1000/1000
Hot
Most Recent

Host genetic diversity tends to limit disease spread in nature and buffers populations against epidemics. Genetic diversity in wildlife is expected to receive increasing attention in contexts related to disease transmission and human health.
Throughout generations, interactions with pathogens produce evolutionary changes in host populations [39][72]. In a host population, pathogen infections favor genotypes with higher resistance (ability to limit pathogen burden [73][74]) or tolerance (ability to limit disease severity induced by a given pathogen burden [73]). In addition to changes in resistance or tolerance, diseases can have other impacts on host populations: population reduction, changes of age structure, alteration on life-history parameters, or effects on genetic diversity [75]. However, the presence of pathogens can also cause changes to host behavior.
For instance, individuals can avoid infected mates to reduce pathogen transmission [76][77][78]. However, host–pathogen interactions have induced evolutionary processes that are responsible for the functioning of other mate choice-related behaviors. On the other hand, individuals (mainly females [79][80]) can choose genetically dissimilar mates to promote genetic diversity of descendants and, hence, their capacity to resist or tolerate pathogens [81][82][83]. However, the existence of infectious diseases might boost individuals to choose mates with the same level of infection, a behavior that tends to favor genetically similar mating and loss of population genetic diversity [84][85].
Studies have also investigated the effects of infectious diseases on dispersal behavior that, in turn, influence the genetic structure of populations [86][87]. Demographic declines that follow disease outbreaks increase resource availability and decrease dispersal advantages. Consequently, the low need for dispersal reduces gene flow and enhances genetic differentiation. However, the expected reduction in genetic diversity as a consequence of low dispersal might be counteracted by the effect of balancing selection acting on immune genes during the disease outbreak [88].
Pathogen– host coevolutionary dynamics may be characterized by fluctuating selection (FS), where host genotypes may be at any moment more resistant to contemporary, compared to past or future pathogens, or by arms races (AR), where both hosts and pathogens tend to increase resistance/infectivity over time [89]. The FS dynamic is based on specialized interactions and, hence, it is typical of spatially structured environments. Mixing locally adapted phenotypes may shift the coevolutionary interactions from FS to AR, due to exposure to a higher range of genotypes selected for a wider range of resistance/infectivity in both coevolutionary counterparts [90].
Since spatial variation causes the evolution of locally adaptive immunity, individuals might tend to reduce the contact or refuse mating with genetically different conspecifics harboring dangerous pathogens [91][92]. Genetically different individuals might tolerate pathogens for which local immune systems might not be prepared. They also include genes related to immune system that have not been selected under the local pathogen–host coevolutionary dynamics. These characteristics might make local individuals reluctant to contact and mate with genetically different individuals proceeding from distant populations.
Among mammals, ungulates (a paraphyletic group that includes Artiodactyla and Perissodactyla orders) include a high proportion of wild species with zoonotic diseases. [93] found that 32% of wild ungulate species (73/247 species) were zoonotic hosts. The high rates of disease transmission from wild ungulates to human have been mainly driven by our contact with these species throughout human history [94][95][96]. In addition to promoting species conservation, the maintenance of high levels of genetic diversity in wild ungulates should reduce risks regarding the emergence of infectious diseases, these risks being some of the most important threats to human health and the global economy [97][98][99].
Genetic diversity of ungulates is being altered by human-mediated processes acting on gene flow or effective size of populations. Due to hunting, competition with livestock, and lost habitat, many ungulates occur in small or bottleneck populations [100][101]. Sex-biased harvesting changes population structures and reduces effective population sizes [102][103]. These processes tend to decrease ungulate genetic diversity and, hence, affect species conservation and the probability of infectious disease emergence and spread.
Despite human-mediated alterations of gene flows and effective population sizes, studies addressing genetic diversity of ungulate populations focus primarily on conservation prospects, rather than on its effects on disease emergence and spread (Figure 1). This search showed studies that relate host genetic diversity to the spread of infectious diseases [104] (see below); studies on population genetic structure that highlight the importance of genetic diversity on disease emergence, spread, or development [105][106][107][108][109][110][111][112][113][114]; studies that analyze genetic loci or genetic metrics related to the ability of individuals and population to deal with pathogens and diseases [115][116][117]; studies that associate low genetic diversity with the presence of non-infectious diseases, alterations, or distinctive traits [118][119][120][121][122][123]; and studies on heterozygosity–fitness correlations that show expected [124] or unexpected results [125][126]. Therefore, a notable disconnection appears between studies treating ungulate genetic diversity and those dealing with infectious diseases in ungulates. The emergence and spread of infectious diseases, as well as their threats for human health and economy, might be explicitly added to conservation arguments when dealing with genetic diversity of wildlife.
