1000/1000
Hot
Most Recent

Fibroblast growth factors (FGFs) comprise a large family of growth factors, regulating diverse biological processes including cell proliferation, migration, and differentiation. Each FGF binds to a set of FGF receptors to initiate certain intracellular signaling molecules. Accumulated evidence suggests that in early development and adult state of vertebrates, FGFs also play exclusive and context dependent roles.
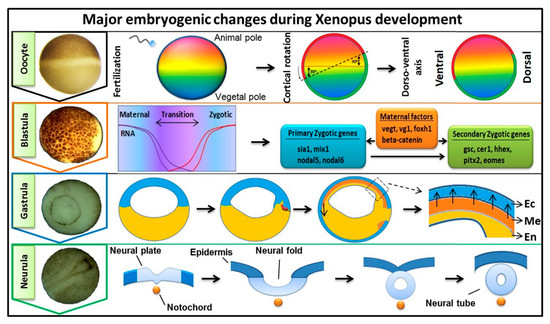
Early embryogenesis in vertebrate embryos involves the irreversible developmental process. As the ovum receives the male haploid genome from a sperm to become a diploid cell, the process of fertilization is started with the fertilized ovum then being referred to as a zygote. The zygote goes through several key developmental stages including mid-blastula transition (MBT), gastrulation (germ layer formation), and neurula, for establishing the overall body axis and generating the anterior CNS and posterior PNS ( Figure 1 ). These are tightly controlled spatiotemporal events led by several signaling pathways and occur in conjunction with maternal or zygotic morphogen gradients throughout the embryos. FGF signaling is known to play an essential role during embryonic development [1][2][3], and in this review, our discussion was focused on involvement of FGF signaling in early embryogenesis. The first discovered FGF ligand, FGF2 (also known as basic FGF/bFGF), was purified from brain tissue in 1975 and was defined for its stimulatory activity in fibroblasts [4]. Since then, a total of 22 FGF members have been identified in humans and similar numbers in vertebrates. Except for the intracellular FGF11 subfamily, these interact with particular FGF receptors (FGFRs) to activate intracellular effector proteins. FGF/FGFR signaling regulates a plethora of cellular processes mediated by activation/modification of cytosolic effectors and followed by transcriptional regulation of target genes. Dysregulation of FGF signaling has been reported to promote several human diseases and disorders, and their severity can vary based on active ligands/receptors and tissues involved [5]. Several lines of evidence also indicate a crucial role for FGF signaling in early embryogenesis for germ layer formation [1][2] and organogenesis [6]. During primordial germ layer specification, FGFs modulate fate determination as autocrine and paracrine signaling agents. Alterations in tightly regulated FGF expression patterns, as with altered FGFR splicing or mutation and changes in spatiotemporal FGF-FGFR interactions, may result in flawed and defective development for multiple congenital diseases and the onset of various cancer types [5][7][8]. In FGF/FGFR genes, genetic mutations that lead to several congenital diseases have been described reviewed elsewhere [9]. In this review, we summarized the information and our understanding of the functional role(s) of FGFs in early embryonic germ layer specification and axis formation during embryogenesis.
The FGFs are a large family of growth factors consisting of 22 members in humans and mice, and 19 members identified in Xenopus ( Table 1 ). In this family, there are seven subfamilies described in vertebrates, namely FGF1, FGF4, FGF7, FGF8, FGF9, FGF11, and FGF19 (reviewed [10][11]). Subfamily members have high similarity in their amino acids sequences. Except for the intracellular FGF11 subfamily, extracellular secretion of a given FGF is required for its signaling and function. Based on their secretion profile, FGF members can also be placed into two groups. The first are those whose secretion takes place through a classic endoplasmic reticulum-Golgi secretion pathway in the cells as these FGFs contain N -terminal hydrophobic peptides and they include FGF3, 4, 5, 6, 7, 8, 10, 17, 18, 19, 21, and 23 [10]. The second group of FGFs are those that do not contain N -terminal hydrophobic peptides and are endoplasmic reticulum-Golgi-independent for their secretion. These include FGF1, 2, 9, 16, and 20 [10][11][12]. As an exception, FGF22 remains attached to the cell surface by its N -terminal signal peptide rather than being secreted (reviewed [10]). All members of FGF11 subfamily ( Table 1 ) are known as non-secretory FGFs and are strictly intracellular proteins. Even though these FGFs share structural homologies with other secreted FGFs, they do not share any functional similarities [13][14]. As intracellular entities, FGF11 subfamily members have been documented as being components of certain protein kinase-mediated signaling pathways and they also interact with membrane channels to regulate cell fate [15][16].
