1000/1000
Hot
Most Recent

CSCs are small numbers of cells that exist in the tumor microenvironment (TME). Lung cancer TME is composed of a various group of non-cancer cells, such as tumor-associated macrophages (TAMs stromal cells), regulatory T cells (Tregs), tumor-infiltrating lymphocytes (TILs), dendritic cells (DCs), natural killer (NK) cells, natural killer T (NKT), myeloid-derived suppressor cells (MDSCs), along with cancer cells: mature cancer cells and CSCs. As yet, the complexity of the interactions between the cells in the immune TME has not been exhaustively described.
Lung cancer is still a leading cause of cancer-related deaths globally among men (23% of all cancer-related deaths) and women (22% of all cancer-related deaths) [1]. Worldwide, cigarette smoking alone accounts for over 80% of all lung cancer cases [2]. Other factors, such as air pollution, emission fuel combustion indoor, environmental exposure to radon and asbestos, contribute to the development of lung cancer [3]. As all these risk factors can be prevented through smoking cessation and air purification activities, it is possible to diminish lung cancer incidence and mortality through population-based preventive strategies [2].
Lung cancer is categorized into two main histological types: small cell lung carcinoma (SCLC, 15% of all lung cancers) and non-small cell lung carcinoma (NSCLC, 85% of all lung cancers). NSCLC comprises main histological subtypes: adenocarcinoma (ADC), squamous cell carcinoma (SQCLC) and large cell carcinoma (LCC) [4]. The collected data suggest that lung cancer is a group of histologically and molecularly heterogeneous disorders even within the same histologic subtype [5].
Most NSCLC patients are diagnosed at advanced stage, when the various treatments cannot be curative. Thus, this requires greater readiness of primary care physicians to carefully screen patients at high risk, even with non-specific symptoms [2]. In the course of the disease, lung cancer spreads when cells detach from a tumor and pass through the bloodstream or the lymph vessels to distant areas of the body and grow. The most common sites of the spread of lung cancer are the: lymph nodes, liver, bones, brain, adrenal glands [6].
Metastatic solid tumors, such as lung cancer, remain largely incurable despite improvements in cancer therapies. Aging is still the most important risk factor for lung cancer development. NSCLC is a heterogeneous illness with unique combinations of somatic molecular changes in individual patients, as well as significant differences in populations around the world with respect to mutation spectra and frequencies. Interestingly, it has been proven that aging is tightly associated with developing EGFR mutation in lung cancer [7][8].
Alterations in these pathways have been described in different chronic lung diseases, including lung cancer [9][10][11]. Participation of stem cells in this process is highly probable. Depending on the nature and damage size, cytostatic, cytotoxic, or mutagenic lesions arising in stem cells have the potential to lead to cells senescence, apoptosis, or transformation (Figure 1). What is more, normal stem cells can become cancer stem cells (CSCs) through the acquisition of mutation and genetic or epigenetic alterations [12].
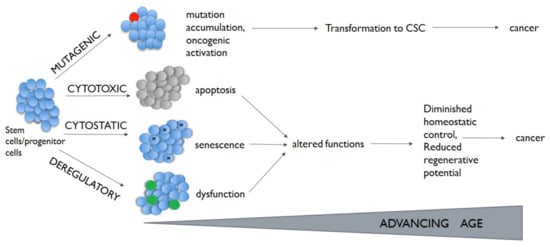
Some author suggests that driver mutations increase the cancer cell stemness, even if it is not possible to be said unequivocally that a cell with a higher number of driver mutations corresponds to less uninvolved phenotype of the cells as in the case of normal stem cells [13][14]. There are some data confirming the presence of driver mutations in CSCs (e.g., KRAS in colorectal CSCs [15]). It should be pointed out that CSCs are a diverse population of cells with relatively high plasticity and a reversible phenotype. However, an insufficient number of pre-clinical and clinical studies on CSCs in NSCLC patients make it impossible to evaluate whether the new targeted therapy may reduce the number of lung CSCs.
Nevertheless, the identification of stem cell origin in the lungs presents a challenge, because the tracheal and bronchiolar epithelia are quiescent, having a low proliferative fraction [16]. Thus, in some simplification, the origin of lung CSCs has been traced back to cells on specific anatomical sites on lungs (Figure 2). The basal cells of proximal airway, such as trachea and bronchi, are linked with SQCLC showing stem-cell-like behavior [16]. ADC is linked with normal stem cells from the bronchoalveolar duct junction area [17].
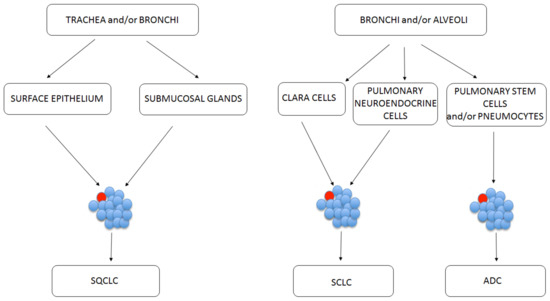
Figure 2. Possible CSC (marked red) initiation sites depending on the lung cancer subtype.
