FoxO1 is a conserved transcription factor involved in energy metabolism. It is tightly regulated by modifications on its mRNA and protein and responds to environmental nutrient signals. FoxO1 controls the transcription of downstream genes mediating metabolic regulation. Dysfunction of FoxO1 pathways results in several metabolic diseases, including diabetes, obesity, non-alcoholic fatty liver disease, and atherosclerosis.
1. Introduction of FoxO1
Forkhead Box O (
FoxO) genes belong to a transcription factor (TF) family characterized by the presence of a conserved DNA-binding domain (the Forkhead box) in the N-terminal end of the protein
[1][2][3] (A). FOXOs bind the consensus DNA-binding element within the promoter of their target genes and regulate their transcription in response to external signal stimulations
[4][5][6] (B). There are four
FoxO genes identified in mammals, including
FoxO1,
FoxO3a,
FoxO4, and
FoxO6 [7]. In the 1990s, the
FoxO1 gene was first identified in studies of chromosomal translocations found in human tumors, including rhabdomyosarcomas and acute myeloid leukemias
[1][3][8][9]. In addition to its important regulatory roles in oncogenesis
[10],
FoxO1 also transcriptionally mediates pathways behind many metabolic diseases, including gluconeogenesis, glycogenolysis, adipogenesis, thermogenesis, and feeding behavior
[11][12][13][14][15], which is the focus of this review article.
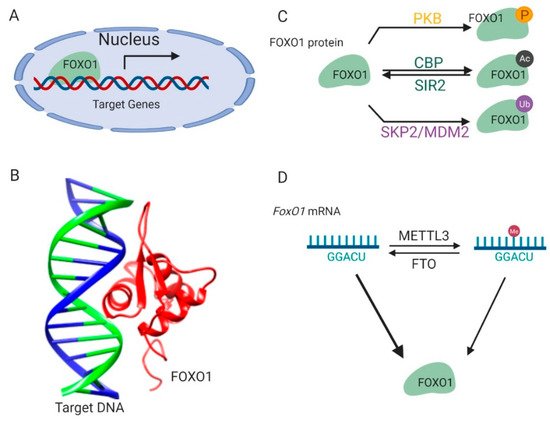
Figure 1. Regulation of
FOXO1 transcription activity. (
A) FOXO1 binds to the promoters of target genes and stimulates their transcription in nucleus. (
B) The complex structure of the FOXO1 protein and its target-binding DNA (Protein Data Bank id: 3CO6). (
C) The post-translational modifications of the FOXO1 protein. (
D) Methylations on the
FoxO1 mRNA regulate its expression. This figure was created with UCSF Chimera
[16] and
BioRender.com.
2. Regulation of FoxO1
2.1. Protein Modifications
FOXO1 protein is sophisticatedly and strictly regulated by modifications on its protein and mRNA, which ensures that transcription of its downstream target genes is tightly responsive to environmental signals
[17] (C). Kinase-mediated phosphorylation of the FOXO1 protein in response to insulin or growth factors is a form of regulation
[18] (C). Phosphoinositide 3-kinase/protein kinase B (PI3K/PKB) phosphorylates the FOXO1 protein at three conserved residues, including Threonine 24, Serine 256, and Serine 319. These modifications result in disrupted interactions between the FOXO1 protein and its target DNA and lead to the translocation of the FOXO1 protein from the nucleus to the cytoplasm, thus suppressing FOXO1-dependent transcription. The FOXO1 protein can also be phosphorylated by c-Jun N-terminal kinase (JNK) or macrophage-stimulating 1 (Mst1)
[19][20]. This phosphorylation results in the import of the FOXO1 protein from the cytoplasm to the nucleus, thereby antagonizing the action of PI3K/PKB.
FOXO1 protein activity is also regulated by reversible acetylation modification on lysine residues (C). They are acetylated by histone acetyltransferase cAMP-response element-binding protein (CREB)-binding protein (CBP) and deacetylated by NAD-dependent histone deacetylase silent information regulator 2 (Sir2) on conserved residues (Lys 242, Lys 245, and Lys 262)
[21][22][23]. The positive charge of these lysine residues in FOXO1 contributes to DNA binding. Thus, acetylation at these residues attenuates the ability of FOXO1 to bind DNA and suppress transcription. Interestingly, it was also found that acetylation regulated the function of FOXO1 by influencing its sensitivity for phosphorylation.
The FOXO1 protein is polyubiquitinated by E3 ligases (C) such as S-phase kinase-associated protein 2 (SKP2) and mouse double minute 2 homolog (MDM2), and targeted for protein degradation in human primary tumors and cancer cell lines
[24][25][26]. The ubiquitination and proteasome degradation of the FOXO1 protein are essential in tumorigenesis and represent a viable target for cancer treatment.