| Human | Mouse | Xenopus | ||||
|---|---|---|---|---|---|---|
| Subfamily | Members (Other Name) |
Ref. | Members (Other Name) |
Ref. | Members (Other Name) |
Ref. |
| FGF1 | FGF1 (aFGF) | [17] | FGF1 (FGFa) | [18] | xFGF1 | [19] |
| FGF2 (bFGF/FGF-β) | [4] | FGF2 | [20] | xFGF2 (bFGF) | [21] | |
| FGF4 | FGF4 (eFGF) | [22] | FGF4 (KFGF) | [23] | xFGF4 (eFGF, fgf4-a, fgf4-b) | [24] |
| FGF5 | [25] | FGF5 | [26] | xFGF5 | [19][27] | |
| FGF6 (HST2, HSTF2) | [28] | FGF6 (HSTF2) | [29] | xFGF6 | [19][27] | |
| FGF7 | FGF3 | [30] | FGF3 (Int-2) | [31] | xFGF3 (INT-2, FGF3A) | [32] |
| FGF7 (KGF) | [33] | FGF7 (KGF) | [34] | * | ||
| FGF10 | [35] | FGF10 | [36] | xFGF10 | [37] | |
| FGF22 (UNQ2500/PRO5800) | [38] | FGF22 | [38] | xFGF22 | [27] | |
| FGF8 | FGF8 (AIGF) | [39] | FGF8 (AIGF) | [40] | xFGF8 (FGF8a, FGF8b) | [41] |
| FGF17 (UNQ161/PRO187) | [42] | FGF17 | [42] | * | ||
| FGF18 (UNQ420/PRO856) | [43] | FGF18 | [43] | * | ||
| FGF9 | FGF9 | [44] | FGF9 | [45] | xFGF9 (GAF, HBGF9) | [46] |
| FGF16 | [47] | FGF16 | [48] | xFGF16 | [27] | |
| FGF20 | [49] | FGF20 | [50] | xFGF20 | [51] | |
| FGF11 | FGF11 (FHF3) | [52] | FGF11 (FHF3) | [52] | xFGF11 | [27] |
| FGF12 (FGF12B, FHF1) | [52] | FGF12 (FHF1) | [53] | xFGF12 | [54] | |
| FGF13 (FHF2) | [55] | FGF13 (FHF2) | [53] | xFGF13 | [56] | |
| FGF14 (FHF4) | [57] | FGF14 (FHF4) | [58] | xFGF14 (FHF4) | [19] | |
| FGF19 | * | FGF15 | [59] | * | ||
| FGF19 (UNQ334/PRO533) | [60] | * | xFGF19 | [27] | ||
| FGF21 (UNQ3115/PRO10196) | [61] | FGF21 | [61] | * | ||
| FGF23 (HYPF, UNQ3027/PRO9828) | [62] | FGF23 | [62] | xFGF23 (fgf23.1, FGF23.2) | [27] | |
| FGF receptor family | ||||||
| FGFR1 | [17] | FGFR1 | [63] | xFGFR1 | [64] | |
| FGFR2 | [65] | FGFR2 | [66] | xFGFR2 | [67] | |
| FGFR3 | [68] | FGFR3 | [69] | xFGFR3 | [70] | |
| FGFR4 | [71] | FGFR4 | [72] | xFGFR4 | [73] | |
FGFs interact with specific FGF receptors to initiate intracellular signaling and there are four cell membrane tyrosine kinase FGF receptors, FGFR1, 2, 3 and 4, which are members of the larger receptor tyrosine kinase (RTK) group [74][75]. Each FGFR is a single-pass transmembrane (TM) protein that includes an N -terminal extracellular ligand FGF binding domain and a C -terminal intracellular tyrosine kinase domain. The extracellular domain contains 3 immunoglobulin-like subdomains (D1, D2, and D3 domains) [76]. There is an also an acidic box between D1 and D2 domains. The D2 and D3 domains facilitate FGF binding [76]. Heparan sulfate (HS) is a coreceptor for FGF binding to an FGFR, and it is essential for FGF binding and signaling. HS is one of the abundant polysaccharides found in the extracellular matrix of mammalian cells [77], and it interacts with the cationic patch found in both FGF and D2 subdomain of FGFR [77]. Ligand binding induces a conformational change in FGFR, leading to its dimerization and activation of the its intracellular kinase [78]. For FGFR1, FGFR2, and FGFR3, two standard isoforms (b-and c-isoforms) are generated by splicing [79]. The splicing variants have altered ligand affinity for various FGFs, except for FGF1, acknowledged as a universal ligand and being able to interact with both FGFR isoforms [74][77].
