Biofilm growth and evolution are very complex interactions among physicochemical and biological processes. Mathematical models are critical to modern biotechnology—both in research and in the engineering practice. Thus, many models of biofilms have been developed to include various biofilm reactor modules. However, considerable challenges exist in modelling microbial processes where mesoscopic dynamics of nutrient transport must be coupled with microscopic bacteria growth and their elementary biochemical reactions at reactive or enzymatic interfaces, in addition to the microbiological and/or ecological aspects of the “micro” organisms involved in biofilms. Lattice Boltzmann Method (LBM) treats flows in terms of fictive parcels of particles which reside on a mesh and conduct translation according to collision steps entailing overall fluid-like behavior. The goal of this review is to discuss and identify the opportunities of applying different LBM-based models to specific areas of biofilm research as well as unique challenges that LBM-based models must overcome.
1. Introduction
Bacteria mainly live together, forming complex and heterogeneous aggregations of structured communities connected together by their excretion of extracellular polymer substances (EPS). Such aggregates of microorganisms are called biofilms. Biofilms can consist of bacteria and various non-cellular materials such as mineral crystals, corrosion particles, clay or silt particles, or blood components, depending on the environment in which the biofilm has developed. Biofilms often contain one or more microbial species, which adhere to each other; to surfaces (living or non-living); and to solid–liquid, liquid–air, liquid–liquid and solid–air interfaces
[1]. Biofilms can cause many industrial and biological problems, such as chronic infections, food contamination, biofouling and equipment corrosion, although they have been applied for a variety of industrial applications, such as in biohydrogen production, fermentative food processing, power generation through microbial fuel cells, biological wastewater treatment and microbial-enhanced oil recovery
[2][3][4][5][6][7][8][9][10][11][12][13][14][15]. As such, it is important to understand the mechanisms of biofilm formation, growth and detachment, as well as their interactions with their surrounding environments, the social interactions between single-species and multispecies biofilms and the potential applications of biofilms for environmental protection and enhanced biotechnology. Extensive efforts have been made to understand the mechanisms of biofilm growth and their interactions with the affective signals
[3][16][17][18][19][20][21][22][23][24]. Due to the very complex interactions among physicochemical and biological processes, direct measurement of the impacts of bioactivity on transport dynamics in 3D structures is very difficult due to sample opacity
[25][26]; thus, many numerical models have been developed to examine biofilm growth, which can be categorized into discrete or particle-based models and continuum models
[27][28].
Biofilm modeling dates back to the 1970s. Biofilms were originally described as uniform biomasses of single microorganism species
[29][30]. The first-generation biofilm models can be implemented quickly and easily but they cannot capture certain details of biofilm structures. Later, stratified biofilms were dynamically modeled
[31]. These one dimensional (1D) stratified models were evolved to describe multi-substrate and multi-species interactions within the biofilm as the second-generation models; however, the second-generation models were not capable of describing the characteristic structural heterogeneity of biofilms. As such, discrete models of biofilm growth were developed in the 1990s and have continued until the current day as the third-generation mathematical models. Cellular automata (CA) or individual-based models (IbM) are two of the discrete model types, which have been used for simulations of inherently stochastic and individual cells and for cell spreading at the local level
[32]. The rules used to model the interactions at the local level can be determined not only through biological principles, but also through mathematical and physical frameworks using CA or IbM approaches. Hydrodynamics and multispecies transport are solved using the finite difference method (FDM), finite volume method (FVM), or finite element method (FEM). As bottom-up models, the third-generation biofilm models allow the description of the sophisticated two- (2D) and three-dimensional (3D) structures and morphologies of microbial biofilms. They can incorporate not only substrate transport and transformations
[33][34], but also hydrodynamics
[35] and population dynamics
[36]. The large-scale morphology dynamics results from the interactions of the small-scale individual cells and the environment; however, the third-generation biofilm models have difficulty in accounting for complex non-equilibrium interface dynamics, multiphase flow with phase changes and complex geometries, such as porous media.
