1. Vaccines/Adjuvants
Vaccines have been viewed as one of the most successful health measures of all time in preventing infectious diseases caused by viruses
[1]. Certain cancers, such as human papillomavirus (HPV), are caused by viruses and, therefore, vaccines were developed to help protect against infection caused by the viruses
[2]. An adjuvant is a substance included in a vaccine that stimulates the immune system by activating pathways that the hosts naturally use to recognize bacteria or viruses. In adjuvant vaccines, the adjuvant is carefully selected combine with a specific antigen to generate specific and long-lasting immunity against the target antigen
[3][4][5]. The development of nanoparticles encapsulating specific antigens can be effective as adjuvants because they provide sustained release of the antigen
[6]. Cage protein (CP) nanoparticles are self-assembled protein structures that attract attention as a vaccine platform and have the potential to improve vaccine efficacy by promoting antigen localization and enhancing endocytosis of antigens by antigen-presenting cells (APCs). Unlike virus-like nanoparticles (VLPs), CP nanoparticles do not come from viral sources, even though they have virus-like structures and geometries
[7]. Immunotherapy utilizing vaccines have been one of the many cancer therapeutic treatments based on tumor-associated antigens (TAA) inducing specific and desired T cell responses without harming normal cells
[5].
T cells play a central role in the immune response. They originate from hematopoietic stem cells, produced from the bone marrow. Each cell develops its own T cell receptor (TCR) that is specific for a particular antigen, maturing in the thymus, and leaving to circulate through the peripheral lymphoid organs, ready to encounter specific genes and be activated
[8]. T cell lymphocytes, especially cytotoxic T cell lymphocytes (CTL), play an essential role in recognizing and killing their targets once the antigens are detected, producing cytokines, and regulating the immune response. Hence, many researchers have been taking advantage of their roles as cell killers to develop macro- and nanoparticles to be used as the antigen/drug delivery system. When delivered into the body, the cargo, usually drugs, is protected from degradation, therefore increasing the half-life of the encapsulated antigen and the immunomodulators in vivo
[9]. These particles are specifically engineered to target specific cell types, reducing off-target side effects. An example is demonstrated by the Serda group
[10] where they showed PLGA encapsulated antigen-induced T cell response with a 1000-fold lower dose compared to the nonencapsulated antigen. Different vaccine particle sizes may go through different uptake routes as described in Khong et al. in the paper published in 2016
[9]. While the complex fabrication process to develop natural proteins as nanocarriers or drugs is still impeding the use of these nanocarriers, it is worth noting that they may greatly improve the response rate and the potential advantages they possess may be the motivation behind many investigations of these nanocarriers
[11].
Although vaccinations can enhance tumor-specific immunity, it was observed that many patients do not seem to respond due to the lack of tumor-reactive T cells or poor T cell infiltration to the tumor. The need for combining different adjuvants into a single vaccine may be the next logical step to increase vaccine efficiency. It was demonstrated that various adjuvants come with both desired and undesired traits. It is hopeful that, by combining different adjuvants, the immune response can be skewed toward the favorable one. Combination therapies may be possible, where the synergy between peptide vaccines and checkpoint inhibitors can be obtained
[9], with a note given to not induce excessive T cell sequestration. Implantable and integrated smart release drug delivery and sensor devices were proposed for the distribution of insulin via feedback-controlled systems
[9].
2. Monoclonal Antibodies (MAbs)
Another immunotherapeutic treatment using monoclonal antibodies (mAbs) is called adoptive cell transfer (ACT). Patient T cells grown in the lab can recognize and kill cancer cells more efficiently compared to native immune cells. The process involves specific tumor-associated antigens (TAAs) being recognized by the genetically modified T cell, more commonly known as chimeric antigen receptors (CARs). In 2017, only two CAR T cell immunotherapies were approved by the Food and Drug Administration (FDA), one for children with acute lymphoblastic leukemia (ALL) and another for adults with advanced-stage lymphomas
[12]. Initial T cell developments start with using the patient’s own cells, which are collected and genetically modified via ex-vivo transduction using a product-specific vector. After several cell expansions, with carefully controlled manufacturing processes, they are then administered back to the patient. The personalized, specificity, and selectivity receptors minimize off-target toxicity and enhance antitumor efficacy
[13]. However, the problem occurs when the patients themselves are not healthy; therefore, impeding the proliferation rate of the cells, resulting in suboptimal cell numbers for some patients. To adapt to the fast-evolving and novel technologies, genetic engineering is utilized, by manipulating genetic codes of cells to attain better specificity and selectivity in detecting and destroying cancer cells.
