Historically, adeno-associated virus (AAV)-defective interfering particles (DI) were known as abnormal virions arising from natural replication and encapsidation errors. In this study, from analyzing the genome configurations of DI particles, a major category of DI particles is revealed, which contains a double stranded DNA genome in a “snapback” configuration. Such molecules may enhance the capsid protein expression and modulate rep expression. The subgenomic particles play an important role in the life cycle wild type AAV.
1. Introduction
AAV is a replication-defective parvovirus which requires a helper virus to complete its life cycle
[1]. The virus is best known for its small genome, which is tightly packaged inside a capsid that is 20–25 nm in diameter. Its single-stranded DNA genome contains approximately 4700 nucleotides and encodes the
rep and
cap genes which represent the non-structural and capsid proteins, respectively. AAV replication is mediated by the rep proteins, which are capable of nicking the inverted terminal repeats (ITR) and therefore initiates AAV replication. In the presence of a helper virus, AAV undergoes its lytic infection. Without a helper virus, AAV integrates into the host genome and maintains a relatively stable, latent state.
AAV infection is highly regulated in both latent and lytic infections. In latent infection, AAV genes remain silent. In the lytic life cycle,
Rep78 and
Rep68, under control of the P5 promoter, are expressed early, followed by expression of the packaging related genes, such as
Rep52/Rep40 and capsid genes
VP1,
VP2 and
VP3. In the late stage of AAV lytic infection, expression from the P5 promoter is downregulated to facilitate virus encapsidation, which was also demonstrated in recombinant AAV replication and packaging
[2]. A variety of host, viral, and helper virus factors are involved in the temporal and spatial regulation of viral replication and packaging.
Although the canonical AAV particle typically contains both ITRs and the entire coding region, AAV populations are often found to be heterogenous
[3][4][5][6][7]. Upon being processed through a CsCl gradient, the full AAV particles are present in the fraction with 1.4 g/mL density. Lighter density AAV particles (1.32 and 1.35 g/mL) are largely deemed as empty particles or defective interfering (DI) particles. DI particles contain aberrant, shorter AAV DNA genomes, and are named for their ability to inhibit AAV replication intracellularly. Template switch was proposed to explain the nonunit-length AAV genome
[6]. However, their composition and molecular conformation are elusive.
2. Cell Lines and Transfections
HEK 293 cells were maintained in Dulbecco’s modified essential medium (DMEM) containing 10% bovine calf serum. PolyJet™ DNA In Vitro Transfection Reagents (SignaGen Laboratories, Frederick, MD, USA) were used to deliver DNA into HEK 293 cells. Cells are plated into 10-cm-diameter culture dishes 18 to 24 h prior to transfection so that the monolayer cell density reaches to the optimal 70~80% confluency at the time of transfection. Complete culture medium with serum is freshly added to each plate 30 min before transfection. We prepared the PolyJet™-DNA Complex for transfection according to the ratio of 3 µL PolyJet™ to 1 µg DNA using serum-free DMEM to dilute DNA and the PolyJet™ reagent. We incubated them for 10~15 min at room temperature and added the PolyJet™/ DNA mixture onto the medium. We removed the PolyJet™/DNA complex-containing medium and replaced with fresh serum-free DMEM 12~18 h post transfection.
3. Expression Plasmids
The pH29 negative expression plasmid was made by deleting a 786-bp sbf1 and BsiwI fragment from pH29 plasmid. This plasmid expresses a normal level AAV Rep in rAAV production system but do not express AAV-2 capsid gene. The pssAAV-CB-RepCap plasmid was made by cloning the 4436-bp rep&cap fragment from pCI-RepCap into the backbone of pssAAV-CB-GFP at Bgl II site. The AAV 5′-SBG dimer was generated through the intramolecular ligation of the ITR-CB-Rep fragment, which contains AAV ITR and partial CB-Rep fragment, and isolated from pssAAV-CB-RepCap by NaeI-BamHI digestion. AAV 3′-SBG monomer was made by intramolecular-ligation of BamHI-SmaI fragment isolated from psub201. AAV 3′-SBG dimer was made by intermolecular ligation of BamHI-SmaI fragment isolated from psub201.
