1000/1000
Hot
Most Recent

Autism spectrum disorder (ASD) is a complex neurodevelopmental condition affecting behavior and communication, presenting with extremely different clinical phenotypes and features. ASD etiology is composite and multifaceted with several causes and risk factors responsible for different individual disease pathophysiological processes and clinical phenotypes. From a genetic and epigenetic side, several candidate genes have been reported as potentially linked to ASD, which can be detected in about 10–25% of patients. Epigenome signature is a promising field in ASD clinical and translational research.
Currently, autistic disorder, along with pervasive developmental disorder not otherwise specified (PDD–NOS), and Asperger syndrome represent the complex set of human neurodevelopmental diseases collectively known as autism spectrum disorder (ASD) [1]. ASD patients typically experience difficulty with social communication and interaction, restricted interests, and repetitive behaviors [2]. Indeed, despite the available technological advances and the innovative experimental approach of study, there is still not a definite and direct causal relationship or pattern to completely understand the pathogenesis of this complex disorder [3]. ASD has a strong heritable component making it essentially a genome-based human disease [4].
In terms of ASD risk factors definition, there is unanimous agreement on the multifactorial framework of the etiology, suggesting that in most cases autism results from the interaction of multiple genetic and environmental factors, as often demonstrated for other complex diseases [5][6][7][8][9][10][11]. In this line, the role and interplay between genetics and environment with effects on epigenetics and epigenomics have become the subject of intensified researches [3][12][13].
Among those genes associated with modification of disease susceptibility, the ones belonging to the folate homeostasis and methionine-homocysteine recycling are gaining interest in the context of different neuropsychiatric/neurobehavioral disorders [14][15]. In the past decades, several studies indicated that low folate levels and vitamin B12 together with high homocysteine (Hcy) levels were associated with neurodevelopmental disease, especially cognitive decline in psychogeriatric and psychiatric patients [16].
One-carbon metabolism is crucial in epigenetic regulation during embryo development and it is an integrated complex system composed of three main pathways: the folate cycle, the methionine cycle, and the transsulfuration pathway [17]. Overall, folate is crucial during neurodevelopment and it represents an efficient mediator of the crosstalk between genetics and epigenetics [18][19]. Deficiencies or unbalancing of the mutual levels of the different intracellular folate isoforms may negatively act during fetal growth and promote pediatric cancers, leukemia, neurodevelopmental disorders [20][21][22][23][24].
Folate biochemistry has been deeply investigated in the past [25]. Briefly, once in the cells, folate is converted into its tetrahydrofolate (THF) active form by a two-step process dependent on NADPH and DHFR. As shown in Figure 1, 5,10-methylene-THF can be converted to 5-methyl-THF by the MTHFR enzyme and recycled back to THF by methionine synthase (MS). 5-methyl-THF may transfer its methyl group to cobalamin (B12) and the resulting methyl-cobalamin can act as a methyl-donor for Hcy to generate MS-mediated methionine.
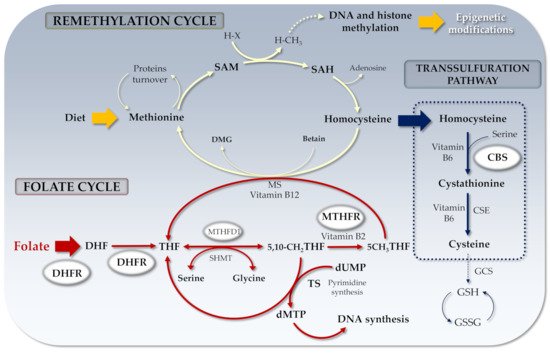
Figure 1. Folate cycle and related genes. DHF: Dihydrofolate; THF: Tetrahydrofolate; SAM: S-adenosyl methionine; SAH: S-adenosyl Homocysteine; MTHFR: Methylenetetrahydrofolate reductase; RCF: Reduced folate carrier; DHFR: Dihydrofolate reductase; dUMP: Uridine monophosphate; dTMP: Thymidine monophosphate; DMG: Dimethylglycine; MS: Methionine synthase; CBS: Cystathionine-β-synthase; H-X: Methyl acceptor; H-CH3: Methylated acceptor; TS: Thymidylate synthase; MTHFD1: Methylenetetrahydrofolate dehydrogenase-1 (NADP+ dependent); CSE: Cystathionine γ-lyase; GCS: γ-glutamylcysteine synthetase; GSH: Glutathione; GSSG: Glutathione disulfide.