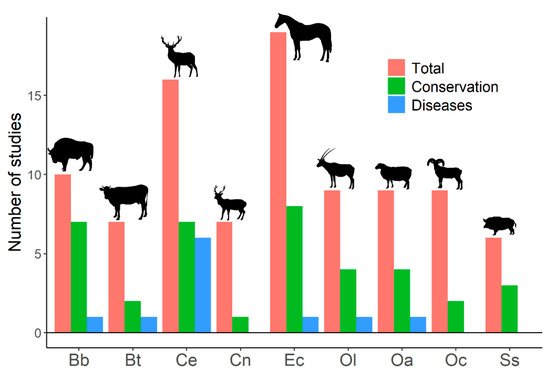
Figure 1. Number of published studies on genetic diversity for the most frequently studied ungulates. Results from a search on the Web of Science with the following search terms: genetic diversity, inbreeding, and ungulates (217 studies were obtained). Studies on genetic diversity of ungulate populations published in scientific journals were selected (204 papers). Total: number of studies on genetic diversity of ungulate populations published in scientific journals. Conservation: number of studies that explicitly related genetic diversity to conservation (papers in which the word ‘conservation’ appeared in the title, abstract, or the name of the journal). Diseases: number of studies that explicitly associated genetic diversity with diseases (papers in which the title, abstract, or name of the journal used at least one of the following terms: ‘disease’, ‘pathogen’, ‘parasite’, any variation of ‘immunity’, or the name of any disease). Bb: Bison bonasus, Bt: Bos taurus, Ce: Cervus elaphus, Cn: Cervus nippon, Ec: Equus caballus, Ol: Oryx leucoryx, Oa: Ovis aries, Oc: Ovis canadensis, Ss: Sus scrofa. The search was last consulted on 15 April 2021.
Wild boar and red deer are important game species in Spain, where most populations are in private hunting estates (typically 750–3000 ha). In these private estates, wild boar and red deer can coexist with other wild ungulates, such as fallow deer, or domestic ungulates, such as cattle [127][128]. These management actions alter ecology and behavior of individuals that, in turn, affect gene flow and effective population size. Consequently, some populations can present low levels of genetic diversity or high inbreeding, reference [104][108][129][130][131][132][133][134][135] which are associated with low antler development in red deer [121] and high predisposition to TB progression in both species [104][129][130].
A management action, presumably affecting genetic diversity, is the placement of high perimetral fences around the estates to maintain wild ungulates inside the owned land [136][137][138]. As a result of this practice, in some of the properties, we can find two types of hunting estates: open (without perimetral fences) and fenced (with perimetral fences). Additionally, small effective population sizes at the moment of fence placement might cause a founder effect with important consequences on genetic diversity and inbreeding. The lack of gene flow and founder effects might make wild boar and red deer in fenced estates present low levels of genetic diversity and might make potentially dangerous populations.
Additionally, studies comparing red deer populations in open and fenced estates have found that there are not differences in genetic diversity between both types of estates [131][134][135]. Fences might allow a certain degree of individual movements among estates and, hence, genetic diversity in fenced estates can be maintained. In addition to fence permeability, behavioral processes might avoid the loss of genetic diversity in both species [139][140][141] (see below). In spite of the existence of these processes tending to maintain reservoir genetic diversity, fenced estates, mainly those in which wild reservoirs and livestock coexist, bear potential risks that should be monitored.
Owners and managers might increase genetic diversity and reduce inbreeding in fenced estates by translocating individuals from different populations [142]. However, translocations imply the existence of additional risks that might overcome their benefits in maintaining reservoir genetic diversity. Moreover, translocated individuals swamp the genetic variation of the native populations and, hence, increase homogenization and reduce genetic diversity of a species scale [143][144][145]. In order to minimize the disadvantages of translocations, owners and managers should select individuals from populations that occur in nearby, similar habitats, and with low genetic divergence [146][147].
In open estates, managers promote the harvesting of the maximum number of males before they are hunted by neighbors. The low rate of migration rates of males among open estates might hinder the maintenance of genetic diversity and explain why open and fenced estates do not present different levels of genetic variation. Therefore, genetic diversity in open estates can be, to some extent, maintained by female dispersal. Nevertheless, a reduction in hunting harvesting over males can be recommended to equilibrate population structure and recover natural dispersal of males.
In addition to fences and population structure, landscape might also influence dispersal and gene flow in wild boar and red deer. For red deer, however, landscape features significantly affect movements [148]. In Spain, forest continuity has been shown to favor red deer movements and dispersal [134]. Therefore, to avoid loss of generic diversity in red deer populations, refuge (forest) continuity might be recommended to facilitate individual movements between open estates.
In addition to processes related to dispersal, game management affects the mating system of Spanish red deer and wild boar populations. However, high levels of mate competition in fenced estates favor the success of those males with higher levels of genetic diversity [141]. It is worth highlighting that the genetic diversity contributed by males tends to be higher than that transmitted by females in populations with high levels of mate competition [141]. The higher effective population size and genetic diversity contributed by males might help to explain why red deer populations in fenced estates have similar genetic diversity than that in open estates.
Another mating system related behavior that can be affected by game management is dissimilar mating. In open estates, the low proportion of males, that are mainly young and philopatric, hinders the action of dissimilar mating. Therefore, dissimilar mating in fenced estates, where the proportion of adult males is high, might favor genetic diversity conservation and might also help to explain the lack of differences in genetic diversity between fenced and open estates. However, this argument loses importance in those open estates where females disperse to avoid inbreeding [138].
Inbreeding depression is an important selective pressure affecting behavioral processes related to mating system [149][150]. These processes tend to compensate the loss of genetic diversity of populations and their action depends on the existence of equilibrated population structures. Male-biased hunting in red deer causes altered population structures that hinder the action of behaviors favoring genetic diversity conservation. Whenever possible, hunting regimes favoring equilibrated population structures in red deer populations might be recommended.
In wild boar, mating system related processes affecting genetic diversity have been found. Firstly, multiple paternity (different males siring offspring within the same litter) tends to maintain genetic diversity [151][152] and it has been found in wild boar populations [139]. Multiple paternity and male mate competition, both tending to favor genetic diversity conservation, are expected to act mainly in equilibrated population structures with high proportion of adult males. Nevertheless, management actions ensuring equilibrated population structures might be recommended to favor genetic diversity conservation.