The Dead-End (DND1) protein can interact with different messenger RNAs (mRNAs) in the cell. It uses multiple mechanisms to regulate expression of proteins from their cognate mRNAs. High levels of DND1 are found in the progenitor cells that develop into the egg and sperm. Defects in DND1 can cause tumors in the testes and ovaries of vertebrates. DND1 may also participate in human cancer development in cells other than those of the testesand ovaries.
1. Introduction
The story of Dead-End 1 (DND1) in cancer of mouse germ cells can be traced back almost half a century. A new spontaneously arising mutation led to generation of the new inbred subline of mice (129/Sv-
Ter) that exhibited primordial germ cell (PGC) deficiency, accompanied with a high incidence of congenital testicular germ cell tumors (TGCTs) or teratomas
[1][2][3]. These recessive phenotypes mapped to mouse chromosome 18, or the
Ter locus
[4][5]. In 2005, twenty years after the 129/Sv-
Ter mouse line was isolated, positional cloning of
Ter identified the genetic defect to a mutation in
Dead-End 1, a mouse ortholog of the zebrafish
dead-end gene required for PGC migration and survival
[6]. Mouse
Dnd1 contains four exons.
Ter is due to a single nucleotide substitution occurring spontaneously in exon 3 of
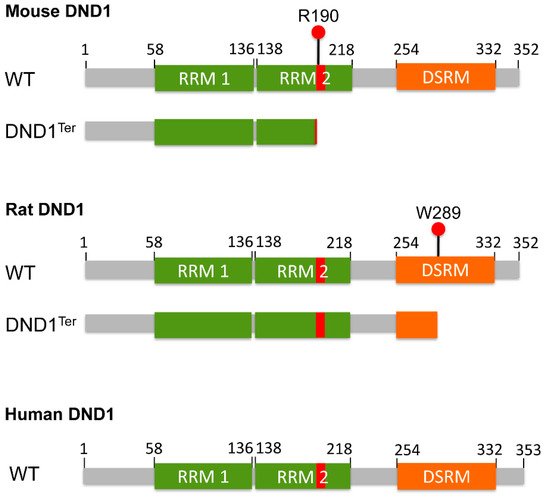
Dnd1, which transforms the arginine residue at amino acid (aa) 190 into a premature stop codon, thus causing either truncation or loss of DND1 expression (
Figure 1). This mutation results in the phenotypes of PGC loss, male and female sterility, and the high incidence of TGCT in males of the 129/Sv mice.
Figure 1. Schematic representation of the structure of mouse, rat, and human DND1 proteins. The RNA recognition motif 1 (RRM1), RNA recognition motif 2 (RRM2) and double stranded RNA-binding motif (DSRM) are shown. Ter mutation transforms arginine at amino acid (aa) 190 in mouse and tryptophan at aa 289 in rat DND1 into a premature stop codon. The red bar in RRM2 represents the HRAAAMA motif spanning from aa 189 to 195 in all three species. The predicated mouse and rat DND1Ter protein are also shown underneath the wild-type (WT) protein, respectively. Please note that whether DND1Ter causes truncation or loss of DND1 expression is still controversial. The NCBI Reference Sequence numbers for mouse, rat and human DND1 proteins are NP_775559.2, NP_001102849.1 and NP_919225.1, respectively, accessed on 20 July 2021.
DND1 is an RNA-binding protein that contains two RNA recognition motifs (RRMs) in tandem, spanning approximately aa residues 58–136 and 138–218 (
https://www.uniprot.org/uniprot/Q6VY05, accessed on 18 July 2021), respectively, followed by a double stranded RNA-binding motif at the carboxyl (C)-terminus (
Figure 1). The amino (N)-terminal RRM1 is canonical and essential for binding specific mRNAs
[7][8]. The RRM2 is less conserved and contains the HRAAAMA motif (
Figure 1, described below). Over the years, DND1 has been found to possess diverse molecular functions. The most studied is its role in translation regulation
[7][8][9][10]. In addition to TGCTs, DND1 has also been shown to potentially impact ovarian teratomas (OTs) and somatic tumors
[11][12][13][14][15][16][17][18][19]. Because DND1 has been described as a protein primarily expressed in germ cells, the multiple research reports and database evidence of DND1 involvement in somatic cancers are unexpected.