2.2. mRNA Methylation
In addition to these post-translational modifications on protein,
FoxO1 is also regulated by methylation modification on its mRNA
[27][28][29] (D). There are two specific adenosine sites in the coding sequence of the
FoxO1 mRNA
[27][28]. The adenosine (A) at these sites can be methylated to N
6-Methyladenosine (m
6A) by methyltransferase like 3 (METTL3) protein, and m
6A can be demethylated to A by fat mass and obesity-associated protein (FTO). These reversible m
6A modifications participate in regulating
FoxO1 gene translation
[29].
3. FoxO1-Regulating Mechanism Behind Diseases
3.1. Glucose Production in the Liver
FoxO1 regulates many important metabolic pathways in the liver, fat tissue, and hypothalamus
[30][31]. It is well established that
FoxO1 regulates hepatic gluconeogenesis and glycogenolysis in response to an insulin signal in the blood (). High concentration of insulin decreases blood glucose level by promoting glucose absorption after feeding and inhibits glucose production by hepatic gluconeogenesis and glycogenolysis in the fasting state.
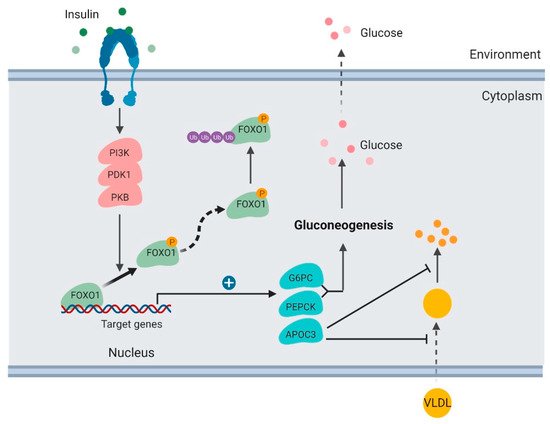
Figure 2. Regulation of glucose production and lipoprotein uptake by
FOXO1 in the liver (created with
BioRender.com).
In the liver, insulin activates the PI3K/PKB signaling pathway and results in FOXO1 protein phosphorylation and degradation
[32]. FOXO1 TF binds and promotes transcription of
glucose 6-phosphatase (
G6PC) and
phosphoenolpyruvate carboxykinase (
PEPCK), which are key enzymes stimulating gluconeogenesis and glycogenolysis
[33][34]. When FOXO1 is suppressed, transcription of
G6PC and
PEPCK subsequently decreases, which consequently inhibits the rates of glucose production in the liver.
It has been well established that
FOXO1 is a key mediator in the signaling pathway of insulin regulating hepatic gluconeogenesis. Hepatic
FOXO1 loss-of-function mutant suppresses
G6PC and
PEPCK expression, decreases hepatic gluconeogenesis, and improves fasting glycemia in diabetic
db/db mice
[32].
3.2. Lipoprotein Uptake in the Liver
Apolipoprotein C-III (
ApoC3) is another downstream target of
FoxO1 that functions directly in plasma triglyceride metabolism
[31][35].
ApoC3 is secreted by the liver and enriched in very low-density lipoprotein (VLDL). It was reported to suppress hepatic uptake of VLDL and inhibit lipoprotein lipase
[36].
ApoC3 overexpression in humans functions in atherosclerosis
[37].
FOXO1 binds to the ApoC3 promoter and enhances its transcription
[35] (). FOXO1 overexpression increases hepatic ApoC3 expression and elevates plasma triglyceride levels. FOXO1 loss-of-function mutation interferes with
ApoC3 expression in response to insulin stimulation. Insulin deficiency or resistance results in unrestrained ApoC3 expression and impaired triglyceride metabolism in the pathogenesis of atherosclerosis and hypertriglyceridemia.
3.3. Lipogenesis in the Liver
Clinically, it was observed that therapeutically decreasing blood glucose usually caused increased lipogenesis in the liver
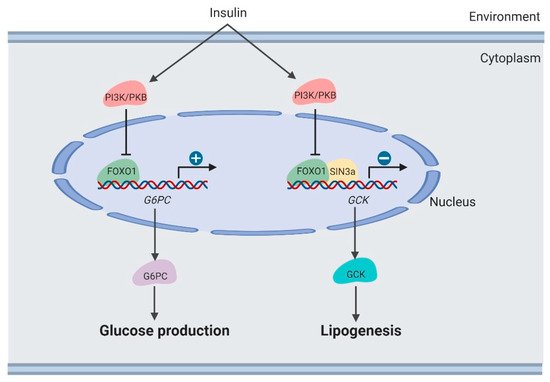
[11][38]. Lipogenesis is induced by suppressing the
FoxO1-dependent inhibition of
glucokinase (
Gck)
[39] (). Hepatic
Gck expression is associated with lipogenesis and fatty liver in humans. Increased
Gck activity induced fatty liver and its metabolic consequences in humans.
