Fusarium oxysporum f. sp. niveum (Fon) is the causative agent of Fusarium wilt disease of watermelon; it is the most serious soil-borne pathogen around the globe.
1. Introduction
Fusarium is a complex genus and the most diverged species in the Eumycota for its worldwide distribution, causing diseases in plants, animals, and humans as well as the profuse presence of non-pathogenic
Fusarium in the natural ecosystem
[1].
Fusarium oxysporum species complex (Fosc) is an economically devastating species of
Fusarium and is globally dispersed in various habitats, along with indoors, soil, and marine environments
[2][3]. It is a crucial ubiquitous soil-borne phylogenetic diversified fungus with a vast host range including horticultural and grain crops that cause diseases like wilt, rot, and damping-off
[4][5]. Members of this species complex are not the only source of uncontrollable vascular wilt diseases in various plants, but also the source of contagious diseases in humans and create a serious challenge to food security and public health
[6]. In terms of the economic importance of the fungus, the pathogen was ranked fifth among the top 10 plant pathogenic fungi
[7]. This seed and soil-borne plant pathogen cause serious detrimental effects on contaminated transplants showing symptoms like chlorosis, necrosis, immature leaf fall, vascular system browning, and finally wilting, which causes tremendous yield reduction. Additionally, if infection occurs earlier or during the harvesting period, some of them can produce mycotoxins in agricultural products
[8][9]. Cereals and other food grains can be contaminated by
Fusarium toxins and causes many diseases like feed refusal syndromes in mammals, mouldy sweet potato toxicity, and poisoning in bean hulls and different other living organisms
[10].
In the early 1880s, Smith identified the wilt disease of watermelon from South Carolina and Georgia after that of cotton
[11][12]. Watermelon wilt fungus was termed
Fusarium niveum by Smith (1899) and also suggested that it was a variety of
Neocosmospora vasinfectum var.
niveum. After that, Wollenweber and Reinking (1935) gave a new name of watermelon
Fusarium wilt of
F. bulbigenum var.
niveum Woll
[13]. Based on this classification, Leach and Currence (1938) reflected that the
Fusarium of watermelon and melon are various forms of
F. bulbigenum var.
niveum (form 1 and 2, correspondingly). Previously, Hansford (1926) first proposed that all species in the
Fusarium unit assembled as a sole species,
F. oxysporum [13]. Due to extreme host specificity exhibited by numerous pathogenic isolates of
F. oxysporum, finally, Snyder and Hansen (1940) restated that all species within the unit Elegans be reflected a sole species,
F. oxysporum, and proposed specialized forms (i.e.,
forma specialis (f. sp.)) that can distinguish particular virulence to one host or another
[13]. Accordingly,
F. oxysporum f. sp.
niveum is named from the watermelon wilt form 1 and
F. oxysporum f. sp.
melonis from the melon wilt form 2. After that, the concept of formae speciales achieved widespread recognition, which led to the grouping of 10 species into a single unit with many pathogenic formae speciales, and further physiological races were derived from F. sp.
[14][15].
Fusarium wilt pathogen is one of the most widely studied and devastating soil-borne pathogens around the world with both saprophytic and pathogenic members
[13][16]. Non-pathogenic and pathogenic
F. oxysporum strains remain in the soil, but the pathogenic strain causes severe vascular wilt disease in more than 150 economically major agricultural crop species. The most important crops that are likely to be infected by vascular wilt disease are banana, tomato, melon, watermelon, and cotton
[17]. In the Cucurbitaceae family, eight various f. sp. have been identified; among them,
F. oxysporum f. sp.
cucumerium (Foc; cucumber),
F. oxysporum f. sp.
niveum (Fon; watermelon), and
F. oxysporum f. sp.
melonis (Fom; melon) are enormously important. Out of this, Fon is the most destructive pathogen of watermelon around the world
[18]. The pathogen is responsible for yield losses of around 30–80% or even more
[19][20], and presently is a major hindrance in watermelon cultivation. Moreover, difficulties faced by plant pathologists include reliable identification of the causal agents of the disease according to the epidemiological related parameters such as severity, levels of species, formae speciales, pathovars, biovars, and races. The properties of these characteristics would foster appropriate and relevant control measures
[21].