Males typically show higher Hcy circulating levels compared to females (about 10–15% higher), though the sex gap is normally reduced by women menopause transition. Folate deficiency is associated with hyperhomocysteinemia, a recognized risk factor for several pathological conditions, with a strong inverse relation driven by the number of variant alleles in the MTHFR gene [25][22]. In a recent study performed in Israel where no national folic acid fortification programs exist, males, as expected, showed the lowest folate levels compared to females [26]. The consequence of low folate on Hcy levels was that males had higher Hcy than females also in presence of normal B12 levels and independently from age and estrogen effect [26].
In many complex diseases, neurological conditions included, sex differences are associated with prevalence, course of the disease and outcome [27][28]. Among behavioral abnormalities, epidemiological studies consistently reported higher ASD prevalence in males compared with females (male: female ratio about 4:1), also considering the different role of abnormalities detected in the mother or father of ASD children [29][27][30][31]. Moreover, ASD females show a less severe disease phenotype highlighting sex differences also in a genetic view [32].
Information and experimental data on sex differences between human brain developments according to the brain epigenome are very limited. Since some of such differences are evident in the brain before birth, they should happen during pregnancy in which sex imprinting actions can be mainly ascribed to the mother and less to the developing fetus. After birth, the genome, epigenome, and gender of the newborn take place, mutually interacting in determining the ultimate brain epigenome (Figure 2).
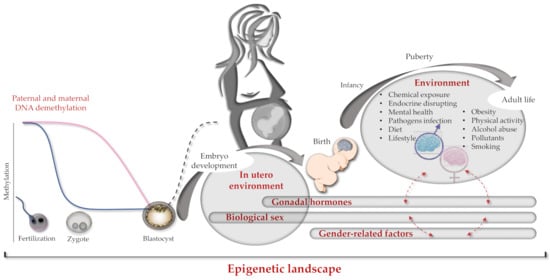
Figure 2. Brain epigenome imprinting. Schematic representation of genetics and epigenetics interactions occurring from conception to individual adult life. On the left, paternal and maternal hemigenome DNA demethylation processes occurring during fertilization (blue and pink lines respectively). In the middle, de novo global genome methylation at the blastocyst phase is shown as a dark dashed line. On the right, dashed red arrows indicate cross-interactions between different epigenetics factors (e.g., sex and gender).
Although the key role of folate isoforms balance/dysregulation in some complex pathological conditions has been demonstrated, more efforts and dedicated studies are needed to conclusively identify the most critical factors involved in ASD establishment. In particular, there is the mandatory need to elucidate the precise mechanisms by which sex-specific factors can modulate disease onset, severity and the different disease phenotypes at presentation, as well as how they can help and guide the choice of useful or unsafe dedicated treatments. To face these challenges, the synergy between advanced technologies and experimental/clinical investigations including pharmacogenetics/genomics/OMICS approaches [33][34][35][36] and personalized medicine [37] will allow a progression in the pathophysiological understanding of complex neurodevelopmental/neurological diseases. Earlier identification of informative molecular and biological biomarkers and an appropriate therapeutic choice are the unique effective strategy to follow (Figure 3). Accordingly, the sex gap in ASD prevalence reported to be about 4:1 (males-to-females respectively) appears to be more realistically close to 3:1 due to a potential diagnostic gender bias that does not efficiently include ASD girls, that are therefore at high risk of not receiving a proper and early diagnosis [38].
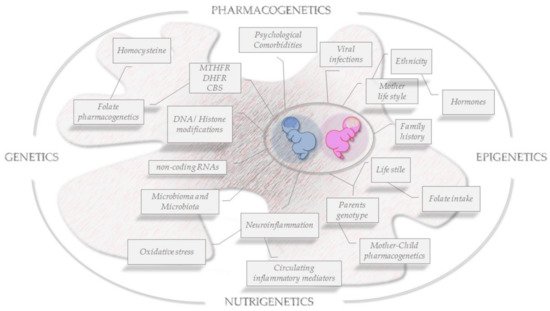
Figure 3. Folate OMICS-environment interactions on brain epigenomics. Snapshot of key risk factors for ASD development and key transgenerational pathophysiological features to be faced in a sex/gender approach.
In conclusion, due to an unavoidable mutual influence between the psychological/social gender and the biological sex in human life, and that they both interact on the brain development by genetics and epigenetics actions, we cannot easily separate the effect of sex or gender on the brain epigenome establishment. Then we must begin to consider sex and gender combined together any time a difference appears in the clinical phenotype between women and men [39].