In this section, we briefly summarized the activation modes of FGFR associated cytosolic effectors and linked components that act as intermediates. Several partner receptor proteins may be associated with the cytosolic domain of an FGFR, such as cell adhesion molecules (CAMs) and G-protein coupled receptors (GPRCs) [80]. Signal-induced activation of FGFR typically activates multiple cytosolic signaling pathways. An FGFR is mainly associated with its intracellular signaling intermediates, including phospholipase C (PLCγ), FRS1, FRS2/FRS2α, and FRS3/FRS2β (reviewed [81]). FRS2 recruits the adaptor GRB2 (growth factor receptor-bound 2) [82], and once GRB2 is bound to the functional domain of the FRS2, it can interact with either SOS or GAB1 and form a complex [83]. Upon FGF ligand binding to FGFR and heparan sulfate, multiple cytosolic events occur; these are mostly activational phosphorylations. Once the FGFR complex is activated, GRB2/SOS exchanges the GDP to GTP for Ras; GTP-Ras in turn activates and stimulates Raf (also known as MAPK kinase kinase) as part of the Ras/MAPK pathway (for a detailed mechanism, refer to reviews [84][85][86][87]). Similarly, GRB2 switches on PI3K/Akt signaling cascade, as activation of PLCγ/PKC and JAK/STAT pathways are also directly linked to FGFR activation [84][85][86][87]. These pathways generate the signals leading to targeted transcription factors regulating transcription of their target genes.
Known FGFs and FGFRs in human, mice, and Xenopus.
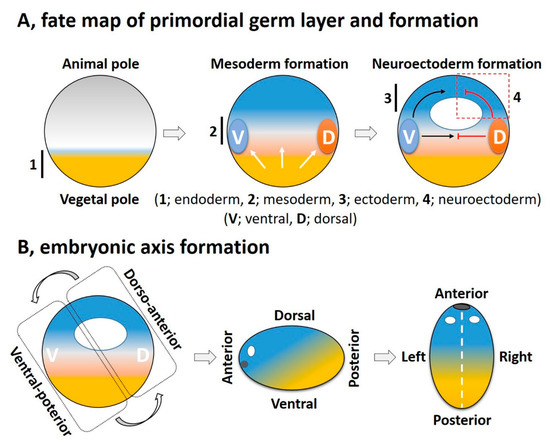
Cellular diversification is an essential process in generating a complex, multicellular organism. This process begins with gastrulation as the three germ layers of endoderm, mesoderm, and ectoderm are specified. For an amphibian embryo, the first embryonic germ to specify is the endoderm from the vegetal pole. The formation of mesoderm, the second germ layer, is the result of active mesoderm inducing signals, originating from the vegetal region or the prospective endoderm, which triggers the mesoderm specification. During the mesoderm specification, generally two distinct signaling centers, namely the ventral and dorsal signaling centers, are established in the marginal region; these are based on presence and activation of maternal or zygotic factors at ventral and dorsal half of the embryo ( Figure 2 A). Once the ventral and dorsal signaling centers are established, they both collectively drive further germ layer specification. The dorsal centers (also called the dorsal mesoderm, dorsal organizer, and organizer) produce dorsal mesoderm and promote neuroectoderm formation by inhibiting ventral signaling. On the opposite end, the ventral center provides the ventral mesoderm and the ectoderm (for epidermis formation) ( Figure 2 A). In this section, we try to summarize primordial germ layer formation and the active role of FGF signaling in this process.
