When plants reach a developmental stage for flowering competence,
FLC must be silenced. Multiple chromatin modifiers and RNA-binding proteins participate in
FLC transcriptional and posttranscriptional regulation during this process
[10][29]. Two possible mechanisms have been proposed. The first one considered a set of antisense long noncoding RNAs (lncRNAs) collectively called
COLD INDUCED LONG ANTISENSE INTRAGENIC RNAs (
COOLAIR), which are transcribed from the 3’-end of
FLC and proximately polyadenylated by FCA, FPA, and FY, and spliced by PRP8
[30]. Some of the
COOLAIR variants could lead to FLOWERING LOCUS D (FLD) recruitment, which demethylates H3K4me2, reducing the transcription of both sense and antisense
FLC mRNAs
[31]. Alternatively, there is evidence that FCA binds to CURLY LEAF (CLF), a Polycomb repressive complex (PRC2) subunit, in the presence of
COOLAIR for H3K27me3 deposition at the
FLC locus
[32].
3. Endogenous Signals in Flowering Transition
Plant’s endogenous cues that participate in flowering transition are present independently of the season, although environmental signals influence them. GAs hormones play an essential role in this process, as well as T6P, which constitutes a sensor of sugar availability and hence the plant’s reserves to support reproduction
[8][16]. Both signals are interconnected with the miR156 - SQUAMOSA BINDING PROTEIN LIKE (SPLs) - miR172 regulatory module
[15] and the MADS-domain TFs to induce flowering transition.
As the plant ages, miR156 levels gradually decrease, allowing
SPL mRNAs accumulation. In turn, SPL9, SPL10, and SPL15 induce reproductive transition by regulating genes related to flowering and binding to the
MIR172B promoter
[38][39]. Subsequently, the accumulation of miR172 indirectly promotes flowering transition by targeting
APETALA 2 (AP2) and
AP2-like flowering repressors, which repress
SPL3,
SPL4, and
SPL5 (
SPL3/4/5) genes
[38][40].
GAs constitute a group of tetracyclic diterpene compounds that have multiple functions in plant development. It was Lang (1957) who first described the effect of these hormones as bloom inducers
[41]. Specifically, high levels of GA
4 correlate with strong induction of
LFY and
AP1 at the incipient FM
[42].
Induction of
SPL3/4/5 by GA in the SAM is mediated by SOC1
[8][43]. Furthermore, GAs are required to release SPL15 from DELLAs, REPRESSOR OF
ga1-3 (RGA), and GIBBERELIC ACID INSENSITIVE (GAI) inhibition. SPL15 then associates with SOC1, which, in collaboration with the REF6 and MED18 complex, induce
FUL and
MIR172B [39]. Additionally, GAs are important for upregulation of
SPL10, which indirectly contributes to
FT induction
[44][38].
Interestingly, DELLA/GA signaling affects SPL9 function by two opposing mechanisms. On one hand, GA-induced ubiquitination of RGA is required for SPL9 to induce
SOC1 [45]. On the other hand, SPL9 seems to require the presence of DELLAs to induce
AP1 transcription, since SPL9-mediated expression of
AP1 strongly decreases when plants are treated with GAs. The latter mechanism indicates that DELLAs also participate as co-activators
[45].
Besides SPLs, GA signaling contributes to flowering transition by regulating the expression and activity of some MADS-box TFs in different manners (). For example, FLC binding to
SOC1 and
FT regulatory regions is enhanced in the presence of RGA; thereby, its degradation by the addition of GA
3 reduces
FLC repressive action
[46]. Moreover, GAs induce
SOC1 through the activity of NF-Y, or by downregulation of
GATA NITRATE-INDUCED, CARBON METABOLISM INVOLVED (
GNC), and its paralog
GNC-LIKE (
GNL), which encode direct repressors of
SOC1 [47][48].
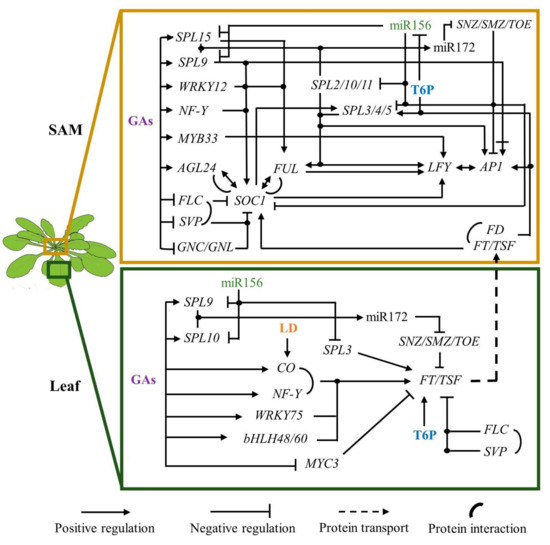
Figure 1. Gene regulatory network that promotes flowering transition in response to endogenous cues. Intricate regulations among the miR156-SPLs-miR172 module, and GAs and T6P signaling lead to FT induction in the leaves and LFY and AP1 at the flanks of the IM. MADS-box, SPLs, AP2-like, and WRKY TFs have a preponderant role in flowering transition. SPL9 dual regulation on AP1 (positive and negative bars), implies that in the presence of DELLAs, AP1 is induced, but GA treatment drastically reduces it. Line intersections with dots indicate molecular interactions.