Figure 3. Regulation of lipogenesis by
FOXO1 in the liver (created with
BioRender.com).
Moreover,
SIN3 Transcription Regulator Family Member A (
SIN3A) was identified as an insulin-sensitive
FOXO1 corepressor of
Gck.
SIN3A knockout interferes with
Gck expression regulated by
FOXO1 in response to environmental nutrients, while not affecting expression of other genes targeted by
FOXO1, such as
G6PC. These results may provide the opportunity to develop selective modulators of the
FoxO1 pathway, which removes adverse effects of hepatic lipogenesis in therapeutically treating diabetes
[39].
3.4. Lipid Metabolism in Adipocytes
FoxO1 binds to the promoter sites of
peroxisome proliferator-activated receptor gamma (
PPARγ) DNA and represses its transcription
[40]. In response to insulin stimulation, FOXO1 protein is degraded and subsequently unable to prevent transcription of
PPARγ. Because high levels of PPARγ protein initiate adipogenesis,
FoxO1 suppresses adipogenesis
[41].
Moreover, the FOXO1 protein is a repressor of
uncoupling protein 1 (
Ucp1) gene transcription
[31][42]. UCP1 protein is a mitochondrial inner membrane proton channel linked to energy use that serves as a well-known biomarker of the thermogenesis state of adipose tissues
[43][44][45].
FoxO1 inhibition increases UCP1 expression, thereby augmenting thermogenesis and fat loss. Selectively inhibiting FTO by inhibitors decreases FOXO1 expression and reduces body weight and fat mass in a high-fat diet-induced obesity (DIO) mouse model. Suppressing FOXO1 expression increased the energy expenditure of mice. Thermogenesis in adipose tissue was induced by reduced FOXO1 expression, which was a cause of the decreased body weight and increased energy expenditure
[29].
3.5. Feeding Behavior in the Hypothalamus
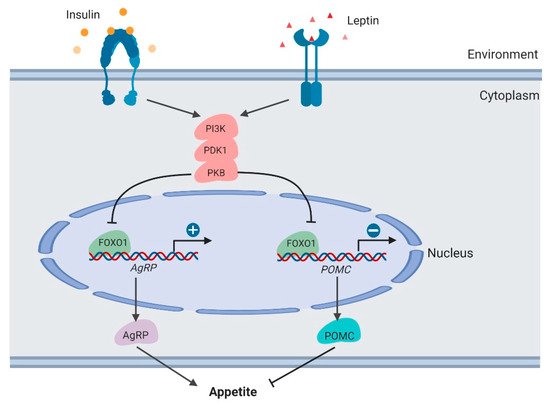
FoxO1 in the central nervous system, mainly the hypothalamus, functions directly in integrating signals from peripheral tissues and mediating feeding behavior. Insulin and leptin are well-studied nutrient signals, integrating peripheral energy status to the hypothalamus. In the arcuate nucleus (ARC) of the hypothalamus, two neuronal populations express specific feeding-related neuropeptides, including pro-opiomelanocortin (POMC) and agouti-related peptide (AgRP)
[46]. POMC suppresses appetite and decreases body weight. AgRP enhances food intake and increases body weight
[47][48].
The
PI3K/PDK1/PKB–FoxO1 signaling axis functions to integrate leptin and insulin signals to regulate POMC and AgRP secretion
[49][50][51] (). As the downstream target of these kinases, the FOXO1 protein is phosphorylated and inactivated in neurons, thus promoting POMC transcription and suppressing AgRP transcription. In mouse models,
PDK1 knockout in POMC neurons suppresses FOXO1 phosphorylation and degradation, thus suppressing POMC transcription. By contrast, in AgRP neurons,
PDK1 deletion results in enhanced AgRP expression and increased food intake. In the
PDK1-knockout mouse models,
FoxO1 inactivation rescues these phenotypes, which demonstrates the crucial roles of
FoxO1 in regulating feeding behavior.
Figure 4. Regulation of appetite by
FOXO1 in the hypothalamus (created with
BioRender.com).
Furthermore, in a DIO rat model, the mechanism of
FoxO1 in regulating feeding behavior was confirmed
[52]. In these rats, because of insulin resistance, the
PI3K/PDK1/PKB-FoxO1 regulatory axis was damaged, resulting in increasing the amount of FOXO1 protein and food intake. However, the intracerebroventricular (i.c.v.) micro-infusion of
FoxO1-antisense oligonucleotides (
FoxO1-ASOs) largely decreased the hypothalamic
FoxO1 expression, food intake, and body weight gain. These suggested that pharmacological inactivation of
FoxO1 may be used to suppress appetite and treat obesity.