2. Molecular Mechanisms of DND1 Function
2.1. mRNA Stabilization
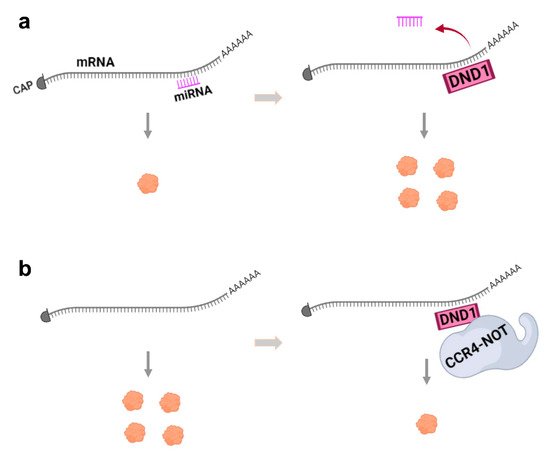
Several studies have demonstrated that DND1 can bind specific mRNAs and block microRNA (miRNA) access from the 3′-untranslated region (3′-UTR) of target mRNAs and inhibit miRNA-mediated mRNA degradation, thus up-regulating translation (
Figure 2a). The target mRNAs of DND1 that have been studied are often those involved in regulation of the cell cycle or apoptosis. For example, DND1 binds to the 3′-UTR of
p27 mRNA and blocks miR-221 so as to up-regulate p27 protein expression
[7]. Increase in the level of this cyclin-dependent kinase inhibitor is well known to decrease cell proliferation and induce apoptosis
[20]. Uridine-rich regions (URRs) on the 3′-UTR of the
p27 transcript mediate the DND1-3′-UTR interaction, as mutating the URRs impaired DND1-mediated up-regulation of
p27 translation
[7]. In another example, zebrafish Dnd1 protein binds the URR in the 3′-UTR of the mRNA of the DNA replication inhibitor,
geminin, and increases its translation
[21]. In addition to directly regulating cell cycle via stabilizing
p27 or
geminin, DND1 can also impose indirect regulation of the cell cycle. For example, through blocking miR-26a from
enhancer
zeste
homolog
2 (
EZH2) mRNA that encodes a methyltransferase responsible for histone H3 lysine 27 (H3K27) trimethylation (me3), DND1 up-regulates the translation of
Ezh2 [22]. In turn, EZH2 protein represses cyclin
CCND1 expression to inhibit entry into the cell cycle.
Figure 2. The two major mechanisms of DND1 function. (a) Interaction of DND1 with mRNA blocks miRNA access to mRNA to up-regulate translation and protein expression. (b) DND1 can also bind to mRNA to target it for destruction by the CCR4-NOT complex, thus decreasing translation and protein expression. These mechanisms may be cell-type- or mRNA-specific.
During
Xenopus embryogenesis, Dnd1 has been reported to bind to the 3′-UTR of
trim36, which is essential for vegetal cortical microtubule assembly, promoting localized high concentration of Trim36 protein in the vegetal cortex in the
Xenopus egg. Thus, Dnd1 also appears to have an essential role in vegetal cortical microtubule assembly during axis specification during embryogenesis
[23].
DND1 function of blocking miRNA access is also subject to negative regulation or DND1 itself is subject to miRNA regulation. For example,
Xenopus dnd1 mRNA can be cleared by
miR-18 in the soma and specifically protected from degradation in germ line of
Xenopus embryos via Elr-type proteins
[24]. Another example in squamous cell carcinoma cell lines demonstrated that miR-24 down-regulates DND1
[13]. The DNA cytosine deaminase,
apoli
poprotein
B mRNA-
editing enzyme,
catalytic polypeptide-like
3 (APOBEC3), was found to interact with DND1
[25] and counteract DND1 function by restoring the suppressive activity of miRNA on
p27,
LATS2 and
CX43 mRNAs
[26].
