1000/1000
Hot
Most Recent

Filamentous fungi are known to biosynthesize an extraordinary range of azaphilones pigments with structural diversity and advantages over vegetal-derived colored natural products such agile and simple cultivation in the lab, acceptance of low-cost substrates, speed yield improvement, and ease of downstream processing. Modern genetic engineering allows industrial production, providing pigments with higher thermostability, water-solubility, and promising bioactivities combined with ecological functions.
| Name (No). | Producing Strains | Activity |
|---|---|---|
| Aspergillus | ||
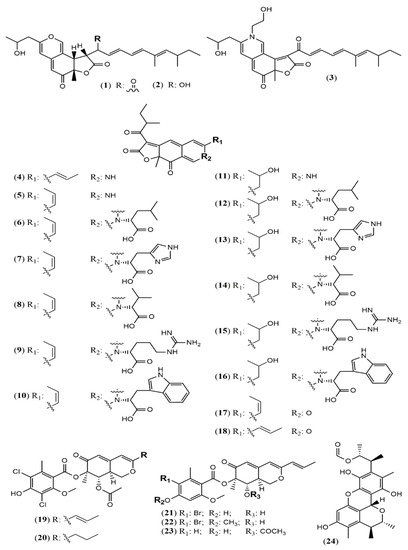
| Sassafrin E-F (1–2) | A. neogabler IBT3020 [19] |
Data not reported |
| Sassafrinamine A (3) | ||
| Trans-cavernamine(4) | A. cavernicola [20] | Data not reported |
| Cis-cavernamine (5) | ||
| Cis-cavernamines-Leu, His, Val, Arg, Trp (6–10) |
||
| Hydroxy-cavernamine (11) | ||
| Hydroxy-cavernamines-Leu, His, Val, Arg, Trp (12–16) |
||
| Cis-cavernines (17) | ||
| Trans-cavernines (18) | ||
| Falconensins O (19) | A. falconensis [21] | Anti-inflammatory (MDA-MB-231 cells line for NF-κB inhibition: 15.7 µM) |
| Falconensins P (20) | Not tested | |
| Falconensins Q (21) | Anti-inflammatory (MDA-MB-231 cells line for NF-κB inhibition: 11.9 µM) | |
| Falconensins R (22) | Anti-inflammatory (MDA-MB-231 cells line for NF-κB inhibition: 14.6 µM) | |
| Falconensins S = 8-O-Acetil-falconensin I (23) | Anti-inflammatory (MDA-MB-231 cells line for NF-κB inhibition: 20.1 µM) | |
| Penicitrinol Q (24) | A. terreus [22] | Antimicrobial (S. aureus: 4.3 mg/mL; B. subtilis: 6.2 mg/mL) |
| Chaetomium | ||
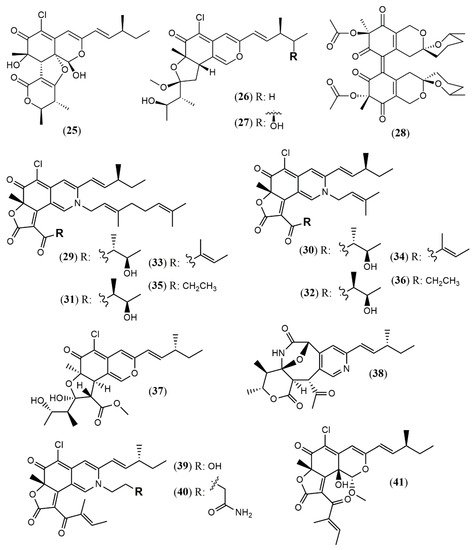
| Chaephilone C (1R,7S,8R,8aR,9E, 11S,40R,50R) (25) |
C. globosum TW1–1 [23] |
Anti-inflammatory (inhibit NO production: 15.12 µM) |
| Chaephilone D (26) | Anti-inflammatory (inhibit NO production: 20.65 µM) | |
| Chaephilone C * (27) | C. globosum [25] | Cytotoxic (HepG-2: 38.6 µM); BST (68.6% of letality at 10 mg/mL) |
| Cochliodone J (28) | C. globosum [26] | Cytotoxic (HeLa: 17.3 µM) |
| (4′R,5′R,7S,11S)-N-(3,7- dimethyl-2,6- octadienyl)-2-aza- 2-deoxychaetoviridin A (29) |
C. globosum MP4-S01–7 [27] |
Antitumor (MGC803 and AGS gastric cells lines: 0.78 and 0.12 µM, induced apoptosis) |
| 4′-epi-N-(3,7-dimethyl-2,6-octadienyl)-2-aza-2- deoxychaetoviridin A (30) | Antitumor (MGC803 and AGS gastric cells lines: 0.46 and 0.62 µM, induced apoptosis an altered the cell cycle distribution) | |
