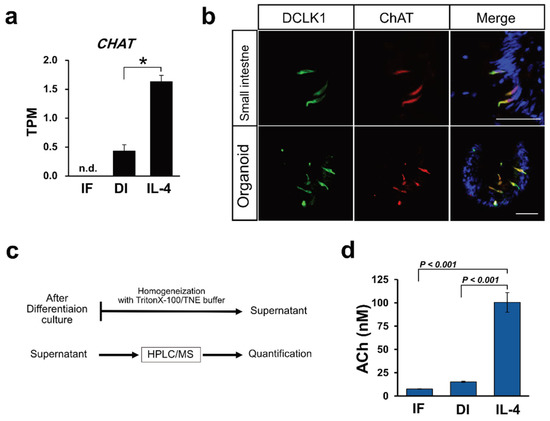
Figure 3c). The ACh concentration was evaluated from the standard curve of d4-ACh, as described in the Materials and Methods. From the standard curve, the ACh concentration range was 2.27–183.50 nM (0.4 to 33.3 ppb of ACh-Cl). We also measured the ACh concentration of cytosolic samples prepared from the organoids of control (IF), differentiation (DI), or IL-4 induced (IL-4) conditions, and found that the average concentration of ACh in each culture condition was 7.58, 15.18, and 100.45 nM, respectively. The average value (
N = 3) of ACh concentration was statistically analyzed, and a bar graph was generated to show that ACh was significantly up-regulated in IL-4-induced conditions compared with the other conditions (
Figure 3d).
Figure 3. Acetylcholine production correlates with tuft cell differentiation. (a) CHAT mRNAs were induced by IL-4 induction. Data are shown as means ± SEM (N = 3); * adjusted p-value < 0.0001 vs. DI group, R package “DESeq2”. (b) Immunohistochemical analysis of tissues from the small intestine and from macaque organoids show that DCLK1 positive cells (green) and ChAT positive cells (red) were colocalized. Nuclei were stained with DAPI. Scale bar, 50 µm. (c). A schematic flow chart of ACh quantification. (d) Quantification of Ach in the organoids cultured in IF, DI, or IL-4 medium. Samples obtained from the organoids were analyzed by HPLC/MS and ACh level in each sample was calculated. Data are shown as means ± SEM (N = 3); p-values are shown in each figure, Tukey’s HSD.
2.4. Granule Secretion Induced by ACh in Organoid Cultures
We observed ACh accumulation in the intestinal organoids of macaques when tuft cell differentiation was induced by IL-4. A previous study showed that Paneth cells express muscarinic ACh receptors
[13]. Upon carbachol stimulation, Paneth cells release antimicrobial substances
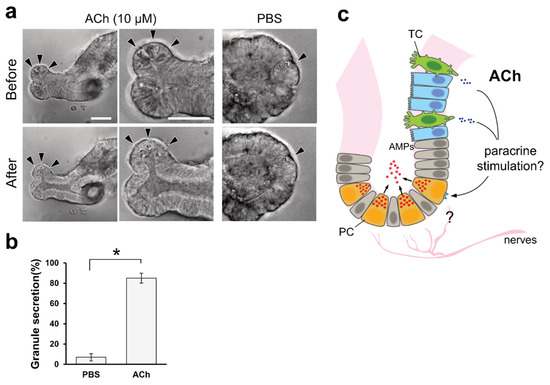
[14]. As carbachol is a synthetic agonist of the cholinergic receptor, we speculated that ACh could also stimulate granule secretion from Paneth cells. Thus, we investigated the effect of ACh on Paneth cells using mouse organoids with distinguishable vesicles in Paneth cells. Upon the basolateral stimulation of mouse intestinal organoids with 10 µM ACh, we detected the release of vesicular contents from Paneth cells (
Figure 4a). This phenomenon lasted for 5 min, until the area of granules in Paneth cells was significantly decreased compared with the control PBS stimulation (
Figure 4b).
Figure 4. ACh induces granular secretion from Paneth cells in mouse intestinal organoids. (a). Representative images of Paneth cells (arrowheads) before and after 5 min of 10 µM ACh stimulation. PBS was used as a negative control. Granules in the apical part of the Paneth cells were released inside the organoid upon ACh stimuli, and very few granules were observed. Scale bar, 100 µm. (b). Granular secretion was statistically measured by calculating the area of granules, as described in the Materials and Methods. The secretion ratio was described as the ratio of the post-stimulation area to the pre-stimulation granule area. * p < 0.001, Welch’s t-test. (c). Hypothetical model derived from this study. ACh is basolaterally secreted from tuft cells and acts as a paracrine stimulus on Paneth cells to secrete granules that contain antimicrobial peptides in the intestine. TC, PC, and AMPs represent tuft cells, Paneth cells, and antimicrobial peptides, respectively.