Although the present knowledge on lung CSCs functions is restricted, a number of CSC markers have been proposed. These are often markers belonging to CD (cluster of differentiation). Many studies have confirmed the presence of the following molecules on lung CSCs: CD133, CD44, CD90, epithelial cell adhesion molecule (EpCAM), C-X-C chemokine receptor type 4 (CXCR4)
We identified lung CSCs exhibiting CD133, CD44, CD90, EpCAM, CXCR4 in lung cancer patients in our previous studies [18][19][20]. The rationale for the identification of CSCs according to some authors is that they seem to be more stable, guarantee resistance to systemic therapies and, in our view, are capable of bringing information for the modification of immune response in the site of the tumor [16][21].
Patient-derived organoids can be applicable to identify treatment resistance signatures of cancer stem cells in treated organoids. Unfortunately, the majority of cancer organoid models are limited to adenocarcinomas [22]. However, as we broaden our understanding of tumor development and more, cancer stem cell markers are identified, organoids can only become a stronger research tool covering all types of cancer [22]. Thus, combined with other in vivo experiments, such as xenotransplantation assays of CSCs, organoid cultures of human CSCs have a high potential to advance our understanding of human cancer biology [23].
It is thought to be related to their activation of different signaling pathways such as: Wnt (Wingless-type), Notch, and Hedgehog [24]. A growing body of evidence supports the association of Notch signaling dysregulation with various types of malignancies, including NSCLC [25]. Notch signaling pathway play a role in stem cell maintenance in NSCLC; aberration in that pathway may result in increasing the number of CSCs resistant to platinum-based therapy [26]. What is more, Hedgehog pathway is involved in tumor drug resistance in NSCLC, as cytotoxic chemotherapy, targeted therapies and radiotherapy [27].
Lung cancer TME is composed of a various group of non-cancer cells, such as tumor-associated macrophages (TAMs stromal cells), regulatory T cells (Tregs), tumor-infiltrating lymphocytes (TILs), dendritic cells (DCs), natural killer (NK) cells, natural killer T (NKT), myeloid-derived suppressor cells (MDSCs), along with cancer cells: mature cancer cells and CSCs [28][29]. Indeed, each cell involved, immune cell or tumor cell may influence the immunological behavior of the other cells, either distant or adjacent, within the TME. Some studies showed also that CSCs may activate mechanisms to circumvent a possible attack from the immune cells: loss of cancer antigen expression, initiation of oncolytic pathways, and promotion of immunosuppressive milieu [30]. It provides the ground to induce or recruit the differentiation of suppressive immune cells, including suppressive macrophages (M2 type) or Tregs [31][32].
MDSCs represent the heterogeneous group of immature myeloid cells (precursors of macrophages, dendritic cells and granulocytes). MDSCs show pro-angiogenic activity, induce the production of metalloproteinases and contribute to the formation of pre-metastatic niches that facilitate the colonization of cancer cells [33][34]. Reports from recent years indicate that MDSCs affect the expression of oncogenes in CSCs and induce their proliferation [35][33]. Furthermore, MDSCs can regulate CSCs by the secretion of pro-inflammatory cytokines: IL-1,
TME elicits differentiation of the CD4+ T cells into different subsets of T cells, particularly suppressive Tregs, and T helper 17 (Th17) cells. Despite this, the exact role of the last cells in tumor immunity is still unclear depending on tumor stages and histological subtypes. Intriguingly, recent reports demonstrate that Tregs, under certain circumstances, express IL-17, which together with hypoxia plays a pivotal role in the regulation of CSCs [36]. Nonetheless, the interactions between CSCs and Tregs, which play an important immunosuppressive role in the TME, are still poorly understood.
The understanding of the immunological profile of CSCs and their interaction within TME has provoked the investigation of the immunological targeting of these cells (Table 1).
| Type of Immunotherapy | Condition | Study Intervention | Reference |
|---|---|---|---|
| DCs vaccination | SQCLC, melanoma | ALDHhigh CSC-pulsed DCs | [37] |
| DCs vaccination | Squamous cell cancer, melanoma | CSCs lysate-pulsed DCs | [38][39] |
| T-cell therapy | Colon cancer | CD8+ cytotoxic T-cells, specific for the CSCs antigen | [40] |
| T-cell therapy | Prostate cancer | CAR T-cells against EpCAM antigen | [41] |
| Virotherapy | Glioblastoma | Oncolytic adenovirus targeting CD133+ CSCs | [42] |
| Virotherapy | Ovarian cancer | Oncolytic vaccinia virus targeting ID8-T tumor model that harbors CSCs | [43] |
| Virotherapy | Hepatocellular carcinom | Oncolytic measles viruses: targeting CD133+ CSCs | [44] |
| Virotherapy | Breast cancer | Oncolytic vaccinia virus targeting ALDHhigh CSCs | [45] |
| Combined therapy | Bladder cancer | CSCs vaccine combinated with anti-PD-1 | [37] |
| Monoclonal antibody | Breast cancer | Anti-CD44 antibody | [46] |
| CSC-CAR T | Prostate | EpCAM-specific CAR T cell | [37] |
| Targeting signaling pathway | Lung cancer | Hedgedog pathway inhibitor | [47] |
| CSC-primed T cells | Lung cancer | CD8+ cytotoxic T-cells, ALDHhigh specific CSCs | [48] |