The active molecules with mesoderm inductive activity from the vegetal hemisphere include members of Xenopus nodal-related factors (Xnr1, 2, 4, 5, and 6) and TGFβ family (activin βB), now known as endogenous mesoderm inducing factors in various animal models, including Xenopus, zebrafish, and mouse [88][89][90][91] and the signaling pathways utilized by these signals are often due to activin ligands [88][89][90][91]. This mesoderm inducing effect by activin treatment in metazoan embryogenesis is now widely accepted. The ectopic expression of activin/Nodal downstream intracellular effectors Smad2/3 activate xbra , chordin (chrd) , noggin (nog) , and goosecoid (gsc) expression [92]. Activin-mediated mesoderm induction critically depends on FGF signaling, as demonstrated when DNFR led to complete loss of mesodermal genes expression due to activin [93]. In comparison, ectopic expression of Smad2 significantly elevates expression of various FGFs and FGFRs including fgf3 , fgf8 , fgf20 , fgfr1 , and fgfr2 [94]. Supporting the role of FGFR signaling in mesoderm induction is also observed with the expression profile of several FGF ligands in early mesoderm of vertebrate embryos. Fgf3 expression is primarily in the ring around blastopore lip (mesoderm) of Xenopus gastrula, which then resolves into neural or brain tissues at a later stage of development [95]. Similarly, fgf4 , fgf8 , and fgf20 are largely expressed in early mesoderm or late mesodermal lineages [96][97][98][99]. However, integration of FGF and activin/nodal signaling in mesoderm specification is a potential subject for a future investigation.
Brachyury (Bra/Xbra) is a T-box transcription factor and signature mesodermal (also known as a pan-mesodermal marker) factor, and is actively involved in gene regulation process required for mesoderm induction and differentiation [100][101]. Similarly, Eomesodermin (Eomes) is another T-box transcription factor, an important mesodermal factor crucial for mesoderm induction and differentiation [102]. Both proteins are necessary for mesoderm induction; however, Xbra is also involved in expression of ventral/posterior specific genes (e.g., ventx1.1 and xhox3 ) [100][103][104][105]. Xbra is indispensable for mesoderm formation and maintenance since knockdown of Xbra converts its mesodermalizing character to a neuralizing one [106][107]. Eomes has been reported to induce dorsal/anterior mesodermal ( gsc , chrd , and nog ) genes in Xenopus embryos [102]. Xbra early expression was reported to depend on FGF signaling as embryos treated with SU5402, showed no xbra expression [97]. In turn, FGF2 and FGF4 have been reported to induce xbra transcription in animal explants of Xenopus embryos, wherein Xbra makes an autocatalytic regulatory loop to maintain the Fgf4 expression [106][108]. Xbra can maintain the expression fgf4 , but its activation is most likely thought to depend on Nodal signaling; this is similar to fgf8 activation largely depending on nodal/activin signaling [97]. FGF8, have two alternative splice variants (protein isoforms), namely FGF8a and FGF8b, and these show different inductive features. FGF8a has been reported to have mostly neural inducing activity with little mesoderm inducing capability, whereas FGF8b is important for mesoderm induction and differentiation [41].
A spatiotemporal expression of FGF ligands and receptors is seen in Xenopus embryos, suggesting diverse expression patterns among ligands. During gastrulation, fgf1 , fgf2 , fgf4 , fgf8 , fgf20 , and fgf22 are highly expressed, implying a requirement for mesoderm specification and extension [19]. Several transcription factors have been reported to incorporate FGF signaling in mesoderm specification and embryonic patterning. As an example, mef2d is transcriptionally activated by FGF4 and FGF8 in the marginal region of Xenopus embryos, and from the vegetal region, nodal (Xnr5 and 6) also induces mef2d expression. Once Mef2d protein is made, it can form a positive expression feedback loop with FGF4 and FGF8, leading to the expression of mesodermal genes like xbra , chrd , gsc , and nog [109]. A similar example of interaction with FGF signaling is for Egr1, which has been reported as a downstream target of FGF signaling and plays an important role in embryonic development. The Egr1 activates the transcription of myod , and represses xbra transcription [110]. Collectively, the presented evidence suggests that FGF signaling may interact with several other signaling pathways (e.g., activin, nodal, BMP, and RAR) during mesodermal differentiation in a context-dependent manner (for example in the ventral and dorsal mesoderm). However, the details for the different ligands, their receptors, and their mechanistic integration with other key signaling entities remain to be fully elucidated.
