1000/1000
Hot
Most Recent

Natural products (NPs) are evolutionarily optimized as drug-like molecules and remain the most consistently successful source of drugs and drug leads. Due to traditional discovery approaches’ limitations relying on untargeted screening methods, there is a growing trend to employ unconventional secondary metabolomics techniques.
Natural products (NPs) originating from plants, animals, marine organisms, and particularly from microbial sources continue to inspire novel discoveries in chemistry [1], biology [2], and medicine [3]. They possess immense structural and chemical diversity with a wide variety of biological properties. Most pharmacologically relevant antimicrobial, antiviral, anti-inflammatory and analgesic, and antitumor agents and approved small molecule drugs have either been NPs, their derivatives, synthetic compounds with NP pharmacophore, or their synthetic mimics. Notably, more than half of the new small molecule drugs have been developed from microbial NPs over the past decades [4][5].
However, finding novel NPs has become more difficult as the rediscovery of known NPs is still an increasing challenge. A high rate of the discovery of NPs was yielded by classical methods that recover only a fraction or even none of the desired secondary metabolites. The sharp decline in discoveries arose with limitations of the traditional top-down screening approaches. Those approaches, including bioassay- and chemical signature-guided isolations, have largely been exhausted and may no longer be capable of delivering novel lead compounds [6].
In the search for alternative methods, advancements made in bioinformatics and chemical analysis might hold the key to lead a renaissance in the field of microbial NP discovery. The growing knowledge of different biosynthetic machinery, drug targets, and resistance mechanisms has served as a launch platform to a new era in the methodological approach for drug discovery [7][8]. Given the rising limitations imposed by uncultivable strains and silent gene clusters, the integrative approach of bottom-up targeted screening, employing advanced analytical methods and guided by bioinformatics analysis, provides a promising alternative for unlocking the microbial metabolomes on an unprecedented scale. This approach eventually leads to disclosing the potential of microbial NP discovery [9][10][11].
Genomics and metagenomics (which has also been described as environmental genomics, relating to the genomic DNA from an environmental sample) revealed the remarkable biosynthetic potential of microbial NPs and their vast chemical inventory that can be prioritized and systematically mined for novel or new secondary metabolites with desirable bioactivities. The growing application of bioinformatics into a standard practice in discovery projects has varied approaches to identify novel lead structures [12].
Fueled by the fast development of genome sequencing technologies, genome mining evolved during the last decades and is currently an essential part of drug discovery efforts. The genome mining approach detects and analyzes the BGCs of the chemical compounds automatically (computationally) and subsequently connects those genes to molecules. Furthermore, the significance of this approach associated with other techniques leading to drug discovery, especially of microbial NP origin, has been extensively described elsewhere [13][14][15][16]. Although the genome mining approach showcases the full biosynthetic potential of a strain, it is not very worthwhile without linking the predicted secondary metabolite BGCs to their product. Moreover, to take full advantage of NP diversity, BGCs must be prioritized by product novelty or function. BGCs hold the key information to understanding and predicting a specific or a group of related metabolites. By identifying open reading frames (ORFs) in a gene sequence, one can set the borders of the protein-encoding genes, and therein the protein sequence can be predicted through bioinformatics tools. As in some cases, bioinformatics can reveal BGCs with high similarities as a fast evaluation for target novelty; consequently, the time invested with computational work would save extensive resources and efforts only to re-isolate a previously described compound [11][16][17][18][19].
Moreover, the other culture-independent approach of metagenomics has also been established. Metagenomics relies on sampling environmental DNA (eDNA) and assessing their metabolomics independent from the producing organism. This has great implications when considering strains challenging to isolate or cultivate, such as strains from extreme environments and symbionts of marine organisms [12][20]. The revolution in this approach encompassing the phenotypic and homology DNA screening strategies in situ has been ameliorated by the advancement of next-generation sequencing (NGS) technologies [21][22][23]. By directly capturing eDNA from the environment and subsequently identifying, isolating, and expressing BGCs in a heterologous host, metagenomics has the potential to bring biosynthetic diversity from the environment into drug discovery pipelines.
The key step in compound detection and identification relies directly on analytical instrumentation and data processing software for increased sensitivity and accuracy. Given the need for increased sensitivity in metabolomics, mass spectrometry (MS) is a predominant analytical technique with wide applicability in high-throughput screening programs. It has the potential to uncover elemental composition; structural information, i.e., mass-to-charge ratios (m/z); isotopic patterns; and abundance, as well as fragmentation patterns of molecules. Current separation techniques, including high-performance liquid chromatography (HPLC) or ultra-high-pressure liquid chromatography (UPLC), as well as gas chromatography (GC), are routinely coupled to MS towards efficient detectability of the generated ions. This coupled system has proved a powerful technique that has contributed towards metabolic profiling [24][25][26].
Recent advances in MS that integrate molecular networking (MN) of the MS/MS data have allowed for more rapid dereplication of known molecules from complex mixtures (Figure 4), which in turn have enabled not only the identification of related analogs but also contributed towards unraveling novel compounds by avoiding re-isolation of known compounds.
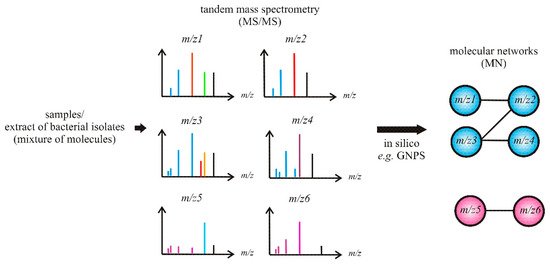
Figure 4. Schematic of molecular networking (MN)-based dereplication.
Remarkably, advancements in bioinformatics tools, genomics, and bioanalytics (particularly in MS) have recently enhanced the field of microbial NP research. These strategies outlined above offer alternatives to accelerate NP drug discovery over conventional methods efficiently. With continued significant progress in both genomics and metabolomics approaches and/or combined with synthetic biology, the microbial NPs discovery field shows strong signs of developing and is ready to lead at the forefront of delivering drugs or drug leads.