2. Lattice Boltzmann Equation in Modeling of Biofilms
The lattice Boltzmann method (LBM)-based models are more recent in the field of biofilm modeling research than others using FDM or FVM. LBM approaches are particle-based, bottom-up models that are considered suitable for tracing the dynamics and properties of individual bacterial cells. Conversely, the behavior of individual cells is not directly tracked in continuum models and the biofilm is modeled using volume- or mass-averaged behaviors of different functional groups. The first attempt to model biofilm growth using LBM can be dated back to 1999 by Picioreanu et al.
[35][37]. In the 2000s, many researchers used the LBM approach to study biofilm growth
[34][38][39][40][41]. In LBM-CA models, the CA describes the spreading possess for a limited number of directions and with ad hoc or probabilistic rules
[42][43]. In the LBM-based biofilm models, biofilm growth is simulated through coupling with either IbM (e.g., LBM-IbM)
[28][44] or with CA (e.g., LBM-CA)
[3][19][35][37]. In recent years, LBM-based models have been extended to the modeling of several key processes, such as multispecies competition and cooperation
[3], the thermal effects on biofilm growth
[19][20], the effects of pH values on biofilm growth
[21] and heterogeneous permeability in biofilms
[28]. The LBM-based models are very effective in modeling complex multilateral interactions among biofilm growth, transport phenomenon and hydrodynamics, allowing better and more detailed description of processes in different industrial or academic applications. We call the LBM-based biofilm models as the fourth-generation models.
Biofilm growth, formation and detachment are multiscale processes in nature
[45]. Biofilm growth links cell and population scales and interacts with hydrodynamic and nutrient. Their multiscale nature requires to incorporate more intracellular states, behaviors and interplay among biofilm growth, reaction-diffusion and hydrodynamics in quantitative modeling of biofilms.
2.1. Lattice Boltzmann Equation
Compared to traditional computational fluid dynamics (CFD) using FDM or FVM, LBM, due to its fundamental kinetic nature, has a number of key benefits, such as easier parallelization, handling of pressure field (no need to solve the Poisson equation that is computationally expensive), complex boundary conditions and geometries and interface interactions. These advantages make LBM an ideal scale-bridging scheme to integrate simplified kinetic models with hydrodynamics in the simulation of microscopic/mesoscopic processes and biofilm growth.
In biofilm modeling, the system must be governed by the mass, energy and momentum conservations with their initial and boundary conditions. The substrate conservation is described by the reactive transport equation. The flow field is expressed by the Navier–Stokes equation and the continuity equation. The set of conservation equations is mathematically complicated and difficult to be handled analytically. Thus, the set of conservation equations have been solved using numerical methods, such as FDM and LBM.
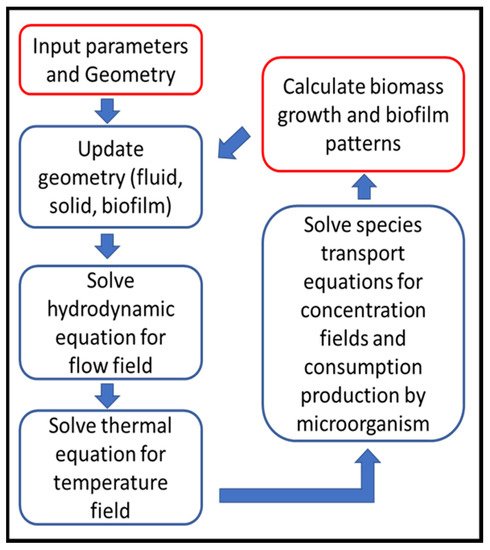
Although each biofilm simulation has its own known and unknown parameters that affect the simulation process, a general simulation procedure would generally be applicable to LBM simulations, shown in
Figure 4 [19]:
Figure 4. LBM simulation flow chart (Reprinted from
[19] with permission from American Chemical Society).