2. Disease Cycle and Epidemiology
The Fon is a predominant, monocyclic, soil-borne, worldwide diversified fungus including saprophytic and pathogenic entities
[4][5]. Fon is a host-specific to watermelon, it even cannot intimately infect concomitant cucurbits crops such as cucumber and cantaloupe with some limitation, which has been studied in greenhouse conditions
[22]. The microorganism dispersing media are soil, plant debris, farm machinery
[23], and seeds
[24] and are known to survive more than 15 years without host plants
[25]. Water and contaminated farm equipment can spread the pathogens over short distances, and for extensive areas, the spread of the disease has to be through contaminated soil, seeds, or seedlings. Normally, once a region becomes contaminated, it persists so emphatically
[26]. The fungus infects the tissues of the plants as a germinating spore, and the growing hyphae penetrate the plant tissue through the wounds or openings near the site of elongation for root hair
[27]. The fungal hyphae eventually penetrate the vascular tissue and produce microconidia
[27]. The microconidia are then released into the xylem, which travels upward with the water and begins to colonize the watermelon’s vascular tissue of the plant
[28]. Afterwards, there is an infection of the watermelon plant by the pathogen frequently inhabited within it, which remains until death or decay, as presented in
Figure 1. At the last stages of the disease, the fungus produces thick mats of white mycelia and plenteous macroconidia. Under stressful environmental situations, chlamydospores develop from the aforementioned pathogen structures and conjoin into the soil.
Fusarium wilt disease is generally achieved by spreading chlamydospores, which is the primary way for the pathogen's survival
[26]. Chlamydospores are the lowest manageable attribute of
Fusarium wilt infection and can live for more than 10–15 years.
Fusarium wilt does not spread from plant to plant within season due to the absence of spore production above the ground in the field. Martyn and Vakalounakis (2017) indicated that it could also be spread by seeds
[29]. Fon was first isolated from infected seed in 1928. Since then, several researchers have confirmed the seed-borne nature of Fon. Still, the spreading mechanism of the seed-borne nature of watermelon seed is mostly undiscovered and infection rates are normally less than 5%
[13]. Another mechanism for the survival of pathogens is in the plant debris and the establishment of non-host plants that are alive
[26]. At the advanced stages of infection, permanent wilt and yield losses on watermelon could reach around 30–80% or even more
[19][20]. It becomes worse in sandy soils with a temperature range of 77° to 80 °F and a pH range of 5.5–6.5. At the anatomical level, the colonization procedure of pathogenic
Fusarium species has been described by several researchers
[30][31]. Disease development and symptom expression of host plants depend on the colonization of vessels by the pathogen
[32].
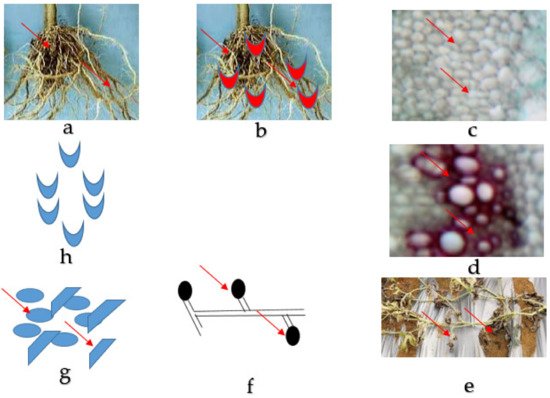
Figure 1. Fusarium wilt disease life cycle of watermelon caused by Fusarium oxysporum f. sp. niveum. (a) The healthy root of watermelon. (b) Germinating spore in contact with the root for penetration into the tissues. (c) Collapsed and distorted vessels in the xylem of the watermelon plant. (d) Gum produced by mycelia in the vessels. (e) The entire watermelon plant wilts and dies. (f) Spore formed by mycelia in the soil. (g) Micro and macroconidia present in the soil. (h) Germinating spore.