A recent study dissected the individual functions of the two RRMs of Dnd1 in regulating
nanos1 translation in
Xenopus germline
[8]. The N-terminal RRM1 is critical for interaction with mRNA targets
[7][8][10]. The RRM2 possesses ATP-dependent helicase activity
[8][27] required to resolve the secondary structure of bound mRNAs to promote translation
[8]. The above-mentioned HRAAAMA motif is presumably part of the putative ATPase domain. Furthermore, CRISPR-Cas9-mediated base-editing generated mutations of four aas in RRM1 of mouse DND1 (E59, V60, P76 and G82), which functions in mRNA binding, depleted PGCs in mouse embryos, indicating the essential role of these RRM1 aas for DND1 function during PGC development
[28].
2.2. mRNA Repression or Degradation
Other studies show that DND1 can also function as a translation suppressor
[9][10][29]. It binds a UU(A/U) trinucleotide motif predominantly in the 3′-UTR regions of mRNA and in addition recruits the CCR4-NOT deadenylase complex, to degrade target mRNAs (
Figure 2b). The mRNA targets of DND1 destabilized by CCR4-CNOT include genes associated with apoptosis, inflammation, and pathways that regulate stem cells, such as of the TGF, WNT and PI3K-AKT signaling pathways
[10]. In germ cells, DND1 has also been shown to partner with NANOS2
[9][30][31], an evolutionarily conserved RNA-binding protein specifically involved in germ cell development, and then interact with CCR4-NOT. DND1 is required for loading unique RNAs into the Nanos2-CCR4-NOT complex
[9], likely accounting for the synergistic increase of incidence of TGCT in double mutants for DND1 and NANOS2
[31]. On the other hand, Dnd1 has also been shown to bind mRNA and repress their translation without affecting their stability during zebrafish embryogenesis
[29]. Further studies to elucidate the exact cellular and physiological context of these mechanisms are needed.
DND1 has a long list of target mRNAs
[10][23][32][33][34], which is expected to grow longer. In germ cells, DND1 down-regulates many genes associated with pluripotency and active cell cycle (e.g.,
Rheb,
Rhob,
Ccne1, and
Trp53 etc.) whereas it up-regulates chromatin regulator genes (e.g.,
Kat7,
Rbbp5,
Kdm5a, and
Setdb1 etc.)
[34], suggesting that post-transcriptional up- or down-regulation may exist concurrently in cells and could be mRNA target-specific.
2.3. Other Molecular Functions of DND1
In addition to overcoming the inhibitory action of miRNA, DND1 can also activate germline-specific translation through promoting initiation. In
Xenopus oocytes, Dnd1 binds to the mRNA of germline gene
nanos1 and interacts with eukaryotic initiation factor 3f (eIF3f) which is a repressive component in the 43S preinitiation complex
[35]. This Dnd1-eIF3f interaction counteracts the repressive activity of eIF3f and leads to
nanos1 translation.
DND1 was also found to possess a unique function of modifying the activity of target protein. In mouse embryos, DND1 interacts with the activator protein 1 (AP1) subunit c-Jun
[36]. Co-transfection with
Dnd1 and
c-Jun plasmids in GC-1 mouse spermatogonia cells significantly increased the transcriptional activity of AP1.
DND1 has been detected in both the cytoplasm and nucleus
[37][38][39], in line with the function of DND1 as a post-transcriptional
[7][9][10][29] or transcriptional regulator
[36]. What determines the multifaceted regulatory roles of DND1 is still unknown. Whether the different reported functions of DND1 are species-specific also remains to be clarified. A recent study revealed an interesting 3D domain swapped dimerization of the DND1-RRM2 domain
[40], which increases surface area for multimeric interactions and may allow DND1 to exert different functions in different context. Further investigations are warranted to elucidate the mechanisms and causes underlying the multifaceted roles of DND1.
3. Dnd1Ter Mutation in Testicular and Ovarian Teratomas
In vertebrates, PGCs segregate from pluripotent epiblast cells and migrate into the developing gonads, where they specify into either male or female germ cells. In mice,
Dnd1 expression is specifically induced in PGC precursors at embryonic day (E) 6.5–6.75 and maintained until the pre-meiotic spermatogonia stage in adult testes
[10]. Immunoblotting using mouse-specific anti-DND1 antibodies has detected DND1 in the spermatids of adult mouse testes but not in Sertoli cells
[39]. Additionally, examination of The Human Protein Atlas indicates
DND1 mRNA is enriched in spermatogonia and early spermatids (
https://www.proteinatlas.org/ENSG00000256453-DND1/celltype, accessed on 18 July 2021) from human testis
[41][42][43].