-
Geometry definition: as the first step, the computational domain is defined regarding to the bioreactor geometry.
-
There are usually three types of zones in the domain as fluid, solid, biofilm that are updated in this step due to initial geometry or zones calculated in previous time step.
-
Solve the hydrodynamic equations (Equations (2) and (3)) to capture local velocity vectors.
-
Solve energy/temperature equations (Equations (7) and (8)) to find local temperature and update fluid and species properties, if they are considered temperature dependent.
-
Solve the species transport equations (Equations (11) and (12)), that gives local species concentration and consumption by bacteria regarding to local fluid velocity, temperature and concentrations of nutrients.
-
Biomass calculation (its concentration, detachment and shrinkage) in each grid in order to determine the biofilm growth and spread of growth and spread.
-
Update biofilm pattern, affecting flow connection channels.
-
Return to Step (2) for the next time step, calculations continue up to desired total simulation time.
2.2. Applications of LBM-Based Biofilm Models
LBM-based biofilm models are based on the idea that flow field, substrate fields and temperature field can be solved by LBM while biofilm growth can be characterized using IbM or CA. Therefore, an LBM-based approach is essentially a hybrid scheme. Because modeling biofilm growth, detachment, attachment and shrinkage have two approaches: IbM and CA, an LBM model can couple with one of them to solve interactions among biofilm growth, flow and reactive transport and temperature. These hybrid models have shown their capability in representing the biofilm structure heterogeneity.
LBM-based models are more recent in time in the biofilm modeling research than others using FDM or FVM. It is common that CA is discretized spatially as same grid of rectangular elements as LBM. Each grid element has four first-order neighbors and another four second-order neighbors in the 2D square space discretization. A grid cell is allowed to be filled up to a predetermined maximum value and a simple rule is employed to locate the extra biomass in a new compartment. As one of the first studies in the file of biofilm growth using LBM-CA, Picioreanu et al.
[35][37] presented a two- and three-dimensional model for biofilm growth considering both convective and diffusive mass transfer and cellular automata approach for the growth calculation. A two- and three-dimensional solute transport were solved using LBM when convection term was considered, or the FDM if without convection term. A CA approach was used for the biofilm growth calculation. This platform simulated bacterial growth, biomass spreading and biofilm detachment for multispecies and multi substrate biofilms.
Biofilm growth in porous media has many important applications, such as, bioremediation of contaminated sites, bio-barriers development in groundwater protection, biodegradation, microbially-enhanced oil recovery, dynamic membrane filters and biofilters of wastewater treatment
[21][28][40][41][46][47]. Due to the highly heterogeneous biofilm growth within porous media, this can cause a decrease of pore space, leading to a reduction in porosity and permeability of the system and increasing hydrodynamic depression. Therefore, understanding biofilm growth in porous media is of interests to achieve an efficiency of industrial applications matter.
3. Conclusions
We present the fundamentals of LBM-based biofilm modeling first and two main integrated approaches: the integrated LBM with CA or IbM. Then, we focus on progresses of LBM-CA and LBM-IbM approaches and their applications. The LBM-based models are very promising to handle complex multilateral interactions which occur in a bioreactor and have successfully simulated many complicated phenomena in the biofilm’s dynamics and interactions between biofilm growth and hydrodynamics in different industrial or academic applications. However, long-term challenges exist in modeling biofilms when bridging different spatiotemporal scales, such as multispecies competition and cooperation, community assembly, cell-to-cell signaling, quorum sensing, antimicrobial resistance and cellular motility. Addressing these challenges would inevitably need a combined approach that not only selectively couples multiple relevant models by adding their strengths, but also the incorporation of new findings. LBM-based models doubtlessly offer new ways in understanding the dynamics of biofilms and controlling the biofilm formation and growth in public health, biotechnology and bioengineering and microbiology.