Initially, the infected watermelon plant showed symptoms like loss of pressure (turgor pressure) of vines and leaves, which could be recovered at night; exclusively single or some vines may be affected
[33]. The progress of the disease in infected seedlings changed from dull green to yellow and finally necrotic
[34]. When the fungus continues to colonize the xylem vessel, the plant forms more tyloses, finally limiting the water movement and the vine begins to wilt
[26] (
Figure 2).
Figure 2. Wilting disease symptoms of watermelon plants caused by Fusarium oxysporum f. sp. niveum under field condition. (a) Showing brown necrotic lesions at the base of the stem. (b) Showing a severe form of wilting, laterally the whole plant wilted and died. Pictures are from the personal collection of M. Z. Rahman collected during the field survey.
3. Races and Vegetative Compatibility Groups
Based on the aggressiveness of the pathogen or host cultivar’s resistance performance, Fon has been classified into four physiological races (0–3) (
Table 1)
[16][26]. Crall (1963) first reported Fon race 0 from Florida, USA, and stated that all modern varieties had resistant
F0-1 gene to the race 0, so Fon race 0 loses its economic importance as a pathogen
[13]. Fon race 1 is considered to be the widely prevalent race around the commercial watermelon-producing areas in the world. Race 1 was first identified by Smith (1894) from South Carolina, USA, and discrimination between races 0 and 1 could be more quantitative and not qualitative, and race 0 changed to race 1 based on aggressiveness
[35]. At present, many diploid (seeded) and some triploid (seedless) cultivars have developed resistance toward Fon races 0 and 1
[13][26]. Later on, Fon race 2 was discovered in Israel and subsequently identified from the United States in 1981
[13]. The prevalence of race 2 was so high in some locations and showed more virulence than Fon race 1, which can infect commercial diploid seeded and triploid seedless varieties
[22][36]. After 37 years later, another new race 3 was identified in Maryland, where it was reported to be the highest in terms of aggressiveness than races 0, 1, and 2
[16][37]. The main sources of Fon race 3 could be from contaminated seed or seedlings, selection, or mutation from races 0, 1, and 2
[16].
Table 1. Different watermelon varieties used to separate four races of Fon.
| Variety |
Response of Disease |
| Race 0 |
Race 1 |
Race 2 |
Race 3 |
| All Sweet |
R |
R |
S |
S |
| Crimson sweet |
R |
S |
S |
S |
| Calhoun Gray |
R |
R |
S |
S |
| Sugar Baby |
S |
S |
S |
S |
| Black Diamond |
S |
S |
S |
S |
| Charleston Gray |
R |
S |
S |
S |
| PI-296341-FR |
R |
R |
R |
S |
Traditionally,
F. oxysporum f. sp.
niveum can be distinguished from other formae speciales and saprophytic strains of
F. oxysporum only by its virulence on watermelon. Distinguishing races require the screening of pathogenic isolates on cultivars of varying levels of resistance. These tests are laborious and often inconsistent or inconclusive. Results can be greatly influenced by environmental factors, hosts age, inoculum level, and inoculation methods
[38]. An alternative approach to the classification of strains of
F. oxysporum is based on vegetative compatibility
[39][40] where isolates are grouped into a specific phenotypic class
[41]. Therefore, VCGs are useful for characterizing the genetic diversity within a formae speciales and, in some cases, for distinguishing pathogens from non-pathogens
[42]Genetic exchange and the sexual stage is absent in
F. oxysporum, therefore it is limited to genetic transformation and the parasexual cycle. Asexual
Fusarium oxysporum needs heterokaryosis for genetic exchange, which is regulated by a set of heterokaryon loci, and it helps in the formation of vegetative compatibility by the fusion of hypha and cell lysis
[43].
Fusarium oxysporum with the capability to produce stable heterokaryon is in the same vegetative compatible group (VCG) or more likely genetically the same or clonal lineage
[44][45].