The
Dnd1Ter mutation causes PGC loss in all mouse genetic backgrounds, leading to infertility in both genders. However, only male
Dnd1Ter mice on the 129 strain background develop the cancerous TGCT. In these mice, some of the remaining PGCs acquire the intermediate pluripotent status
[44] to transform into highly proliferative, undifferentiated pluripotent embryonal carcinoma (EC) cells. At the time of birth, most EC cells differentiate into random mix of various cells and tissues that compose the teratomas of TGCTs. Although the incidence of teratomas in wild-type (WT) strain 129/Sv males is 1.4%, the
Dnd1Ter mutation increases teratoma incidence to 17% in heterozygous mutants and 94% in homozygotes
[2]. Thus,
Dnd1Ter is a potent modifier of spontaneous TGCT susceptibility on the 129 strain background.
More strikingly, in the WKY/Ztm rat strain, homozygous mutation in
Dnd1 led to formation of congenital testicular and ovarian teratomas, as well as infertility with complete penetrance in both genders
[11]. The rat
Dnd1Ter was determined as a point mutation in exon 4 of rat
Dnd1, which introduces a premature stop codon at aa 289, resulting in a longer truncated protein compared to the predicted mouse
Dnd1Ter mutant (
Figure 1). The HRAAAMA motif is lost in the mouse DND1
Ter mutant due to the premature stop codon at aa 190, but remains intact in rat Ter
[14] (
Figure 1), which may explain for the gender difference in Dnd1
Ter-related teratocarcinogensis between mouse and rat. A series of studies from Tokumoto’s lab also point to the role of DND1 in female germ cells. They determined that the
ett1 locus, on mouse chromosome 18 but separate from the
Ter locus, is responsible for experimentally testicular teratoma (ETT) formation induced in 129/Sv mice when E12.5 fetal testes are transplanted into adult testes
[45]. They subsequently identified a single nucleotide polymorphism that introduces glycine to serine mutation at aa codon 25 (G25S) in melanocortin 4 receptor (
Mc4r) gene within the
ett1 locus
[46]. In a very recent report, they showed that the MCR4 G25S homozygotes also caused OTs) in 48.7% of LTXBJ mice. The incidence of OT was further increased to 66.7% when
Ter locus of the 129 strain was introduced in heterozygous to
Mc4rG25S/G25S mice
[12].
DND1 was found to interact with several pluripotency associated mRNAs,
OCT4,
SOX2,
LIN28 and
BCL2L1, in the germinal vesicle-stage of pig oocytes
[47]. In
Xenopus oocytes, Dnd1 was also subject to ubiquitin-independent proteasomal degradation and translational repression, regulations that prevent premature accumulation of Dnd1 in oocytes
[48]. These reports highlight the role of DND1 and its interactions with specific genetic susceptibility factors that result in germ cell tumors (TGCTs or OTs) in male and female animal model systems.
In contrast to its function in mouse TGCT, germline
DND1 mutations were shown unlikely to contribute significantly to human TGCT susceptibility
[49][50]. In addition, DND1 was not detected among the panel of putative prognostic and diagnostic markers of human TGCT tissues identified by proteomic techniques
[51], indicating DND1 may not play a significant role in human TGCT. The functional discrepancy between mouse and human DND1 in TGCT is probably not surprising, considering that even in mouse, the increase of TGCT incidence by
Dnd1Ter is highly dependent on the strain genetic background
[32] and genetic susceptibility factors other than
Dnd1Ter can also increase TGCT occurrence in mouse
[45][52][53]. On the other hand, mechanistic studies indicate a link between DND1 and miRNAs that characterize human TGCTs. For example, DND1 was found to interact with
LATS2 mRNA to alleviate miR-372-mediated
LATS2 suppression
[7][32][33]. Furthermore, proteins such as APOBEC3 that interact with DND1 can modulate DND1 function of blocking miR-372
[26]. Significantly, miR-372 belongs to the miR-371-373 cluster, which is one of the most relevant biomarkers for TGCTs in patients
[54][55]. Thus, further investigation is warranted to elucidate any putative link between DND1 function and miRNA signatures of human TGCT.