Also, DELLA proteins inactivate some of the WRKY TF family members implicated in flowering. WRKY75 and WRKY71 are direct regulators of
FT, and WRKY71 also binds to
LFY, although it is not clear if its TF activity depends on GA signaling
[49][50]. Furthermore, WRKY12, in association with SPL10, induces
MIR172B, while WRKY13 and SPL10 repress it
[51]. Both of them interact with GAI and RGA-LIKE 1 (RGL1), and the degradation of these DELLAs by GAs enables WRKY12 to upregulate
FUL expression directly and
SOC1 indirectly
[52]. Subsequently, FUL induces flowering partly in collaboration with SOC1
[53]. SOC1 and FUL suppress SVP repression over
GA20OX2. Thus, by lowering the repressor activity of SVP, GA levels rise, reinforcing flowering transition
[54]. SOC1 and AGL24 mutual induction is also enhanced in response to GA
[55]. Finally,
XAANTAL2 (
XAL2) is another member of the MADS-box family involved in flowering, probably in response to GA and independently of
SOC1, since the double mutant
xal2-2 soc1-7 was unable to flower in response to GA
3 treatment after 88 days under SD conditions
[56].
GAs regulation of MADS-box genes leads to the induction of
LFY expression
[57]. However, GAs can upregulate
LFY expression by an independent mechanism. MYB33 accumulates in the shoot apex in response to GA
4 addition
[58]. Furthermore, MYB33 binds to a highly conserved region in the
LFY promoter
[59].
The carbohydrates were long considered to be involved in the vegetative to reproductive transition
[60]. However, the mechanisms by which sugars regulate flowering are not yet completely understood. Nevertheless, T6P signaling is critical for flowering transition regardless of day length
[16]. T6P is produced from glucose-6-phosphate and uridine diphosphate (UDP)-glucose by the TREHALOSE-6-PHOSPHATE SYNTHASE 1 (TPS1)
[61]. T6P cascade induces
FT and its closest homolog
TSF in a CO-independent manner
[16]. Additionally,
TPS1 is highly expressed in the IM, and its overexpression reverts the late-flowering phenotype of
ft-10, suggesting that T6P acts independently or downstream of FT regulation
[16].
Under SD photoperiod, T6P positively regulates the expression of
SPL3/4/5 at the SAM in a partially independent manner of miR156 decreasing by aging
[16]. However, there is also evidence that an increase in endogenous sugar levels represses miR156
[62][63].
The close relationship between SPL and MADS-box TFs, as well as their response to endogenous and environmental cues during plant’s phase transitions, strongly suggest that they are part of the same gene regulatory network (). Moreover, it has been suggested that both groups of proteins act as pioneer transcription factors, binding, and opening inaccessible chromatin by recruiting chromatin remodeling complexes
[64].
4. Long-Day Photoperiod and High-Temperature Function as Inductive Signals
Day length and high permissible temperatures are important signals to induce flowering in A. thaliana. Detection of photoperiod relies primarily on CO protein, a B-box-type zinc finger TF with a CCT domain, that accumulates during the day in the vascular tissue [4]. CONSTITUTTIVE PHOTOMORFOGENIC 1 (COP1) and SUPPRESSOR OF PHYA-105S 1 (COP1/SPA1) complex ubiquitinates CO protein to be degraded by the proteasome at night. During the diurnal phase of long-day (LD)-photoperiod, cryptochromes 1 and 2, and phytochrome A are activated by blue and far-red light, respectively, inhibiting COP1/SPA1 activity. This action allows CO accumulation in the afternoon [65][66].
CO induces FT expression [18] which is translated and systemically travels from rosette leaves to the SAM to induce flowering [67][68]. Once they reach the SAM, FT interacts with the bZIP transcription factor FD via the 14-3-3 growth response factors [69]. Moreover, the FT-FD complex regulates the expression of SPL3/4/5 in the IM, and they directly or in association with FD upregulate FUL, LFY, and AP1 [70][71].
PHYTOCHROME INTERACTING FACTOR 4 (PIF 4) and its orthologs PIF5 and PIF7 are responsible for inducing FT and TSF expression in response to high temperature under short-day conditions [72][73]. Interestingly, at 27 °C, the H2A.Z-nucleosomes levels decreased at the FT locus, relaxing the chromatin and favoring the union of PIF4 to FT DNA [72]. It seems, therefore, that LD-photoperiod and high temperature are signals that can be separated, however, more research is required to establish their possible synergism.
The actual hierarchical flowering model proposed that different inputs converge into the integrators that transduce these signals to the FM identity genes [1]. Alternatively, we propose that endogenous signals can transversally dictate whether the plants remain in the vegetative phase or initiate the reproductive state. In this view, miR156, the DELLA proteins, and possibly low concentration of certain carbohydrates maintain the vegetative state, while miR172, GA, and T6P allow the reproductive phase change. Inductive signals like LD-photoperiod and temperature accelerate the flowering transition process in plants such as Arabidopsis, in part, by upregulating FT in the leaves and a group of the SPLs and MADS-box genes in the apical meristem. This ensures that flowering happens when the external conditions are optimal for those species.