Larkin et al. (1990) extensively studied 250 strains of Fon collected from five different states in the USA, Taiwan, and Australia and observed a significant correlation between vegetative compatibility group (VCG) and physiological races or virulence races. He reported three distinct VCGs (0080, 0081, and 0082) for Fon strains, in which race 1 and race 2 belong to VCG 0080 and VCG 0082, respectively
[46], whereas VCG 0081 comprises only one Fon strain from Florida. He also demonstrated that the pathogenic strains of Fon were incompatible with non-pathogenic
F. oxysporum; additionally, within the same race strains were compatible, but incompatibility was observed with the opposite race. Zhou and Everts (2007) described three VCGs: two were alike and earlier identified by Larkin et al. (1990) (i.e., VCG 0080 and VCG 0082), and the other one was distinct
viz. VCG 0083
[47]. His results differed from those previously described by Larkin et al. (1990) and obtained an insignificant similarity between virulence race and VCG. VCG 0080 and VCG 0082 comprised all three race (0, 1, and 2) strains, and VCG 0083 consisted of only six isolates, which were classified as race 3
[16], but none of the strains was in VCG 0081. The newly identified race 3 is pathogenic to PI 296341-FR, previously stated to be resistant to race 2. Besides, race 3 isolates are compatible vegetatively with one another (VCG 0083) and incompatible with race 1 (VCG 0080) and race 2 (VCG 0082). Among these VCGs, VCG 0080 is considered the main diverged group followed by VCG 0082, VCG 0083, and 0081 distributed in a limited geographic area.
Globally collected Fon isolates were analyzed through mtDNA RFLP and the results indicated that no similarity was reported between geographic origin and race; RFLP was grouped into two common patterns that contained all three Fon races from various regions
[48]. Some other forms of formae speciales (
Table 2) like
F. oxysporum f. spp.
melonis [49],
lycopersici [50],
cubense [51], and
asparagi [52] showed a complex relationship between genetic diversity and virulence. From these findings, it can be concluded that VCG and virulence (race or cross pathogenicity) were in a complex relationship among the strains of Fon
[53]. Perhaps VCG cannot be used to differentiate the races of Fon; instead, it only helps to separate pathogenic strains from non-pathogenic strains of Fon and in the characterization of genetic variability among the Fon strains.
Table 2. Vegetative compatibility groups, formae speciales, and several races of Fo responsible for vascular wilt disease in many important crops.
| Host |
Forma Specialis |
VCG |
Described Races |
References |
| Watermelon |
Niveum |
008- |
0, 1, 2, 3 |
[11] |
| Melon |
Melonis |
013- |
0, 1, 2, 1.2 Y. 1.2 W |
[54] |
| Cucumber |
Cucumerinum
Radicis-cucumerinum |
018-
026- |
1, 2, 3
- |
[55] |
| Bitter gourd |
Momordicae |
- |
- |
[56] |
| Bottle gourd |
lagenariae |
041- |
- |
[57] |
| Vegetable sponge |
Luffae |
- |
- |
[58] |
| Wax gourd |
Benincasae |
- |
- |
[13] |
| Tomato |
lycopersici
Radicis-lycopersici |
0030-
009- |
1, 2, 3 |
[59] |
| Pisum sp |
pisi |
007- |
1, 2, 5, 6 |
[60] |
| Radish |
raphani |
022- |
- |
[61] |
| Cotton |
vasinfectum |
011- |
1, 2, 3, 4, 5, 6, 7, 8 |
[62] |
| Banana |
Cubense |
012- |
1, 2, 3, 4 |
[63] |
| Bean |
Phaseoli |
016- |
1, 2, 3, 4, 5, 6, 7 |
[64] |
| Cabbage |
conglutinans |
010- |
1, 2 |
[65] |
| Chickpea |
Ciceris |
028- |
1, 2, 3, 4 |
[66] |
| Onion |
Cepae |
0420- |
- |
[67][68] |
| Asparagus |
Asparagi |
100- |
- |
[52] |
| Sweet potato |
Batatas |
036- |
1, 2 |
[69] |
| Sugar beet |
Betae |
027- |
- |
[70] |
| Carnation |
Dianthi |
002- |
1, 2, 3, 4 to 11 |
[71] |
| Lettuce |
Lactuace |
030- |
1, 2, 3 |
[72] |
| Gladiolus |
Gladioli |
034- |
2 |
[73] |
| Tobacco |
Nicotianae |
037- |
0, 1, 2, 3 |
[74] |