4. The Emerging Role of DND1 in Somatic Cancers
Although in the zebrafish, Dnd1 is exclusively expressed in germ cells, in mammals it appears that lower levels of DND1 are expressed in a variety of cells and tissue types, mainly of epithelial or mesenchymal origin. DND1 expression has been reported in normal human neutrophils, hematopoietic CD34+ progenitor cells
[16], rat neural stem cells, neurons and astrocytes
[56] and in breast epithelial cells
[17]. Additionally, emerging evidence indicates a role of DND1 not limited to only in germ cell cancers but also in some types of somatic cancers
[13][14][15][16][17][18][19].
Table 1 summarizes the studies of DND1 in different cancer types. Importantly, these studies on DND1 not only used established cancer cell lines but used patient derived cancer tissue samples. We discuss below the studies on DND1 in breast cancer, gastrointestinal (GI) cancers, tongue squamous cell carcinoma (TSCC) and leukemia.
Table 1. Studies on the role of DND1 in somatic cancers.
| Tumor Type |
Endogenous DND1 Status in Human Cell or Tissue Samples |
Phenotypes Caused by Experimental Alteration of DND1 |
Mechanism of DND1 Function |
Reference |
| Breast cancer |
Lower expression of DND1 levels in breast cancer tissue compared to normal. |
Knockdown of DND1 in MCF-7 cells decreased BIM expression and inhibited apoptosis. |
DND1 increases expression of BIM by blocking miR-221 from BIM-3′UTR. |
[17] |
| Hepatocellular carcinoma (HCC) |
DND1 mRNA and protein levels significantly decreased in HCC sphere cells. |
DND1 overexpression inhibited spheroid formation; suppressed HCC cancer cell stemness; inhibited epithelial-mesenchymal transition; increased sensitivity of HCC cells to sorafenib. |
DND1 binds to LATS2 3′-UTR, elevating LATS2 level and YAP phosphorylation and retention in the cytoplasm, therefore diminishing transcriptional activity of YAP. |
[18] |
| Intestinal polyposis |
N/A |
Apc+/MinDnd1+/Ter mice had higher polyp numbers compared to Apc+/MinDnd1+/+ mice. |
N/A |
[14] |
| Colorectal cancer (CRC) |
DND1 expression significantly up-regulated in CRC cell lines. |
Silencing DND1 reduced SW48 cell line viability and overexpression of DND1 promoted cell proliferation. |
DND1 is the potential target of miR-24 in SW48 cells and involved in miR-24 mediated inhibitory effects on cell proliferation. |
[19] |
| Tongue squamous cell carcinoma (TSCC) |
Reduced expression of DND1 in TSCC cells and tissues. |
DND1 knockdown in TSCC cell lines enhanced cell proliferation and reduced apoptosis. Enhanced DND1 expression reduced cell proliferation and increased apoptosis. |
DND1 is a direct target of miR-24. miR-24 suppressed DND1, leading to reduced CDKN1B. |
[13] |
| Acute myeloid leukemia (AML) |
Lower DND1 mRNA levels in AML blasts and CD34+ progenitor cells. |
Inhibition of both RBM38 and DND1 mRNAs significantly attenuated NB4 differentiation and resulted in decreased p21(CIP1) mRNA. |
Activity of RBM38 and DND1 during neutrophil differentiation antagonize the activity of oncomiRs to protect mRNAs, for example p21CIP1 that are important for myeloid differentiation. |
[16] |
| Skin cancer |
Reduced expression of DND1 mRNA in transformed HaCaT cells and loss of DND1 mRNA and protein in tumors from transformed HaCaT cells. |
Expression of DND1 in transformed HaCaT cells interfered with miR-21-mediated repression of MSH2; Knockdown of DND1 reduced MSH2 RNA, an effect further enhanced by miR-21. |
DND1, which decreases sensitivity of MSH2 to miR-21, is down-regulated during tumorigenesis therefore increasing the effectiveness of miR-21 in tumors. |
[15] |
In a study on human breast cancer by Cheng et al., the group’s analysis of The Cancer Genome Atlas (TCGA) microarray mRNA expression profiles for breast cancer found that patients with higher
DND1 levels had longer overall survival
[17]. Furthermore, comparison of 21 pairs of breast tumors and their neighboring mammary normal epithelial tissues found two-fold lower
DND1 mRNA levels in breast cancer tissues compared to adjacent normal breast tissues. The lowest
DND1 levels were detected in a panel of human breast cancer cell lines with highest metastatic capability. In addition,
DND1 expression was found to be positively correlated with the pro-apoptotic effector
BIM expression in human breast cancer. Experimentally, knockdown of
DND1 in MCF-7 cells decreased BIM expression and inhibited apoptosis. DND1 was found to increase BIM expression and stability by blocking miR-221 from
BIM-3′ UTR.