| N-(3- methyl-2-butenyl)-2-aza-2-deoxychaetoviridin A (31) | Antitumor (MGC803 and AGS gastric cells lines: 2.7 and 6.5 µM) | |
| 4′- epi-N-(3-methyl-2-butenyl)-2-aza-2-deoxychaetoviridin A.(32) | Antitumor (MGC803 and AGS gastric cells lines: 3.0 and 2.9 µM) | |
| N-(3,7-dimethyl-2,6- octadienyl)-2-aza-2-deoxychaetoviridin E (33) | Antitumor (MGC803 and AGS gastric cells lines: 0.72 and 0.12 µM) | |
| N-(3-methyl-2-butenyl)-2-aza-2-deoxychaetoviridin E (34) | Antitumor (MGC803 and AGS gastric cells lines: 6.8 and 2.0 µM) | |
| 4′,5′-dinor-5′-deoxy-N-(3,7- dimethyl-2,6-octadienyl)-2-aza-2-deoxychaetoviridin A (35) | Antitumor (MGC803 and AGS gastric cells lines: 2.2 and 1.2 µM) | |
| 4′,5′-dinor-5′- deoxy-N-(3-methyl-2-butenyl)-2-aza-2-deoxychaetoviridin A (36) | Antitumor (MGC803 and AGS gastric cells lines: 5. 8 and >10 µM) | |
| Seco-chaetomugilin (37) | C. cupreum [28] | Anticancer (MCF-7: 75.25% at 50 mg/mL) Increased ROS production: 19.6% at 5 mg/mL |
| Chaetolactam A (38) | Chaetomium sp. g1 [30] |
Cytotoxic (Not detected) |
| 11-epi-chaetomugilide B (39) | Cytotoxic (HL-60: .3.19 µM; A549: 8.37 µM; MCF-7: 4.65 µM; SW480: 4.21 µM; apoptosis induction mediated by caspase 3 in HL-60 cell: 3 µM) | |
| Chaetomugilide D (40) | Cytotoxic (HL-60: .15.92 µM; MCF-7: 17.97 µM; SW480: 14.09 µM; apoptosis induction mediated by caspase 3 in HL-60 cell: 15 µM) | |
| Globosumone (41) | C. globosum [31] | Cytotoxic (Not detected) |
| Hypoxylon | ||
| Hybridorubrin A (42) | H. fragiforme [33] | Antimicrobial (% biofilm inhibition of S. aureus: 81% at 250 mg/mL) |
| Hybridorubrin B (43) | No antimicrobial or cytotoxic activity | |
| Hybridorubrin C (44) | Antimicrobial (% biofilm inhibition of S. aureus: 82% at 250 mg/mL) | |
| Hybridorubrin D (45) | Antimicrobial (% biofilm inhibition of S. aureus: 71% at 250 mg/mL) | |
| Fragirubrin F (46) | Not tested | |
| Fragirubrin G (47) | Not tested | |
| Rutilin C (48) | Antimicrobial (% biofilm inhibition of S. aureus: 58% at 250 mg/mL) | |
| Rutilin D (49) | Not tested | |
| 3′-Malonyl-daldinin F (50) | H. fuscum [34] | Cytotoxic (L929 murine fibroblast: weak; KB 3.1 cervix-cancer cells: weak) |
| Monascus | ||
| Monapilonitrile (51) | M. pilosus BCRC 38072 [35] | Anti-inflammatory (inhibit NO production: 2.6 µM) |
| Monapilosine (52) | Anti-inflammatory (inhibit NO production: 12.5 µM) | |
| N-Ethanolic monapilosine (53) | Anti-inflammatory (inhibit NO production: 27.5 µM); cytotoxic (LPS-induced RAW264.7: cell viability< 65% at 50 µM) | |