Head-to-head dimer DNA molecules containing Gluc gene were generated by ligation of the 1587 bp fragment, which is from pssAAV-CB-Gluc plasmid digested with SmaI and ClaI. The monomer fragment includes a single CB promoter, Gluc gene and a single Poly A signal. Tail to Tail dimer DNA molecules containing Gluc gene were generated by ligation of the 1557 bp fragment, the monomer fragment is from pdsAAV-CB-Gluc digested with SmaI and MluI.
4. Molecular State of AAV Subgenome Particles at a Single-Virus Level
The high GC content and palindromic nature of the AAV ITR has been a major obstacle for analyzing AAV genomes. The full DNA genome of the AAV virus has largely been obtained through assembling multiple fragments to obtain a consensus sequence. Even with next generation sequencing, the library construction procedures often require breaking the sample DNA and reassembling the genomes. At the single-virus level, important information became lost by this maneuver since the viral genomes in a population are derivatives of the same consensus sequences. Here we utilized the Pacbio Single Molecule, Real-Time (SMRT) sequencing platform and mapped the population genomic configuration distribution of the AAV genome at a single virus level (
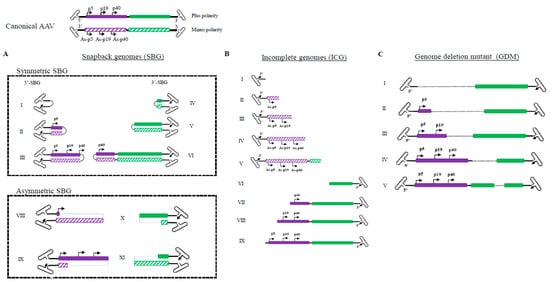
Figure 1). There were no enzymatic or mechanic actions that altered the original viral DNA configuration. From analysis of more than 400,000 ccs reads, four major categories of molecules are found in the AAV population. Category 1: Canonical AAV genomes, which contain the full AAV genome flanked by two copies of the AAV ITR; Both positive and negative strands are encapsidated equally into AAV capsids. Category 2: Snapback genomes (SBG), which contain partial duplex AAV genomes with an ITR. Such a molecule was shown to be able to snap back and anneal to itself upon denaturing and renaturing cycle
[6] SBGs are essentially a self-complementary DNA molecule with either the left moiety (5′-SBG) or the right moiety (3′-SBG) genomes. These snapback genomes were either symmetric or asymmetric. The symmetric SBG has near equal length of top strand and bottom strand when in self-complementary conformation. The asymmetric SBG has varying lengths of top strand or bottom, which leads to a single strand region as a loop. Category 3: Incomplete genomes (ICG), which contain an intact 3′ITR and a partial AAV coding sequence, but the sequences do not reach the 5′ ITR. Category 4: Genome deletion mutants (GDM), in which the middle region of the AAV genomes is missing.
Figure 1. Illustration of molecular configurations for major categories of subgenomic particles in a wild type AAV population. (Summarized from 409919 sequencing reads of two PacBio sequencing runs: 220645 and 189274) Both plus and minus strands of the canonical AAV genome are shown at the top. (A). Varying lengths of snapback AAV genomes (SBG) are illustrated, including both symmetrical snapback genomes (sSBG) and asymmetrical snapback genomes (aSBG). (B). Incomplete genomes (ICG) in the AAV population are missing the 5′ terminal sequences. (C). Genome deletion mutants (GDM) have both 5′ and 3′ ITR and miss in the mid-region of AAV. Only plus strand particles are plotted.