Several agents have been discovered or experimentally developed that are relatively specific in modulating FGF signaling, and chemically, these include proteins and small molecules. Based on their activities, these modulators may be placed into three groups: The first includes the endogenous activator(s) as those agents having a positive or compensatory activity relative to FGF signaling. Second are the endogenous factors showing inhibitory or negative activity towards FGF signaling. Third includes the group of relatively small molecular-weight compounds, chemically synthesized to act as inhibitors and with use intended mostly for therapeutic or research applications. Prominently known endogenous activators/inhibitors of the FGF pathway were chosen to be briefly discussed.
A positive feedback regulatory mechanism plays an important role in embryonic and post embryonic development in vertebrates. In this process, several modulators may act as co-activators, interacting with a specific domain of an FGF or an FGFR; this interaction can in turn significantly accelerate or amplify FGF signaling. Notably, heparan sulfate proteoglycans (HSPGs) and glypicans are essential extracellular key modulators, which interact with a variety of growth factors including various FGFs and morphogens and their receptors [111]. These interactions can be positive as well as negative on target signaling with fine control being exerted in a spatiotemporal manner. Target signaling may include that for Wnt, Shh, and FGF pathways. Currently, several FGF positive modulators have been identified that include anosmin-1 (An1), Sef1, L1CAM, and FLRT3, and can alter FGF signaling at multiple levels (for more details, we refer the reader to the selected articles [111][112][113][114]) ( Table 2 ). Certain small molecules have also been reported to induce FGF signaling. For example, 8-hydroxyquinoline sulfate and pyrithione zinc increases the expression of FGF target genes in zebrafish embryos [113]. However, as these reports provide a thoughtful explanation and clues to the regulatory mechanisms in FGF signaling, detailed mechanisms on the control of FGF signaling in development remain to be fully understood.
| Modulators | Target | Effect on FGF Signaling |
|---|---|---|
| Anosmin-1 | Extracellular domain of FGFR | Positive |
| L1CAM | Extracellular domain of FGFR | Positive |
| HSPGs | FGFs or extracellular domain of FGFR | Positive |
| FLRT3 | Intracellular domain of FGFR | Positive |
| Sprouty | Grb/Raf | Negative |
| Sef’s/Spred/Pyst1/Dusp’s | Modification of several kinases activity like (MAPK, MEK, ERK, and PKB/Akt) | Negative |
A negative feedback mechanism is an essential regulatory strategy for nearly all signaling pathways and this is particularly important during embryonic development. FGF signaling has demonstrated as having an active functional role at multiple steps across embryonic development. To date, a group of endogenous proteins have been identified that show antagonist activity to FGF signals and are generally associated with the negative feedback loop in downregulating FGF signal transduction. One is Sprouty, an intracellular protein that interacts with Drk and Gap1, being components of Ras/MAPK pathway, and blocks their interaction [111][112]. Another is Sef, with a similar expression to various fgf genes, which is capable of interacting with the cytosolic domain of FGFR and blocking the intracellular effectors interacting with FGFR. An ectopic gain of function of sef , for example, has been shown to reduce phosphorylation of Raf1 and MEK1/2 [113]. Other agents that have been reported are Spred [114], Pyst1/Mkp3 [111], Dusp1 [94], and FRS2α/β [115], all endogenous FGF inhibitors/modulators. All of the above mentioned blockers of FGF signaling may have separate or overlapping targets (see Table 2 ) and are generally cytosolic effector proteins.
FGF modulators are essential in early embryogenesis as several studies have reported knockdown of various HSPG genes causing defective development in vertebrates and invertebrates (reviewed [116]). The initial reports from a vertebrate model, namely mouse, has the mutation of “lazy mesoderm” ( lzme ) gene where lzme encodes an enzyme required for glycosaminoglycan biosynthesis) causing defective mesoderm and endoderm migration [117]. Gene expression profiles demonstrated that FGF target genes expression are severely compromised in lzme mutant embryos, whereas the nodal and Wnt3 pathways remained normal [117]. Similar findings have been reported in Xenopus embryos, wherein morpholino-based knockdown of gpc4 (Glypican 4) shows pleiotropic developmental aberrant phenotypes including defected primary axis and forebrain patterning [118]. A detailed discussion of FGF modulators, however, is beyond the focus of current review, and we refer the readers to the related articles [116][117][118][119][120][121].