| Alfalfa |
Medicaginis |
004- |
- |
[75] |
| Potato |
Tuberosi |
035- |
- |
[76] |
| Cyclamen |
Cyclaminis |
015- |
- |
[77] |
| Chrysanthemum |
Chrysanthemi |
005- |
1, 2, 3 |
[78] |
4. Evolutionary Relationship between Races of Fon and VCGs
Restriction fragment length polymorphism (RFLP), isozyme analysis, analysis of intergenic spacer (IGS), and random amplified polymorphic DNA (RAPD) analysis were used to study genetic variability among several formae speciales of
Fusarium oxysporum and demonstrated that isolates in the same VCG were similar in genetic makeup than the isolates in different VCG(s)
[79]. The high similarities of DNA profiles among all three different races of Fon have been stated in previous works
[48][80]. All these races were similar to mtDNA RFLP as well as some common sequences of chromosomal DNA. Generally, a mutation in the gene could cause a modification in developing a new race and its virulence. It was confirmed that the transformational mutagenesis of race 2 isolates of Fon caused a change to race 0 without a definite change in the VCG trait of that isolate
[80]. Additionally, the recent instance of one race coming from the other race in a local community within the evolutionary lineage (VCG) was documented in
Fo f. sp.
lycopersici in California (USA)
[81]. The spread of different races among the VCGs, especially the presence of race 2 in all three different VCGs, implies the probability of the development of a single VCG from the other because of the mutation in a
vic gene. This mutation in the
vic gene led to a modification from incompatibility to compatibility or vice versa and has been confirmed in some other fungi
[82]. The mutation that alters the vegetative compatibility was also reported to describe the origin of two VCGs with similar virulence (race) in Fo f. sp.
melonis [49]. The mtDNA polymorphism similarity between VCGs of
Fo f. sp.
niveum was discovered by Kim et al. (1992) and supported this model of evolution
[48]. Finally, it can be concluded that the transformational mutagenesis of the Fon isolates could cause a change to race without a definite change in the VCG trait of that isolate.
5. Detection of Fon and Other Members of Formae Speciales
Rapid and precise diagnosis of the pathogen is a prerequisite for controlling disease and its management. Traditionally, researchers mainly depend on morphological study and molecular techniques
[83]. Presently, disease assessment persists with the primary technique of discriminating host range and physiological races of an infective Fon isolate
[13]. Differentiation of Fo races and formae speciales is habitually tested by using time-consuming and labour-intensive assessment of disease
[84], therefore, formae-speciales-specific DNA sequencing for molecular screening technique is highly preferable
[85].
To detect and identify various
Fusarium wilt pathogens as well as the various studies among them, several reference genomes are available that can help in the diagnosis of plant pathogens rapidly, accurately, and cost-effectively
[86]. Methods based on DNA polymorphisms, like amplified fragment length polymorphism (AFLP), simple sequence repeat (SSR), random amplified polymorphic DNA (RAPD), insertion-deletion (InDel), utilization of loop-mediated isothermal amplification (LAMP), single-nucleotide polymorphism (SNP), and using specific gene sequences like an internal transcribed spacer, β-tubulin, and calmodulin gene as well as elongation factor-1 alpha (EF-1α) have been reported to be reliable in this regard (
Table 3)
[87]. Diagnostic based genes like ribosomal intergenic spacer (IGS) or EF-1α help to differentiate between several species of fungi and sometimes subspecies separation, however, these may prove uncertain due to the presence of one or several clonal lineages of each forma specialis of