Several studies also indicate that DND1 functions in GI cancers. In hepatocellular carcinoma (HCC) cells, DND1 overexpression was found to inhibit spheroid formation, suppress HCC cancer cell stemness, inhibit epithelial-mesenchymal transition and increase the sensitivity of HCC cells to sorafenib
[18]. The tumor-suppressive function of DND1 in HCC may at least partially result from Hippo signaling activation. It was shown that through binding to
LATS2 3′-UTR, DND1 overexpression led to elevated LATS2 level and YAP phosphorylation and retention in the cytoplasm, diminishing the transcriptional activity of the YAP oncogene.
In another study, mutant
Dnd1Ter was found to significantly increase polyp number and mass in the
Apc+/Min mouse model of intestinal polyposis
[14], indicating the tumorigenic properties of
Dnd1Ter in the intestine. These above studies indicate that normal, WT DND1 has tumor-suppressive type activity with increased DND1 expression being anti-proliferative, pro-apoptotic, or causing reduction of cancer stemness. However, one study contradicts this in that DND1 seems to play a tumor-promoting role in SW48 colorectal cancer cell line
[19], as silencing DND1 suppressed cell proliferation and overexpression of DND1 reversed the tumor-suppressive effects of miR-24.
In TSCC cells, DND1 itself was also identified as a target of miR-24
[13]. Down-regulation of DND1 mediated by miR-24 led to reduced expression of cyclin-dependent kinase inhibitor 1B (CDKN1B), whose translation is up-regulated by DND1. This was accompanied by enhanced cell proliferation.
Lastly, DND1 seems to also play a role in non-solid tumors. The expression of DND1 and another RNA-binding protein RBM binding motif protein 38 (RBM38) is repressed in primary acute myeloid leukemia patients
[16]. Inhibition of
DND1 and
RBM38 mRNA significantly attenuated differentiation of NB4 acute promyelocytic leukemia cells and resulted in decreased
p21(CIP1) mRNA expression.
In summary, different studies on DND1 function found that DND1 is an anti-proliferative, pro-apoptotic tumor suppressor in a variety of cancers, but may also exert oncogenic function in others. The presence of different repertoires of mRNA/protein in different cells or tissues may account for the opposite roles of DND1 detected in the cancer studies, as DND1 stabilizes some mRNAs but suppresses or degrades others
[7][9][10][29] (
Figure 2), and can also modify the activity of target protein
[36]. Moreover, how DND1 functions under different physiological conditions in the cell is unknown. It is also noteworthy that the above-described studies in different cancer types each represent a single study and further investigations are critically needed to verify these findings.
To complement the above-mentioned experimental studies on the role of DND1 in somatic cancers, we queried the frequency of
DND1 gene alteration in human cancers from TCGA database. We examined data on liver, breast, skin, colon neoplasms and AML in cBioPortal (cbioportal.org, accessed on 30 June 2021)
[57][58].