| Muyocopron | ||
| Muyocopronone A (54) | M. laterale ECN279 [36] | Antimicrobial (Not detected) |
| Muyocopronone B (55) | Antimicrobial (methicillin-resistant S. aureus and vancomycin-resistant E. faecalis: MIC at 128 mg/mL) | |
| Lijiquinone 1 (56) | Muyocopron sp. ** [37] |
Antifungal (C. albicans: 79 µM; C. albidus: 141 µM); Cytotoxic (RPMI-8226: 129 µM) |
| Penicillium | ||
| Penicitrinone G (57) | P. citrinum WK-P9 [38] |
Antimicrobial (Not detected) |
| Dangelone A (58) | P. dangeardii [39] | Cytotoxic (Inactive: IC > 20 mmol) |
| Dangelone B (59) | Cytotoxic (HepG2: 6.82 mmol; MCF-7: 14.98 mmol) | |
| Dangelone C-G (60–64) | Cytotoxic (Inactive: IC > 20 µM) | |
| Dangeloside A and B (65 and 66) | Cytotoxic (Inactive: IC > 20 µM) | |
| Didangelone A-H (67–74) | Cytotoxic (Inactive: IC > 20 µM) | |
| Tridangelone A-E (75–79) | Cytotoxic (Inactive: IC > 20 µM) | |
| Penctrimertone (80) | Penicillium sp. T2–11 [40] |
Antimicrobial (C. albicans: 4mg/mL; B. subtilis: 4mg/mL); cytotoxic (HL-60: 16.77 µM; SMMC-7721: 23.03 µM; A-549: 28.62 µM; MCF-7: 21.53 µM) |
| Phomopsis | ||
| Phomopsone A (81) | Phomopsis sp. CGMCC No.5416 [41] |
Antiviral (Not detected); cytotoxic (Not detected) |
| Phomopsone B (82) | Antiviral (HIV-1: 7.6 µM); cytotoxic (A549: 176.7 µM; MDA-MB-231: 303.0 µM); | |
| Phomopsone C (83) | Antiviral (HIV-1: 0.5 µM); cytotoxic (A549: 8.9 µM; MDA-MB-231: 3.2 µM); apoptosis (PANC-1 cancer cells: 28.54% at 17.3 µM | |
| Tersaphilone A-C (84–86) | P. tersa FS441 [42] |
Cytotoxic (Not detected) |
| Tersaphilone D (87) | Cytotoxic (SF-268: 7.5 µM; MCF-7: 7.8 µM; HepG-2: 14.0 µM; A549: 8.3 µM) | |
| Tersaphilone E (88) | Cytotoxic (SF-268: 5.6 µM; MCF-7: 5.4 µM; HepG-2: 9.8 µM; A549: 6.7 µM) | |
| Pleosporales | ||
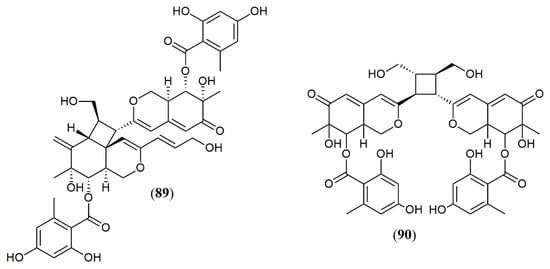
| Dipleosporalone A (89) | Pleosporales sp. CF09-1 [43] |
Cytotoxic (MDA-MB-231: 1.9 µM; HeLa: 2.5 µM; MGC-803: 1.3 µM; MCF-7: 2.1 µM; A549: 1.0 µM) |
| Dipleosporalone B (90) | Cytotoxic (MDA-MB-231: 3.8 µM; HeLa: 3.0 µM; MGC-803: 2.0 µM; MCF-7: >10 µM; A549: 3.5 µM) | |
| Talaromyces | ||
| Trans-PP-O (91) Atrosins S (92), D (93), E (94), H (95), L (96), M (97), Q (98) and T (99) |
T. atroroseus [17] | Not tested |
| Talaralbol A (100) | T. albobiverticillius [44] | Anti-inflammatory (LPS-induced NO production in RAW264.7 cell: 10.0 µM); 31.0% of inhibitory rate) |
| Talaralbol B (101) | Not detected |