F. oxysporum [86][88].
A close relationship in evolution was observed between the five formae speciales causing wilt disease of Cucurbitaceae based on mtDNA RFLPs and the isolates of a single forma specialis was grouped along with many other isolates, not of the same forma speciales, and it was observed that many formae speciales were formed in one branch. Within the five formae speciales, Fon appeared to be the most homogenous, but the most diverged group was
F. f. sp.
cucumerinum [48]. No polymorphisms were noticed among the 13 isolates of Fon including races 0, 1, and 2 collected from Israel and the USA using mtDNA RFLP analysis
[89]. Additionally, the similarity was observed in the mtDNA RFLP map and the estimated size of the mtDNA between f. sp.
melonis and
F. f. sp.
niveum (45.1 kb and 44.5 kb, respectively)
[48][49].
Table 3. PCR primers used to detect F. oxysporum, F. oxysporum formae speciales, and their races.
| Primers Name |
Target Organism |
Target Gene |
References |
| β-tubulin |
F. oxysporum |
β-tubulin (TUB2) rDNA region |
[90] |
| EF1 and EF2 |
F. oxysporum |
Translation elongation factor1-α coding region |
[91][92] |
| ITS1 and ITS4 |
F. oxysporum |
ITS region of rDNA |
[93] |
| NMS1 and NMS2 |
F. oxysporum |
The mitochondrial
small rRNA subunit (mtSSU) region |
[94] |
| FIGS11 and FIGS12 |
F. oxysporum |
The intergenic spacer (IGS) large rRNA subunit gene region |
[95] |
| Cal228F and CAL2Rd |
F. oxysporum |
calmodulin (cmdA) |
[96][97] |
| 7cF and 11aR |
F. oxysporum |
RNA polymerase II second largest
subunit (rpb2) |
[98] |
| Uni F and UniR |
F. oxysporum |
The endo-polyglacturonase gene (Pg1) |
[99] |
| Fon-1and Fon-2 |
F. oxysporum f. sp. niveum |
Derived from the
random amplified polymorphic DNA (RAPD) fragment |
[100] |
| FONSIX6-F and FONSIX6-R |
F. oxysporum f. sp. niveum race 2 |
The SIX6 (secreted in xylemprotein 6) |
[101] |
| FNR3-F and FNR3-R |
F. oxysporum f. sp. niveum race 3 |
Pathogenicity chromosome |
[102] |
| P12-F2B and P12-R1 |
F. oxysporum f. sp. Lycopersici |
Secreted in xylem 1 (SIX1) |
[103][104] |
| SIX2-F2 and SIX2-R2 |
F. oxysporum f. sp. Lycopersici |
Secreted in xylem 2 (SIX2) |
[103] |
| SIX5-F1 and SIX5-R1 |
F. oxysporum f. sp. Lycopersici |
Secreted in xylem 5 (SIX5) |
[88] |
| SIX4-F1 and SIX4-R1 |
F. oxysporum f. sp. lycopersici race 1 |
Secreted in xylem 4 (SIX4) |
[88] |
| SIX3-F1 and SIX3-R2 |
F. oxysporum f. sp. lycopersici race 2 |
Secreted in xylem 3 (SIX3) |
[103] |
| SIX6b_210_F and SIX6b_210_R |
F. oxysporum f. sp. cubense (Foc) race 1 |
Secreted in xylem 6 (SIX6) |
[105] |
| SIX8b_206_F and SIX8b_206_R |
Foc subtropical race 4 |
Secreted in xylem 8 (SIX8) |
[105] |
| SIX1a_266_F and SIX1a_266_2_R |
Foc tropical race 4 (TR4) |
Secreted in xylem 1 (SIX1) |
[105] |
The most important point for diagnostics is the genes that can encode proteins similar to virulence
[106]. In the case of Fo, which attacks the seedlings of a tomato plant, some of the proteins were identified in secreted-in xylem (SIX) and xylem sap
[104][107]. The proteins or other sets of molecules (and small RNAs and secondary metabolites) have been reported to be associated with pathogenesis during disease progression and colonization and are widely called effectors
[106][108].