DND1 gene shows minimal alteration in these cancer types that we analyzed (data not shown), except that it is altered, mostly through deletion, in 9.38% of melanomas. In addition, we found high
DND1 alteration frequency (>5%) in other three cancer types: lung (up to 18.75%), prostate adenocarcinoma (up to 6.49%), and kidney renal clear cell carcinoma (up to 14.12%). Interestingly, while the majority of
DND1 alteration is deletion or mutation in lung and prostate cancers,
DND1 amplification prevails in kidney cancers. It is imperative to further explore the role of DND1 in these cancer types as to whether the changes in
DND1 represent passenger events or whether DND1 directly contributes to cancer susceptibility and progression.
5. Genetically Engineered Mouse Alleles of DND1
In contrast to the cancer cell line studies and cancer genome data analysis, in vivo model systems allow control of genetic background and testing of direct causal relationship, and thus will serve as important tools for elucidating the possible roles of DND1 in somatic cancers. In addition to the above-mentioned mice carrying the mutations of four aa residues in RRM1 of DND1 (E59, V60, P76 and G82)
[28], hereto, several genetically engineered mouse alleles of
Dnd1 have been generated by different laboratories to study germ cell biology and germ cell cancers.
Two
Dnd1 knockout mouse alleles generated by different research groups have reported to manifest different phenotypes and thus whether
Ter simply causes DND1 loss or produces a truncated DND1 protein remains controversial. The conventional
Dnd1-knockout mouse line,
Dnd1∆ [31], was established by administering tamoxifen globally to
Dnd1+/flox; Rosa26+/CreERT2 female mice
[9]. These
Dnd1∆ mice showed phenotypes similar to those of
Ter mutant mice in spermatogenesis, oogenesis, and teratoma incidence, with a slight difference in spermiogenesis. The DND1 protein level in
Dnd1+/Ter mice was half of that in WT embryos and the expression of the shorter, truncated DND1 protein was not detected. These data support that
Ter mutation causes DND1 loss. However, another study indicated that
Dnd1Ter is not functionally equivalent to DND1 loss in mice
[14]. This
Dnd1 knockout allele,
Dnd1KO, caused embryonic lethality in
Dnd1KO/KO homozygotes and
Dnd1+/KO heterozygotes were born with reduced numbers.
Dnd1Ter only partially rescued these phenotypes. In addition, while
Dnd1+/KO heterozygous male mice did not exhibit increased occurrence of TGCT, a single copy of
Dnd1Ter was able to increase TGCT risk regardless of whether the alternative allele was loss-of-function (
Dnd1KO) or WT. The phenotypic discrepancy of the
Dnd1∆ and
Dnd1KO mice may at least be partially due to the slight genetic difference between 129 substrains. In addition, the fact that different portions of the
Dnd1 gene were deleted in these two alleles may also account for the discrepancy. In
Dnd1∆ allele, exons 2-3 were removed, whereas exons 1–2 and most of exon 3 of
Dnd1 were deleted in the
Dnd1KO allele. One cannot exclude the possibility that a truncated DND1 is being produced from the
Dnd1KO allele that retains partial DND1 function and could thus reduce TGCT occurrence.
Mouse alleles that overexpress DND1 have also been generated. The LSL-FM-Dnd1flox ‘knock-in’ allele allows Cre recombinase-mediated expression of genetically engineered, FLAG- and myc-tagged DND1 in a cell-type-specific manner [59]. The role of this allele in cancer remains to be tested. The Dnd1-3XFLAG transgenic allele contains 3×FLAG-tag encoding sequence at the C-terminus of Dnd1, resulting in the expression of DND1-3×FLAG fusion protein under the direct control of the Dnd1 enhancer [9]. As good antibodies for co-immunoprecipitation experiments for DND1 are still unavailable, these alleles allow pull down of DND1 binding partners with FLAG or myc antibodies. Furthermore, the newly reported transgenic mouse line carrying a Dnd1GFP fusion allele enables in vivo imaging of the DND1 protein and easy sorting of DND1 expressing cells [60].
Introduction of these DND1 deleted or overexpressing alleles into mouse models for different cancer types will be highly informative to determine whether and how DND1 deletion or overexpression affects somatic cancers. These mouse alleles will undoubtedly serve as valuable tools for addressing the role of DND1 in vivo and advance our knowledge of DND1 function.