3. Genetic Engineering—CRISPR/Cas
Although gene modification approaches have gone on for many years, its limitations, such as low efficiency of gene targeting, remain one of the challenges that researchers are currently facing. One strategy that has been studied to increase efficiency is to introduce the DNA double-strand breaks (DSBs) at the genomic locus of interest, interest
[14], resulting in the development of site-specific nucleases, such as zinc-finger nucleases (ZFNs)
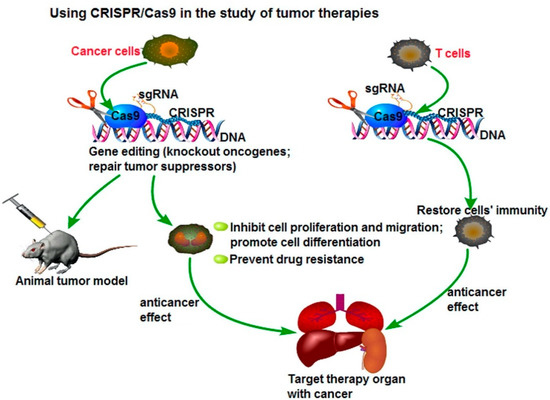
[15][16][17][18]. However, CRISPR-Cas9 methods are cheaper, faster, and more accurate because they only require a short stretch of RNA to target a genomic site. This method can easily and selectively disable or change genes in human cells, providing promising gene therapy treatments to cancer and inherited genetic disorders. A possible application of the CRISPR/Cas9 system to cancer treatment is to directly target the tumor marker in cancer cells and eliminate the genetic alterations of tumor proliferation and metastatic capacity, as seen in . Another potential therapeutic application could be the fusion of dead Cas9 (dCas9) to histone modifiers and proteins involving altering DNA methylation, to target cancer epigenetics, such as acute lymphoblastic leukemia or Ewing sarcoma. The delivery of Cas9 protein with sgRNA is the most straightforward way to achieve gene editing, leading to minimal off-target effects and toxicity due to the transient functionality
[19][20].
Figure 1. Using CRISPR/Cas9 in the study for tumor therapies. Reprinted with permission Figure
[19], Clinical Genetics, 2019.
The CRISPR/Cas9 system, currently one of the more commonly known genetic engineering methods, is an adaptive immune defense mechanism, which is a naturally occurring DNA-cutting system found in bacteria for degradation of foreign genetic material
[21]. In a paper published in
Science in 2012, it was demonstrated that this genetic scissors can be used as a genome-editing tool
[22]. CRISPR stands for clustered regularly interspaced short palindromic repeats and this versatile system consists of two biological components, the CRISPR RNA (crRNA), and the CRISPR-associated endonuclease (Cas9) modules. In general, the endonucleases break the double-stranded DNA and crRNA will then target the specific DNA sequence. The single guided DNA (sgDNA) produced from the crRNA module, and Cas9 can be optimized without altering the function of each other. In mammalian cells, gene knockout is highly prevalent when Cas9 is targeted to the exon regions of specific genes
[15]. CRISPR–Cas9 becomes useful when one can identify a target gene or protein during the drug discovery process. CRISPR–Cas9 allows the editing of genes that cause cancer cell proliferation; it can be used to insert suitable genes after the “cut”. Since its discovery, many researchers have started exploring the roles of genes implicated in cancer initiation, progression, and therapeutic response
[14] and apply CRISPR/Cas9 in their investigations.
The strategies for delivering CRISPR-Cas9 via delivery vehicles are often impeded by the high molecular weight and complexity of the system. However, as mentioned previously, researchers developed new nanoclews, a vehicle covered with positively charged material, such as lipids, which can disrupt the endosomal membrane but remain free inside the cell
[23]. CRISPR–Cas9 systems have also shown multiple advantages over many conventional gene technologies, where researchers can develop genetically modified T cells that enhance their ability to detect and kill cancer cells. The T cells can be modified in a way where they express the chimeric antigen receptors (CARs) on the surfaces, enabling the detection and destruction of specific cancer cells
[24]. Researchers in Sichuan University in China injected a non-small cell lung cancer (NSCLC) patient with genetically modified T cells containing the CRISPR-edited gene in October 2016
[25][26][27]. PD-1 genes, commonly found in NSCLC patients, are an “off-switch” for T cells from damaging healthy tissues, but cancer cells can hijack the system and avoid detection from T cells. Therefore, genetically modified PD-1 knockout T cells are injected back into patients, and subsequently are able to identify and attack the cancer cells. Similar clinical trials using CRISPR as cancer immunotherapy are also ongoing in China for esophageal cancer patients, as well as patients with B-cell lymphoma and leukemia
[28][29].