SIX genes were present across the
formae speciales and the profile of the genes was used to distinguish the distinct
formae speciales, isolates, and races
[85]. The
SIX gene profiling was used to differentiate the three races in
F. f. sp.
lycopersici [88]. The SIX genes (SIX1, SIX9, SIX4, and SIX8) were also present in another species of
F. oxysporum [109][110], and SIX6 was present in
F. f. sp.
vasinfectum [111], which infects
Brassica and
Arabidopsis [112]. Moreover, SIX6 and SIX1 were present in
F. f. sp.
betae [84]; SIX7, SIX10, and SIX1 were present in
F. f. sp.
lini and
canariensis [113]; and SIX7, SIX8, and SIX1 were reported to be present in
F. f. sp.
cubense [114], while SIX5, SIX7, and SIX3 were present in
F. f. sp.
cepae [110]. Additionally, SIX4 plays an important role in the virulence of
F. f. sp.
conglutinans, which caused a yellowish colour in the cabbage plant
[115]. A further difference between races 0, 1, 2, and 3 of Fon could be possible because of the recent studies that have focused specifically on an effector gene (SIX6), which plays a vital role in initiating R-protein-mediated immunity
[101][102]. The gene (SIX6) was reported to be effective in Fon-1 isolates that are called FonSIX6, and it could be the reason to initiate a resistance in some genotypes of Fon-1-resistance, whereas Fon race 2 isolates do not have the gene (FonSIX6 effector gene), which resulted in an escape from the higher disease severity and immune system of the plant
[101]. Besides,
F. f. sp.
cubense and
F. f. sp.
lycopersici can be differentiated from other groups of
formae speciales using PCR primers that are designed to identify the specific SIX effector genes
[109].
Species-specific primer Fn-1/Fn-2 were synthesized from ITS sequences for accurate and rapid identification of pathogenic
F. f. sp.
niveum [116]. Primer set Fn-1/Fn-2 amplified only a single PCR band around 320 bp from Fon, but the primers were unsuccessful to amplify the DNA of several other fungi and
formae speciales f. sp.
cucumerinum. In contrast, primer set FON-1/FON-2 could amplify a single 174 bp DNA fragment, which could differentiate Fon from the other
formae speciales and
Fusarium spp, but could not amplify the DNA of other
formae speciales infecting cucurbits such as
cucumerinum,
melons,
momordicae, and
luffae [117]. Additionally, secreted in xylem protein 6 (SIX6) (i.e., avirulence gene) was identified in Fon races 0, 1, and 3, but absent in race 2. As a result, Fon race 2 was capable of differentiation by using primer set FONSIX6-F/FONSIX6-R
[102]. Finally, Fon race 3 could be distinguished by using primer set FNR3-F/FNR3-R, which amplified the pathogenicity chromosome region (511 bp) of the Fon genome
[102], as presented in
Figure 3.
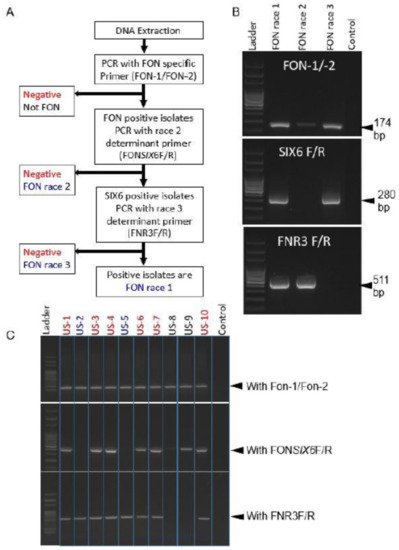
Figure 3. Flow diagram of three marker sets distinguishing Fon races 1, 2, and 3 (
A); Example of differentiation of known Fon races (
B); Identification of unknown Fon races 1, 2, and 3 (
C). Source:
[102].