5. 5′ Snapback Subgenomes (5′-SBG) Function as a Cis-Negative Regulator of Rep Expression
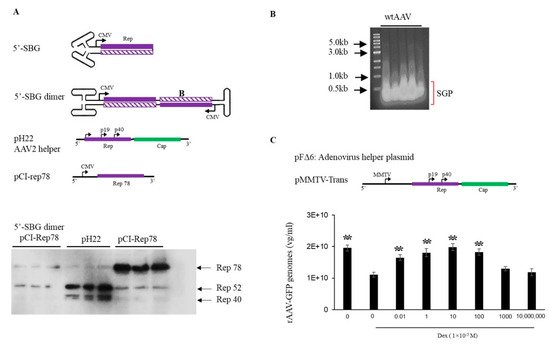
5′-SBG assumed a self-complementary configuration and was capable of replicating in the presence of Rep proteins and adenovirus helper functions, which indicated that the dimeric 5′-SBG molecules consisted of the P5 promoter followed by an inverted, head-to-head coding region. Therefore, logically dsRNA could be expressed from the dimer 5′-SBG when 5′-SBG undergoes DNA replication and becomes a dimer. Such dsDNA overlaps with P5 transcripts in AAV replication which would be able to downregulate P5 promoter expression, mainly Rep78 or Rep 68. To demonstrate its effect on rep expression, we engineered a 5′-SBG dimer with a CMV promoter, a partial rep sequence was constructed to assume a tail-to-tail dimeric configuration (Figure 2A). The RNA transcript would, therefore, assume dsRNA configuration when folding on itself. As shown in Figure 2A, the expression of Rep78 was indeed, significantly reduced in the presence of dimer 5′-SBG, which suggested that 5-SBG, potentially functioned through dsRNA intermediates, exerted its effects against Rep gene as the trans-active regulator.
Figure 2. Effects of AAV 5′-SBG dimers on rep gene expression. (A). Top panel, sketch of a 5′-SBG in self-complementary configuration; 5′-SBG dimer is extended form of 5′-SBG after 2nd stranded DNA synthesis; pH22 is the AAV2 helper plasmid expressing low level of Rep78; pCI-Rep78 is the high Rep78 expression plasmid. Bottom panel, Hek 293 cells were transfected with pCI-Rep78 with or without the presence of 5′-SBG dimers. 5′-SBG dimers were made as detailed in the experiment methods. The transfected cells were harvested at 24 h post-transfection and the expression of proteins were electrophoresed in 10% PAGE gel. The rep proteins were detected by western blot using mouse anti-AAV-Rep 76.3 antibody. Shown in the Figure are triplicate of each testing condition. Effects of AAV subgenomic particles (SGP) on rAAV replication and packaging. (B). AAV SGP were separated in an agarose gel and less than 1 kb SGP DNA identified in an agarose gel, isolated for the transfection. (C). The adenovirus helper plasmid pFΔ6, pMMTV-trans and pssAAV-CB-GFP were co-transfected into 293 cells with or without SGP DNA. Rep expression was under the control of a MMTV promoter in pMMTV-trans, which is inducible by Dexamethasone. ssAAV-GFP was collected at 72 h post-transfection and measured by qPCR. The X axis showed the copy number of SGP molecules used for transfection. “0” means no SGP DNA was added. The cells treated with DEX (1 × 10−5 M) are indicated in the Figure. Y axis: rAAV-GFP yield expressed by vector genomes per ml. ** means p < 0.05, there is statistically significant as compared to 0 M Dex treatment group.
It has been previously shown that down-regulation of
Rep78 gene expression improves rAAV packaging
[2] To test if 5′-SBG may exhibit such regulatory role, we applied extracted 5′-SBG DNA to a rAAV production system containing pMMTV-trans as the AAV rep and cap expression plasmids (
Figure 2B). MMTV is an inducible promoter which can be activated by dexamethasone. In the absence of dexamethasone, low expression of rep expression gave rise to a higher rAAV yield. Conversely, when dexamethasone was added, the vector yield was reduced. When the 5′-SBG molecules were added to the production system in the presence of dexamethasone, rAAV vector yield was increased at low concentration of 5′-SBG genomes. However, when the amount of 5-SBG was increased to more than 10 copies per cell, it started to exhibit an inhibitory effect, and the vector yield was reduced from the peak. Therefore, it was concluded that 5′-SBG had a regulatory function which senses the AAV genome pool. When SBG/AAV ratio is at a low level, it increases AAV packaging efficacy. However, when SBG is present in excess, it had an inhibitory effect and became a true “defective interfering particles”.